Abstract
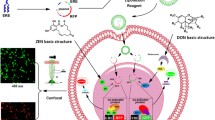
Lysosome is an organelle in the cell, commonly used as biomonitoring tool in environmental pollution. In previous studies, the lysosomal proteome profiling in yeast was analyzed in response to different toxic chemicals, such as sodium meta-arsenite and tetracycline. A recombinant yeast contained the specific biomarker found in the previous studies was constructed for toxic detection. We evaluated green fluorescent intensity of the recombinant yeast exposed to toxic chemicals with dose dependent using by a portable kit designed by our laboratory. The results confirmed the potential of both the yeast and the portable kit to detect each of toxic chemical such as heavy metals, pesticides, or pharmaceuticals at an optimal dose that the intensity of fluorescent protein reached peak at. Therefore, the recombinant yeast and portable kit play an important role to detect various toxic chemicals.
Similar content being viewed by others
References
Miyawaki, A. et al. Fluorescent indicators for Ca2+ based on green fluorescentproteins and calmodulin. Nature 388:882–887 (1997).
Miyawaki, A. Visualization of the spatial and temporal dynamics of intracellular signaling. Dev Cell 4:295–305 (2003).
Zhang, J., Campbell, R. E., Ting, A. Y. & Tsien, R. Y. Creating new fluorescent probes for cell biology. Nat Rev Mol Cell Bio 3:906–918 (2002).
Walmsley, R., Billinton, N. & Heyer, W. Green fluorescent protein as a reporter for the DNA damage-induced gene RAD54 in Saccharomyces cerevisiae. Yeast 13:1535–1545 (1997).
Baumstark-Khan, C., Rode, A., Rettberg, P. & Horneck, G. Application of the Lux-Fluoro test as bioassay for combined genotoxicity and cytotoxicity measurements by means of recombinant Salmonella typhimurium TA1535 cells. Anal Chim Acta 437:23–30 (2001).
Bongaerts, R. J., Hautefort, I., Sidebotham, J. M. & Hinton, J. C. Green fluorescent protein as a marker for conditional gene expression in bacterial cells. Method Enzymol 358:43–66 (2002).
Nguyen, N. T. et al. Toxic detection in mine water based on proteomic analysis of lysosomal enzymes in Saccharomyces cerevisiae. Environ Health Toxicol 29 (2014). http://dx.doi.org/10.5620/eht.e2014019
Liu, L. N., Chen, X. L., Zhang, X. Y., Zhang, Y. Z. & Zhou, B. C. One-step chromatography method for efficient separation and purification of R-phycoerythrin from Polysiphonia urceolata. J Biotechnol 116:91–100 (2005).
Ida, S. et al. Photoluminescence of perovskite nanosheets prepared by exfoliation of layered oxides, K2Ln2Ti3O10, KLnNb2O7, and RbLnTa2O7 (Ln: lanthanide ion). J Am Chem Soc 130:7052–7059 (2008).
Lee, J. H., Park, J. H., Kim, J. S., Lee, D. Y. & Cho, K. High efficiency polymer solar cells with wet deposited plasmonic gold nanodots. Org Electron 10:416–420 (2009).
Lim, E. S. et al. Daphnia magna specific responses to As (III), As (V), and Cd. Toxicol Environ Health Sci 1:196–199 (2009).
Bang, S. H. et al. Acute and chronic toxicity assessment and the gene expression of Dhb, Vtg, Arnt, CYP4, and CYP314 in Daphnia magna exposed to pharmaceuticals. Mol Cell Toxicol 11:153–160 (2015).
Lilius, H., Isomaa, B. & Holmström, T. A comparison of the toxicity of 50 reference chemicals to freshly isolated rainbow trout hepatocytes and Daphnia magna. Aquat Toxicol 30:47–60 (1994).
Peng, Y. et al. Toxic effects of Triclosan on the detoxification system and breeding of Daphnia magna. Ecotoxicology 22:1384–1394 (2013).
Le, T. H. et al. Effects of glyphosate and methidathion on the expression of the Dhb, Vtg, Arnt, CYP4 and CYP314 in Daphnia magna. Chemosphere 79:67–71 (2010).
Schwartz, A. Method of calibrating a fluorescent microscope using fluorescent calibration microbeads simulating stained cells: Google Patents (1989).
Zhang, Y., Kemper, C. R. & Haugland, R. P. Microspheres with fluorescent spherical zones: Google Patents (1998).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Nguyen, NT., Ahn, JY., Bang, S.H. et al. Lysosome based toxic detection in Saccharomyces cerevisiae using novel portable fluorometer. Mol. Cell. Toxicol. 12, 133–138 (2016). https://doi.org/10.1007/s13273-016-0017-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13273-016-0017-y