Abstract
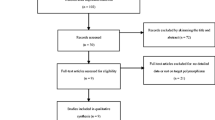
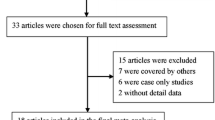
Cytotoxic T lymphocyte antigen 4 (CTLA-4) genetic polymorphisms are implicated to be associated with susceptibility to bone sarcomas, but published studies have reported inconclusive results. The objective of our study was to conduct a meta-analysis investigating the associations between CTLA-4 gene polymorphisms and risk of bone sarcomas. PubMed and Embase databases were searched for all articles published up to June 2, 2013. Odds ratio (OR) with a 95 % confidence interval (95 % CI) was used to assess the association. Finally, 11 individual studies with a total of 2,951 cases with bone sarcomas and 3,396 controls were included in the meta-analysis. There were four studies on the CTLA-4 49G/A polymorphism, three studies on CTLA-4 318C/T polymorphism, two studies on CTLA-4 1661A/G polymorphism, and two studies on CTLA-4 60A/G polymorphism. Overall, CTLA-4 49G/A polymorphism was obviously associated with risk of bone sarcomas (A vs. G: OR = 1.36, 95 % CI = 1.20–1.54; AA vs. GG: OR = 2.24, 95 % CI = 1.67–2.99; AA vs. AG/GG: OR = 2.00, 95 % CI = 1.53–2.62; AA/GA vs. GG: OR = 1.35, 95 % CI = 1.14–1.61). However, CTLA-4 318C/T, 1661A/G, and 60A/G polymorphisms were not associated with risk of bone sarcomas. The current meta-analysis suggests that CTLA-4 49G/A polymorphism is obviously associated with risk of bone sarcomas. More studies are needed to further evaluate the associations between CTLA-4 polymorphisms and risk of bone sarcomas.


Similar content being viewed by others
References
Gaspar N, Di Giannatale A, Geoerger B, Redini F, Corradini N, Enz-Werle N, et al. Bone sarcomas: from biology to targeted therapies. Sarcoma. 2012;2012:301975.
Whelan J, McTiernan A, Cooper N, Wong YK, Francis M, Vernon S, et al. Incidence and survival of malignant bone sarcomas in England 1979–2007. Int J Cancer. 2012;131:E508–17.
ESMO/European Sarcoma Network Working Group. Bone sarcomas: ESMO clinical practice guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2012;23 Suppl 7:vii100–9.
Yarber JL, Agulnik M. Targeted therapies in bone sarcomas: Current approach and future directions. Expert Opin Investig Drugs. 2011;20:973–9.
Grimer R, Athanasou N, Gerrand C, Judson I, Lewis I, Morland B, et al. UK guidelines for the management of bone sarcomas. Sarcoma. 2010;2010:317462.
Chen L, Flies DB. Molecular mechanisms of T cell co-stimulation and co-inhibition. Nat Rev Immunol. 2013;13:227–42.
Ghalamfarsa G, Hadinia A, Yousefi M, Jadidi-Niaragh F. The role of natural killer T cells in B cell malignancies. Tumour Biol. 2013;34:1349–60.
Ramakrishnan R, Gabrilovich DI. Novel mechanism of synergistic effects of conventional chemotherapy and immune therapy of cancer. Cancer Immunol Immunother. 2013;62:405–10.
Deluca DS, Blasczyk R. The immunoinformatics of cancer immunotherapy. Tissue Antigens. 2007;70:265–71.
Ghaderi A. CTLA4 gene variants in autoimmunity and cancer: a comparative review. Iran J Immunol. 2011;8:127–49.
Monjazeb AM, Hsiao HH, Sckisel GD, Murphy WJ. The role of antigen-specific and non-specific immunotherapy in the treatment of cancer. J Immunotoxicol. 2012;9:248–58.
Lens M, Testori A, Ferucci PF. Ipilimumab targeting CD28-CTLA-4 axis: New hope in the treatment of melanoma. Curr Top Med Chem. 2012;12:61–6.
Schmetterer KG, Neunkirchner A, Pickl WF. Naturally occurring regulatory T cells: Markers, mechanisms, and manipulation. FASEB J. 2012;26:2253–76.
Tang ST, Tang HQ, Zhang Q, Wang CJ, Wang YM, Peng WJ. Association of cytotoxic T-lymphocyte associated antigen 4 gene polymorphism with type 1 diabetes mellitus: a meta-analysis. Gene. 2012;508:165–87.
Romo-Tena J, Gomez-Martin D, Alcocer-Varela J. CTLA-4 and autoimmunity: new insights into the dual regulator of tolerance. Autoimmun Rev. 2013;12:1171-6. .
Sun T, Hu Z, Shen H, Lin D. Genetic polymorphisms in cytotoxic t-lymphocyte antigen 4 and cancer: the dialectical nature of subtle human immune dysregulation. Cancer Res. 2009;69:6011–4.
Li M, Zheng H, Li T, Gao P, Zhang XL, Liu DW. Cytotoxic T-lymphocyte associated antigen-4 gene polymorphisms and primary biliary cirrhosis: a systematic review. J Gastroenterol Hepatol. 2012;27:1159–66.
Liu Y, He Z, Feng D, Shi G, Gao R, Wu X, et al. Cytotoxic T-lymphocyte antigen-4 polymorphisms and susceptibility to osteosarcoma. DNA Cell Biol. 2011;30:1051–5.
Wang W, Wang J, Song H, Liu J, Song B, Cao X. Cytotoxic T-lymphocyte antigen-4 + 49G/A polymorphism is associated with increased risk of osteosarcoma. Genet Test Mol Biomarkers. 2011;15:503–6.
Yang S, Wang C, Zhou Y, Sun G, Zhu D, Gao S. Cytotoxic T-lymphocyte antigen-4 polymorphisms and susceptibility to Ewing’s sarcoma. Genet Test Mol Biomarkers. 2012;16:1236–40.
Feng D, Yang X, Li S, Liu T, Wu Z, Song Y, et al. Cytotoxic T-lymphocyte antigen-4 genetic variants and risk of Ewing’s sarcoma. Genet Test Mol Biomarkers. 2013;17:458–63.
Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557–60.
DerSimonian R, Laird N. Meta-analysis in clinical trials. Control Clin Trials. 1986;7:177–88.
Mantel N, Haenszel W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst. 1959;22:719–48.
Yang M, Sun T, Zhou Y, Wang L, Liu L, Zhang X, et al. The functional cytotoxic T lymphocyte-associated protein 4 49G-to-A genetic variant and risk of pancreatic cancer. Cancer. 2012;118:4681–6.
Bharti V, Mohanti BK, Das SN. Functional genetic variants of CTLA-4 and risk of tobacco-related oral carcinoma in high-risk North Indian population. Hum Immunol. 2013;74:348–52.
Gokhale P, Kerkar S, Tongaonkar H, Salvi V, Mania-Pramanik J. CTLA-4 gene polymorphism at position +49 A > G in exon 1: a risk factor for cervical cancer in Indian women. Cancer Genet. 2013;206:154–61.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fan, C., Zhao, X. & Xu, Z. Associations between the cytotoxic T lymphocyte antigen 4 polymorphisms and risk of bone sarcomas. Tumor Biol. 36, 227–231 (2015). https://doi.org/10.1007/s13277-014-2621-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2621-6