Abstract
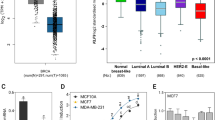
Although the role of core circadian gene cryptochrome 2 (CRY2) in breast tumorigenesis has been demonstrated, the correlations of CRY2 with clinical parameters in breast cancer patients and its involvement in epigenetic processes such as DNA methylation remain relatively unexplored. In the current study, we first queried the Oncomine database and the Gene Expression-Based Outcome for Breast Cancer Online (GOBO) database to identify associations between CRY2 expression levels and clinical parameters in breast cancer patients. We then silenced CRY2 in vitro and performed a genome-wide methylation array to determine the epigenetic impact of CRY2 silencing. The Ingenuity Pathway Analysis software was used to further explore the genes exhibiting altered methylation identified using the array. We found that CRY2 was frequently down-regulated in breast cancer tissue compared to adjacent normal tissue or breast tissue from healthy controls. Lower CRY2 expression was associated with estrogen receptor (ER)-negativity (P < 0.0001), higher tumor grade (P < 0.0001), and shorter overall survival time in breast cancer patients (HR = 1.44, 95 % confidence interval (CI) 1.09–1.91). Genome-wide methylation analysis showed that a total of 515 CpG sites were hypermethylated following CRY2 knockdown, while 730 sites were hypomethylated. The pathway analysis revealed several cancer-relevant networks with genes exhibiting significantly altered methylation following CRY2 silencing. These findings suggest that the core circadian gene CRY2 is associated with breast cancer progression and prognosis, and that knockdown of CRY2 causes the epigenetic dysregulation of genes involved in cancer-relevant pathways, which provide further evidence supporting a role of the circadian system in breast tumorigenesis.



Similar content being viewed by others
References
Kondratov RV, Gorbacheva VY, Antoch MP. The role of mammalian circadian proteins in normal physiology and genotoxic stress responses. Curr Top Dev Biol. 2007;78:173–216.
Reppert SM, Weaver DR. Coordination of circadian timing in mammals. Nature. 2002;418:935–41.
Duffield GE, Best JD, Meurers BH, Bittner A, Loros JJ, Dunlap JC. Circadian programs of transcriptional activation, signaling, and protein turnover revealed by microarray analysis of mammalian cells. Curr Biol. 2002;12:551–7.
Fu L, Lee CC. The circadian clock: pacemaker and tumour suppressor. Nat Rev Cancer. 2003;3:350–61.
Thresher RJ, Vitaterna MH, Miyamoto Y, Kazantsev A, Hsu DS, Petit C, et al. Role of mouse cryptochrome blue-light photoreceptor in circadian photoresponses. Science. 1998;282:1490–4.
van der Spek PJ, Kobayashi K, Bootsma D, Takao M, Eker AP, Yasui A. Cloning, tissue expression, and mapping of a human photolyase homolog with similarity to plant blue-light receptors. Genomics. 1996;37:177–82.
Unsal-Kacmaz K, Mullen TE, Kaufmann WK, Sancar A. Coupling of human circadian and cell cycles by the timeless protein. Mol Cell Biol. 2005;25:3109–16.
Gauger MA, Sancar A. Cryptochrome, circadian cycle, cell cycle checkpoints, and cancer. Cancer Res. 2005;65:6828–34.
Zhu Y, Stevens RG, Hoffman AE, Fitzgerald LM, Kwon EM, Ostrander EA, et al. Testing the circadian gene hypothesis in prostate cancer: a population-based case-control study. Cancer Res. 2009;69:9315–22.
Hoffman AE, Zheng T, Stevens RG, Ba Y, Zhang Y, Leaderer D, et al. Clock-cancer connection in non-Hodgkin’s lymphoma: a genetic association study and pathway analysis of the circadian gene cryptochrome 2. Cancer Res. 2009;69:3605–13.
Hoffman AE, Zheng T, Yi CH, Stevens RG, Ba Y, Zhang Y, et al. The core circadian gene cryptochrome 2 influences breast cancer risk, possibly by mediating hormone signaling. Cancer Prevent Res (Philadelphia, Pa). 2010;3:539–48.
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB, et al. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007;9:166–80.
Turashvili G, Bouchal J, Baumforth K, Wei W, Dziechciarkova M, Ehrmann J, et al. Novel markers for differentiation of lobular and ductal invasive breast carcinomas by laser microdissection and microarray analysis. BMC Cancer. 2007;7:55.
Karnoub AE, Dash AB, Vo AP, Sullivan A, Brooks MW, Bell GW, et al. Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature. 2007;449:557–63.
Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A, et al. X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell. 2006;9:121–32.
Gluck S, Ross JS, Royce M, McKenna Jr EF, Perou CM, Avisar E, et al. Tp53 genomics predict higher clinical and pathologic tumor response in operable early-stage breast cancer treated with docetaxel-capecitabine +/− trastuzumab. Breast Cancer Res Treat. 2012;132:781–91.
Ma XJ, Dahiya S, Richardson E, Erlander M, Sgroi DC. Gene expression profiling of the tumor microenvironment during breast cancer progression. Breast Cancer Res. 2009;11:R7.
Radvanyi L, Singh-Sandhu D, Gallichan S, Lovitt C, Pedyczak A, Mallo G, et al. The gene associated with trichorhinophalangeal syndrome in humans is overexpressed in breast cancer. Proc Natl Acad Sci U S A. 2005;102:11005–10.
Zhao H, Langerod A, Ji Y, Nowels KW, Nesland JM, Tibshirani R, et al. Different gene expression patterns in invasive lobular and ductal carcinomas of the breast. Mol Biol Cell. 2004;15:2523–36.
Finak G, Bertos N, Pepin F, Sadekova S, Souleimanova M, Zhao H, et al. Stromal gene expression predicts clinical outcome in breast cancer. Nat Med. 2008;14:518–27.
McCain J. The cancer genome atlas: new weapon in old war? Biotechnol Healthc. 2006;3:46–51B.
Ringner M, Fredlund E, Hakkinen J, Borg A, Staaf J. GOBO: Gene Expression-Based Outcome for Breast Cancer Online. PLoS One. 2011;6:e17911.
Calvano SE, Xiao W, Richards DR, Felciano RM, Baker HV, Cho RJ, et al. A network-based analysis of systemic inflammation in humans. Nature. 2005;437:1032–7.
Hoffman AE, Zheng T, Ba Y, Stevens RG, Yi CH, Leaderer D, et al. Phenotypic effects of the circadian gene cryptochrome 2 on cancer-related pathways. BMC Cancer. 2010;10:110.
Xiang S, Coffelt SB, Mao L, Yuan L, Cheng Q, Hill SM. Period-2: A tumor suppressor gene in breast cancer. J Circadian Rhythms. 2008;6:4.
Bourguignon LY, Singleton PA, Zhu H, Diedrich F. Hyaluronan-mediated cd44 interaction with rhogef and rho kinase promotes grb2-associated binder-1 phosphorylation and phosphatidylinositol 3-kinase signaling leading to cytokine (macrophage-colony stimulating factor) production and breast tumor progression. J Biol Chem. 2003;278:29420–34.
Orian-Rousseau V, Morrison H, Matzke A, Kastilan T, Pace G, Herrlich P, et al. Hepatocyte growth factor-induced ras activation requires erm proteins linked to both cd44v6 and f-actin. Mol Biol Cell. 2007;18:76–83.
Bourguignon LY, Wong G, Earle C, Krueger K, Spevak CC. Hyaluronan-cd44 interaction promotes c-src-mediated twist signaling, microrna-10b expression, and rhoa/rhoc up-regulation, leading to rho-kinase-associated cytoskeleton activation and breast tumor cell invasion. J Biol Chem. 2010;285:36721–35.
Kai K, Arima Y, Kamiya T, Saya H. Breast cancer stem cells. Breast Cancer. 2010;17:80–5.
Marhaba R, Zoller M. Cd44 in cancer progression: adhesion, migration and growth regulation. J Mol Histol. 2004;35:211–31.
Louderbough JM, Schroeder JA. Understanding the dual nature of cd44 in breast cancer progression. Mol Cancer Res. 2011;9:1573–86.
Vaillant F, Merino D, Lee L, Breslin K, Pal B, Ritchie ME, et al. Targeting bcl-2 with the bh3 mimetic abt-199 in estrogen receptor-positive breast cancer. Cancer Cell. 2013;24:120–9.
Acknowledgments
This work was supported by the National Institutes of Environmental Health Sciences (NIEHS) grant ES018915.
Conflicts of interest
The authors declare that they have no competing interests.
Authors’ contributions
YM carried out the DNA methylation assay, public data search and analysis, and drafted the manuscript. AF and DIJ participated in the data analysis and manuscript preparation. AEH constructed the expression vector and helped with the DNA methylation analysis. MJ and KC participated in the design of the study and helped the statistical analysis. YZ conceived the study and participated in its design and coordination and helped to draft the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1 and 2
(DOC 206 kb)
Rights and permissions
About this article
Cite this article
Mao, Y., Fu, A., Hoffman, A.E. et al. The circadian gene CRY2 is associated with breast cancer aggressiveness possibly via epigenomic modifications. Tumor Biol. 36, 3533–3539 (2015). https://doi.org/10.1007/s13277-014-2989-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2989-3