Abstract
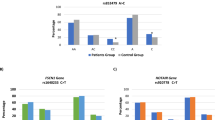
Previous studies indicated that the human X-ray repair complementing group 3 gene (XRCC3) plays an important role in hepatocellular carcinoma (HCC) susceptibility. We aimed to investigate the association of XRCC3 genetic polymorphism with HCC risk. This study was conducted in a Chinese Han population consisting of 300 HCC cases and 300 sex- and age-matched cancer-free controls. Three genetic variants (rs861539, rs12432907, and rs861537) were genotyped by the TaqMan® SNP Genotyping Assay. Our findings suggested that the TT genotype and T allele from rs861539 genetic variants were statistically associated with HCC risk. The TT genotype was statistically associated with the increased risk of HCC compared to CC wild genotype (P < 0.001). And the T allele was more common in the HCC patients than that in the control subjects. (OR = 1.97, 95 % confidence interval (CI) 1.457 ~ 2.659, P < 0.001). Haplotype-based case–control study analysis indicated that TTG haplotype was more frequent in HCC groups than in the control group (odds ratio (OR) = 1.967, 95 % CI 1.456 ~ 2.658); however, the CTG haplotype is more common in the control group than that in the HCC group (OR = 0.550, 95 % CI 0.430 ~ 0.703; P < 0.001). Our data indicated that genetic variants of the XRCC3 gene were statistically associated with HCC risk in a Chinese population.


Similar content being viewed by others
References
Parkin DM, Bray F, Ferlay J, et al. Global cancer statistics, 2002. CA Cancer J Clin. 2005;55(2):74–108.
Llovet JM, Burroughs A, Bruix J. Hepatocellular carcinoma. Lancet. 2003;362(9399):1907–17.
Parikh S, Hyman D. Hepatocellular cancer: a guide for the internist. Am J Med. 2007;120(3):194–202.
But DY, Lai CL, Yuen MF. Natural history of hepatitis-related hepatocellular carcinoma. World J Gastroenterol. 2008;14(11):1652–6.
Yuen MF, Hou JL, Chutaputti A. Hepatocellular carcinoma in the Asia Pacific region. J Gastroenterol Hepatol. 2009;24(3):346–53.
Parkin DM. Global cancer statistics in the year 2000. Lancet Oncol. 2001;2(9):533–43.
Schutte K, Bornschein J, Malfertheiner P. Hepatocellular carcinoma–epidemiological trends and risk factors. Dig Dis. 2009;27(2):80–92.
Duan C, Zhang W, Lu J, Wu H, Liu M, Zhu W. DNA repair gene XRCC3 Thr241Met polymorphism and hepatocellular carcinoma risk. Tumour Biol. 2013;34(5):2827–34. doi:10.1007/s13277-013-0841-9.
Liu C, Wang H. XRCC3 T241M polymorphism is associated risk of hepatocellular carcinoma in the Chinese. Tumour Biol. 2013;34(4):2249–54. doi:10.1007/s13277-013-0765-4.
Gulnaz A, Sayyed AH, Amin F, Khan A, Aslam MA, Shaikh RS, et al. Association of XRCC1, XRCC3, and XPD genetic polymorphism with an increased risk of hepatocellular carcinoma because of the hepatitis B and C virus. Eur J Gastroenterol Hepatol. 2013;25(2):166–79. doi:10.1097/MEG.0b013e328359a775.
Zeng X, Liu S, Yu H, Ji L, Li L, Huang J, et al. DNA repair capacity, DNA-strand break repair gene polymorphisms, and the incidence of hepatocellular carcinoma in southwestern Guangxi of China. DNA Cell Biol. 2012;31(8):1384–91. doi:10.1089/dna.2012.1646.
Girard PM, Graindorge D, Smirnova V, Rigolet P, Francesconi S, Scanlon S, et al. Oxidative stress in mammalian cells impinges on the cysteines redox state of human XRCC3 protein and on its cellular localization. PLoS One. 2013;8(10):e75751.
Lai CY, Hsieh LL, Sung FC, Tang R, Bai CH, Wu FY, et al. Tumor site- and stage-specific associations between allelic variants of glutathione S-transferase and DNA-repair genes and overall survival in colorectal cancer patients receiving 5-fluorouracil-based chemotherapy. PLoS One. 2013;8(7):e69039. doi:10.1371/journal.pone.0069039.
Xie X, Ma YT, Fu ZY, Yang YN, Ma X, Chen BD, et al. Haplotype analysis of the CYP8A1 gene associated with myocardial infarction. Clin Appl Thromb Hemost. 2009;15(5):574–80. doi:10.1177/1076029608329581.
Shi YY, He L. SHEsis, a powerful software platform for analyses of linkage disequilibrium, haplotype construction, and genetic association at polymorphism loci. Cell Res. 2005;15(2):97–8.
Li Z, Zhang Z, He Z, Tang W, Li T, Zeng Z, He L, Shi Y. A partition-ligation-combination-subdivision EM algorithm for haplotype inference with multiallelic markers: update of the SHEsis (http://analysis.bio-x.cn/). Cell Res. 2009;19(4):519–23.
Wu J, Zhang W, Xu A, Zhang L, Yan T, Li Z, et al. Association of epidermal growth factor and epidermal growth factor receptor polymorphisms with the risk of hepatitis B virus-related hepatocellular carcinoma in the population of North China. Genet Test Mol Biomarkers. 2013;17(8):595–600. doi:10.1089/gtmb.2013.0031.
Yu Q, Zhou C, Wang J, Chen L, Zheng S, Zhang J. A functional insertion/deletion polymorphism in the promoter of PDCD6IP is associated with the susceptibility of hepatocellular carcinoma in a Chinese population. DNA Cell Biol. 2013;32(8):451–7. doi:10.1089/dna.2013.2061.
Zhang RC, Mou SH. Polymorphisms of excision repair gene XPD Lys751Gln and hOGG1 Ser326Cys might not be associated with hepatocellular carcinoma risk: a meta-analysis. Tumour Biol. 2013;34(2):901–7.
Hoenerhoff MJ, Pandiri AR, Snyder SA, Hong HH, Ton TV, Peddada S, et al. Hepatocellular carcinomas in B6C3F1 mice treated with Ginkgo biloba extract for two years differ from spontaneous liver tumors in cancer gene mutations and genomic pathways. Toxicol Pathol. 2013;41(6):826–41.
Ng J, Wu J. Hepatitis B- and hepatitis C-related hepatocellular carcinomas in the United States: similarities and differences. Hepat Mon. 2012;12:e7635. doi:10.5812/hepatmon.7635. 10 HCC.
Han X, Xing Q, Li Y, Sun J, Ji H, Huazheng P, et al. Study on the DNA repair gene XRCC1 and XRCC3 polymorphism in prediction and prognosis of hepatocellular carcinoma risk. Hepatogastroenterology. 2012;59(119):2285–9.
Conflicts of interests
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luo, HC., Zhang, HB., Xin, XJ. et al. Haplotype-based case–control study of DNA repair gene XRCC3 and hepatocellular carcinoma risk in a Chinese population. Tumor Biol. 35, 3415–3419 (2014). https://doi.org/10.1007/s13277-013-1451-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-013-1451-2