Abstract
Comparative transcriptomics has gained increasing popularity in genomic research thanks to the development of high-throughput technologies including microarray and next-generation RNA sequencing that have generated numerous transcriptomic data. An important question is to understand the conservation and divergence of biological processes in different species. We propose a testing-based method TROM (Transcriptome Overlap Measure) for comparing transcriptomes within or between different species, and provide a different perspective, in contrast to traditional correlation analyses, about capturing transcriptomic similarity. Specifically, the TROM method focuses on identifying associated genes that capture molecular characteristics of biological samples, and subsequently comparing the biological samples by testing the overlap of their associated genes. We use simulation and real data studies to demonstrate that TROM is more powerful in identifying similar transcriptomes and more robust to stochastic gene expression noise than Pearson and Spearman correlations. We apply TROM to compare the developmental stages of six Drosophila species, C. elegans, S. purpuratus, D. rerio and mouse liver, and find interesting correspondence patterns that imply conserved gene expression programs in the development of these species. The TROM method is available as an R package on CRAN (https://github.com/Vivianstats/TROM) with manuals and source codes available at http://jsb.ucla.edu/trom-transcriptome-overlap-measure.






Similar content being viewed by others
References
Arbeitman MN, Furlong EE, Imam F, Johnson E, Null BH, Baker BS, Krasnow MA, Scott MP, Davis RW, White KP (2002) Gene expression during the life cycle of Drosophila melanogaster. Science 297(5590):2270–2275
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT et al (2000) Gene ontology: tool for the unification of biology. Nat Genet 25(1):25–29
Bolstad BM, Irizarry RA, Åstrand M, Speed TP (2003) A comparison of normalization methods for high density oligonucleotide array data based on variance and bias. Bioinformatics 19(2):185–193
Cunningham F, Amode MR, Barrell D, Beal K, Billis K, Brent S, Carvalho-Silva D, Clapham P, Coates G, Fitzgerald S et al (2015) Ensembl 2015. Nucl Acids Res 43(D1):D662–D669
Davidson EH, Cameron RA, Ransick A (1998) Specification of cell fate in the sea urchin embryo: summary and some proposed mechanisms. Development 125(17):3269–3290
Domazet-Lošo T, Tautz D (2010) A phylogenetically based transcriptome age index mirrors ontogenetic divergence patterns. Nature 468(7325):815–818
Dong Z, Wei H, Sun R, Tian Z (2007) The roles of innate immune cells in liver injury and regeneration. Cell Mol Immunol 4(4):241–252
Fu X, Fu N, Guo S, Yan Z, Xu Y, Hu H, Menzel C, Chen W, Li Y, Zeng R et al (2009) Estimating accuracy of RNA-Seq and microarrays with proteomics. BMC Genom 10(1):161
Gerstein MB, Rozowsky J, Yan KK, Wang D, Cheng C, Brown JB, Davis CA, Hillier L, Sisu C, Li JJ et al (2014) Comparative analysis of the transcriptome across distant species. Nature 512(7515):445–448
Hata S, Namae M, Nishina H (2007) Liver development and regeneration: from laboratory study to clinical therapy. Develop Growth Differ 49(2):163–170
Hicks SC, Irizarry RA (2014) When to use quantile normalization? bioRxiv. doi:https://doi.org/10.1101/012203
Labbé RM, Irimia M, Currie KW, Lin A, Zhu SJ, Brown DD, Ross EJ, Voisin V, Bader GD, Blencowe BJ et al (2012) A comparative transcriptomic analysis reveals conserved features of stem cell pluripotency in planarians and mammals. Stem Cells 30(8):1734–1745
Le HS, Oltvai ZN, Bar-Joseph Z (2010) Cross-species queries of large gene expression databases. Bioinformatics 26(19):2416–2423
Li JJ, Huang H, Bickel PJ, Brenner SE (2014) Comparison of D. melanogaster and C. elegans developmental stages, tissues, and cells by modencode RNA-Seq data. Genome Res 24(7):1086–1101
Li T, Huang J, Jiang Y, Zeng Y, He F, Zhang MQ, Han Z, Zhang X (2009) Multi-stage analysis of gene expression and transcription regulation in c57/b6 mouse liver development. Genomics 93(3):235–242
Necsulea A, Soumillon M, Warnefors M, Liechti A, Daish T, Zeller U, Baker JC, Grützner F, Kaessmann H (2014) The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505(7485):635–640
Pantalacci S, Sémon M (2015) Transcriptomics of developing embryos and organs: a raising tool for evo–devo. J Exp Zool Part B Mol Dev Evol 324(4):363–371
Puniyani K, Faloutsos C, Xing EP (2010) Spex2: automated concise extraction of spatial gene expression patterns from fly embryo ISH images. Bioinformatics 26(12):i47–i56
Shen Y, Yue F, McCleary DF, Ye Z, Edsall L, Kuan S, Wagner U, Dixon J, Lee L, Lobanenkov VV et al (2012) A map of the cis-regulatory sequences in the mouse genome. Nature 488(7409):116–120
Spencer WC, Zeller G, Watson JD, Henz SR, Watkins KL, McWhirter RD, Petersen S, Sreedharan VT, Widmer C, Jo J et al (2011) A spatial and temporal map of C. elegans gene expression. Genome Res 21(2):325–341
Tong X, Feng Y, Li JJ (2016) Neyman-Pearson (NP) classification algorithms and NP receiver operating characteristic (NP-ROC) curves. arXiv preprint arXiv:1608.03109
Tu Q, Cameron RA, Davidson EH (2014) Quantitative developmental transcriptomes of the sea urchin Strongylocentrotus purpuratus. Dev Biol 385(2):160–167
Virmani AK, Tsou JA, Siegmund KD, Shen LY, Long TI, Laird PW, Gazdar AF, Laird-Offringa IA (2002) Hierarchical clustering of lung cancer cell lines using DNA methylation markers. Cancer Epidemiol Biomark Prevent 11(3):291–297
Wang C, Gong B, Bushel PR, Thierry-Mieg J, Thierry-Mieg D, Xu J, Fang H, Hong H, Shen J, Su Z et al (2014) The concordance between RNA-seq and microarray data depends on chemical treatment and transcript abundance. Nat Biotechnol 32(9):926–932
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet 10(1):57–63
Zhao S, Fung-Leung WP, Bittner A, Ngo K, Liu X (2014) Comparison of RNA-Seq and microarray in transcriptome profiling of activated T cells. PloS One 9(1)
Author information
Authors and Affiliations
Corresponding author
Appendix
Appendix
See Tables 2, 3 and Figs. 7, 8, 9, 10, 11, 12, 13, 14, 15, and 16.
Correspondence maps of within-species and between-species TROM scores (calculated based on the z-score thresholds listed in the table). TROM scores are saturated at 6. The names of the species are marked as row or column labels of the corresponding heatmaps. For the Drosophila species, the stages labels 1–13 refer to Embryo 0–2, 2–4, 4–6, 6–8, 8–10, 10–12, 12–14, 14–16, 16–18, 18–20, 20–22, 22–24 and 24–26 h, respectively
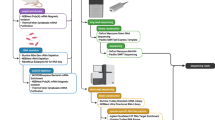
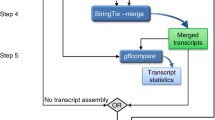
Outline of the package TROM. The three left (and right) heatmaps illustrate the within-species (and between-species) comparison results by using TROM (with z-score threshold 1.5 for both fly and worm), Pearson correlation and Spearman correlation. Greater similarities are shown in darker colors. The results show that compared to the popular Pearson and Spearman correlations, TROM can find clearer correspondence patterns. TROM takes gene expression matrices and orthologous genes of the species of interest as input. The functions select.associated.genes and select.associated.orthologs select the associated genes of different biological samples among all the genes or only among the genes with orthologs in the other species to be compared with. They also provide graphical summaries of the numbers of selected associated genes and orthologs. The functions ws.trom and ws.trom.orthologs perform the within-species transcriptome comparison, find the overlapping associated genes between every two samples and calculate within-species TROM scores. The function bs.trom performs the between-species transcriptome comparison, find the overlapping associated orthologs between every two samples from different species and calculate the between-species TROM scores. The function heatmap.3 visualizes the TROM scores in a heatmap, with various add-on options for customization. The functions find.top.GO.terms and find.top.GO.slim.terms perform gene set enrichment analysis and find top enriched Gene Ontology (GO) terms and GO slim terms in the associated genes. Instead of using the selected associated genes, users may input customized gene lists representing characteristics of different biological samples into the above functions. Please see the package manual and vignette of TROM for details
Correlation measures calculated based on the union of associated genes. Pearson correlation (a) and Spearman correlation (b) for every pair of D. melanogaster stages calculated based on the union of associated genes of all stages. Pearson correlation (c) and Spearman correlation (d) for every pair of D. melanogaster and C. elegans stages calculated based on the union of associated ortholog pairs of all stages. These heatmaps show that correlation measures calculated based on associated genes only still cannot lead to clear correspondence patterns
Comparison of TROM and Pearson/Spearman correlation on simulated data, with a for fly and b for worm. In both panels, the first row gives the true sample relationships (1: high dependence in associated genes; 0: otherwise) defined as in Eq. 6 for varying c. The second row gives the mean receiver operating characteristic (ROC) curves on the 200 simulated gene expression matrices, given the true labels in the first row. The third row gives the mean Neyman–Pearson receiver operating characteristic (NP-ROC) curves, accordingly. The 95 % confidence intervals of the area under the curve (AUC) are marked next to the curves
Top 20 enriched biological process GO terms of D.melanogaster. The enrichment scores in the heatmap are calculated based on stage-associated genes identified from the RNA-seq data (with z-score threshold 1.5) and saturated at 6. For each stage, the common enriched GO terms identified from both microarray (Fig. 13) and RNA-seq datasets are marked in red color (Color figure online)
Top 20 enriched biological process GO terms of D.melanogaster. The enrichment scores in the heatmap are calculated based on the stage-associated genes identified from the microarray data (with z-score threshold 0.5) and saturated at 6. For each stage, the common enriched GO terms identified from both microarray and RNA-seq (Fig. 12) datasets are marked in red color (Color figure online)
Top 10 enriched biological process GO terms of mouse liver. The enrichment scores in the heatmap are calculated based on the stage-associated genes identified from the microarray data (with z-score threshold 1.5). For each stage, the highly relevant GO terms that have been confirmed in previous studies are marked in red color (Color figure online)
Spearman correlations of the developmental stages of D. melanogaster (fly) and C. elegans (worm). a The first panel shows the original Spearman correlations of fly stages, while the rest panels show the Spearman correlations of fly stages under different thresholds. b The first panel shows the original Spearman correlations of worm stages, while the rest panels show Spearman correlations of worm stages under different thresholds. c TROM scores of fly. d TROM scores of worm. All the values under the selected threshold are set to the minimum value of each correlation matrix
Rights and permissions
About this article
Cite this article
Li, W.V., Chen, Y. & Li, J.J. TROM: A Testing-Based Method for Finding Transcriptomic Similarity of Biological Samples. Stat Biosci 9, 105–136 (2017). https://doi.org/10.1007/s12561-016-9163-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12561-016-9163-y