Abstract
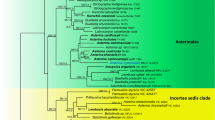
Molecular phylogenetic analyses of LSU rDNA demonstrate monophyly of the genus Melanconiella, and its status as a genus distinct from Melanconis is confirmed. Data of macro- and microscopic morphology, pure cultures and phylogenetic analyses of partial SSU-ITS-LSU rDNA, tef1 and rpb2 sequences revealed 13 distinct species of Melanconiella, six of which are described as new (M. chrysodiscosporina, M. chrysomelanconium, M. chrysorientalis, M. echinata, M. elegans, M. meridionalis). Melanconiella hyperopta var. orientalis is described as a new variety. Diaporthe carpinicola, D. ellisii, D. flavovirens, D. hyperopta and D. ostryae are formally combined into Melanconiella. The name Melanconiella chrysostroma is excluded from Melanconiella, as it is an obligate synonym of Wuestneia xanthostroma. The type of Melanconiella is confirmed as M. spodiaea. Several species are lecto- and/or epitypified. A key to all treated species of Melanconiella is provided, and the circumscriptions of the genera Melanconis and Melanconiella are emended. Most Melanconiella species revealed by molecular phylogenetic analyses can be well characterised by a suite of morphological traits including ascospore shape, length and width, colour, absence/presence and shape of appendages and the anamorph. Anamorph-teleomorph connections were confirmed by pure culture and DNA data, revealing the presence of a single melanconium- or discosporina-like anamorph for each species. Colony growth was found to be characteristic of the respective species. Melanconiella is shown to be confined to the host family Betulaceae, and all species are found to be highly host-specific, mostly confined to a single host species. The biodiversity of Melanconiella was determined to be centred on the genus Carpinus with nine species, five of which have been confirmed on C. betulus. Europe appears to be the geographic centre of Melanconiella biodiversity.

















Similar content being viewed by others
References
Barr ME (1978) The Diaporthales in North America, with emphasis on Gnomonia and its segregates. Mycol Mem 7:1–232
Castlebury LA, Rossman AY, Jaklitsch WJ, Vasilyeva LN (2002) A preliminary overview of the Diaporthales based on large subunit nuclear ribosomal DNA sequences. Mycologia 94:1017–1031
Chaverri P, Samuels GJ (2003) Hypocrea/Trichoderma (Ascomycota, Hypocreales, Hypocreaceae): species with green ascospores. Stud Mycol 48:1–116
Clements FE, Shear CL (1931) Genera of fungi, 2nd edn. H.W. Wilson, New York
de Hoog GS, Gerrits van den Ende AHG (1998) Molecular diagnostics of clinical strains of filamentous basidiomycetes. Mycoses 41:183–189
De Silva H, Castlebury LA, Green S, Stone JK (2009) Characterisation and phylogenetic relationships of Anisogramma virgultorum and A. anomala in the Diaporthales (Ascomycota). Mycol Res 113:73–81
Dennis RWG (1968) British Ascomycetes. J. Cramer, Stuttgart
Druzhinina IS, Kubicek CP, Komoń-Zelazowska M, Mulaw TB, Bissett J (2010) The Trichoderma harzianum demon: complex speciation history resulting in coexistence of hypothetical biological species, recent agamospecies and numerous relict lineages. BMC Evol Biol 10:94
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Ellis MB, Ellis JP (1997) Microfungi on land plants. An identification handbook. Richmond Publishing, Slough
Ellis JB, Everhart BM (1892) North American pyrenomycetes. Ellis & Everhart, Newfield
Fries EM (1849) Summa Vegetabilium Scandinaviae. Sectio posterior. 259–572. A. Bonnier, Stockholm, Leipzig
Fuckel KWGL (1870) Symbolae Mycologicae. Beiträge zur Kenntnis der Rheinischen Pilze. Jahrb Nassau Ver Naturkd 23–24:1–459
Fuckel KWGL (1871) Symbolae Mycologicae. Erster Nachtrag. Jahrb Nassau Ver Naturkd 25–26:289–346
Fuckel KWGL (1874) Symbolae Mycologicae. Zweiter Nachtrag. Jahrb Nassau Ver Naturkd 27–28:1–99
Gazis R, Rehner S, Chaverri P (2011) Species delimitation in fungal endophyte diversity studies and its implications in ecological and biogeographic inferences. Mol Ecol 20:3001–3013
Gerhardt E, Hein B (1979) Die nomenklatorischen Typen der von Th. Nitschke beschriebenen Arten im Pilzherbar des Botanischen Museums Berlin-Dahlem. Willdenowia 9:313–329
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis. program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Jaklitsch WM (2009) European species of Hypocrea part I. The green-spored species. Stud Mycol 63:1–91
Jaklitsch WM (2011) European species of Hypocrea part II. species with hyaline ascospores. Fungal Divers 48:1–250
Jaklitsch WM, Voglmayr H (2004) Hapalocystis occidentalis - a new species of Diaporthales from North America and a key to the species of Hapalocystis. Stud Mycol 50:229–234
Jaklitsch WM, Voglmayr H (2011) Nectria eustromatica sp. nov., an exceptional species with a hypocreaceous stroma. Mycologia 103:209–218
Jaklitsch WM, Komon M, Kubicek CP, Druzhinina IS (2006) Hypocrea voglmayrii sp. nov. from the Austrian Alps represents a new phylogenetic clade in Hypocrea/Trichoderma. Mycologia 97:1365–1378 [‘2005’]
Jaklitsch WM, Stadler M, Voglmayr H (2012) Blue pigment in Hypocrea caerulescens sp. nov. and two additional new species in sect. Trichoderma. Mycologia 104(4):17
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform 9:286–298
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066
Kobayashi T (1970) Taxonomic studies of Japanese Diaporthaceae with special reference to their life-histories. Bull Gov Forest Exp Sta 226:1–242
Liu YL, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol 16:1799–1808
Mejía LC, Castlebury LA, Rossman AY, Sogonov MV, White JF Jr (2011a) A systematic account of the genus Plagiostoma (Gnomoniaceae, Diaporthales) based on morphology, host-associations, and a four-gene phylogeny. Stud Mycol 68:211–235
Mejía LC, Rossman AY, Castlebury LA, White JF Jr (2011b) New species, phylogeny, host-associations and geographic distribution of genus Cryptosporella (Gnomoniaceae, Diaporthales). Mycologia 103:379–399
Montagne C (1834) Notice sur les plantes cryptogames récemment découvertes en France, contenant aussi l’indication précise des localités de quelques espèces les plus rares de la flore Française. Ann Sci Nat Bot Sér 2(1):295–307
Müller E, von Arx JA (1962) Die Gattungen der didymosporen Pyrenomyceten. Beitr Kryptogamenfl Schweiz 11(2):1–922
Munk A (1957) Danish pyrenomycetes. A preliminary flora. Dansk Bot Ark 17:1–491
Nylander JA, Wilgenbusch JC, Warren DL, Swofford DL (2008) AWTY (are we there yet?): a system for graphical exploration of MCMC convergence in Bayesian phylogenetics. Bioinformatics 24:581–583
Otth G (1869) Sechster Nachtrag zu dem in Nr. 15–23 der Mittheilungen enthaltenen Verzeichnisse schweizerischer Pilze. Mitth Natforsch Ges Bern 1868:37–70
Pavlic D, Slippers B, Coutinho TA, Wingfield MJ (2009) Multiple gene genealogies and phenotypic data reveal cryptic species of the Botryosphaeriaceae: a case study on the Neofusicoccum parvum ⁄ N. ribis complex. Mol Phyl Evol 51:259–268
Petrak F (1952a) Fungi beltsvillenses. III. Sydowia 6:5–16
Petrak F (1952b) Fungi beltsvillenses. IV. Sydowia 6:352–360
Petrak F (1962) Mykologische Beiträge zur österreichischen Flora. Sydowia 16:155–198
Reid J, Booth C (1989) On Cryptosporella and Wuestneia. Can J Bot 67:879–908
Riethmüller A, Voglmayr H, Göker M, Weiß M, Oberwinkler F (2002) Phylogenetic relationships of the downy mildews (Peronosporales) and related groups based on nuclear large subunit ribosomal DNA sequences. Mycologia 94:834–849
Saccardo PA (1876) Fungi Veneti novi vel critici. Series V. Nuovo Giornale Botanico Italiano 8:161–211
Saccardo PA (1882) Sylloge Fungorum 1. Padova: published by the author
Saccardo PA (1883) Sylloge Fungorum 2. Padova: published by the author
Saccardo PA (1884) Sylloge Fungorum 3. Padova: published by the author
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W et al. (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA, www.pnas.org/cgi/doi/10.1073/pnas.1117018109
Sieber TN (2007) Endophytic fungi in forest trees: Are they mutualists? Fungal Biol Rev 21:75–89
Silvestro D, Michalak I (2011) raxmlGUI: a graphical front-end for RAxML. Org Divers Evol. doi:10.1007/s13127-011-0056-0
Stamatakis E (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Sutton BC (1980) The Coelomycetes. Commonwealth Mycological Institute, Kew
Swofford DL (2002) PAUP*: Phylogenetic Analysis Using Parsimony (*and Other Methods), Version 4.0b10. Sinauer Associates, Sunderland, Massachusetts
Thiers B (2012) Index Herbariorum: a global directory of public herbaria and associated staff. New York Botanical Garden’s Virtual Herbarium. http://sweetgum.nybg.org/ih/
Tulasne LR, Tulasne C (1863) Selecta Fungorum Carpologia, vol. 2. Paris
Vasilyeva LN, Stephenson SL (2010) Biogeographical patterns in pyrenomycetous fungi and their taxonomy. 1. The Grayan disjunction. Mycotaxon 114:281–303
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246
Voglmayr H, Jaklitsch WM (2008) Prosthecium species with Stegonsporium anamorphs on Acer. Mycol Res 112:885–905
Voglmayr H, Jaklitsch WM (2011) Molecular data reveal high host specificity in the phylogenetically isolated genus Massaria (Ascomycota, Massariaceae). Fungal Divers 46:133–170
von Höhnel F (1918) Mycologische Fragmente. Ann Mycol 16:35–174
von Schweinitz LD (1832) Synopsis fungorum in America boreali media degentium. Trans Am Philos Soc II 4:141–316
Wehmeyer LE (1926) Cultural life histories of Melanconis and Pseudovalsa. Mycologia 18:257–273
Wehmeyer LE (1933) The genus Diaporthe Nitschke and its segregates. Univ Michigan Stud Sci Ser 9:1–349
Wehmeyer LE (1937) Studies of certain species of Melanconis on Carpinus, Ostrya and Corylus. Mycologia 29:599–617
Wehmeyer LE (1941) A revision of Melanconis, Pseudovalsa, Prosthecium and Titania. Univ Michigan Stud, Sci Ser 14:1–161
Werle E, Schneider C, Renner M, Völker M, Fiehn W (1994) Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res 22:4354–4355
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Acknowledgements
We thank Walter Gams for hospitality and excursion support in Italy; Jacques Fournier, Enrique Rubio Domínguez, Sven-Åke Hanson and Larissa Vasilyeva for collecting and communicating Melanconiella specimens; Irmgard Greilhuber and her family for organising and participating in numerous collecting trips together with HV; the fungarium curators of B, BPI, DAOM, G, GZU, K, M, NY, UPS and W for the loan of specimens; Scott Redhead (DAOM) for providing notes of L. E. Wehmeyer and for allowing DNA extraction from the type specimen of M. echinata; Walter Till (WU) for managing the herbarium loans; and the British Mycological Society for invitation to the BMS Spring Foray 2011 in Yorkshire.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Voglmayr, H., Rossman, A.Y., Castlebury, L.A. et al. Multigene phylogeny and taxonomy of the genus Melanconiella (Diaporthales). Fungal Diversity 57, 1–44 (2012). https://doi.org/10.1007/s13225-012-0175-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-012-0175-8