Abstract
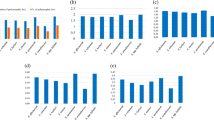
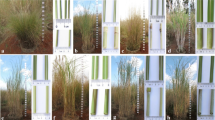
A basic understanding of the cytoplasmic diversity of Saccharum complex comprising of Saccharum species and related genera is essential for broadening the cytoplasmic base of sugarcane cultivars. In the present study markers generated by nine chloroplast microsatellite primer pairs designed from sugarcane chloroplast genome were used to establish the genetic relationship and phylogeny among 43 accessions belonging to different species of Saccharum and related genera representing the Saccharum complex. The polymorphic primer pairs amplified an average of six alleles per loci and possess relatively high polymorphism information content. Molecular diversity was estimated using 54 polymorphic markers. The overall chloroplast genetic diversity in the Saccharum complex was found to be 53 %. The genetic diversity among Saccharum species was 22 %. Among the Saccharum species S. spontaneum was found to be distinct from other species of Saccharum and cpSSR markers could efficiently distinguish S. spontaneum from rest of the Saccharum spcies. Ripidium, one of the most primitive among the Saccharum complex showed the highest genetic diversity (90 %) with Saccharum. Miscanthus, another primitive genus of Saccharum complex was found to be more closely related to Saccharum than Erianthus. In view of the high genetic diversity, the present study supports the utilization of different species of Erianthus in the sugarcane breeding programmes for broadening the cytoplasmic base of the sugarcane varieties.




Similar content being viewed by others
References
Al-Janabi, S.M., M. McClelland, C. Petersen, and B.W.S. Sobral. 1994. Phylogenetic analysis of organellar DNA sequences in the Andropogoneae: Saccharine. Theoretical and Applied Genetics 88: 933–944.
Anderson, J.A., G.A. Churchill, J.E. Autroque, S.D. Tanksley, and M.E. Swells. 1993. Optimizing parental selection for genetic linkage maps. Genome 36: 181–186.
Alwala, S., A. Suman, J.A. Arro, J.C. Vermis, and C.A. Kimbeng. 2006. Target region amplification polymorphism (TRAP) for accessing genetic diversity in sugarcane germplasm collections. Crop Science 46: 448–455.
Asano, T., T. Tsudzuki, S. Takahashi, H. Shimada, and K. Kadowaki. 2004. Complete nucleotide sequence of the sugarcane (Saccharum officinarum) chloroplast genome: A comparative analysis of four monocot chloroplast genomes. DNA Research 11: 93–99.
Bakshi Ram, N.V. Nair, C.M. Radhakrishnan, N. Singh, and B.K. Sahi. 2007. Effect of cytoplasmic diversity on performance of sugarcane hybrids. The Indian Journal of Genetics 67: 229–231.
Burnquist, W.L., M.E. Sorrells, and S.D. Tanksley. 1995. Characterization of genetic variability in Saccharum germplasm by means of restriction fragment length polymorphism (RFLP) analysis. Proceedings of the International Society of Sugar Cane Technologists 21: 355–365.
Clegg, M.T. 1993. Chloroplast gene sequences and the study of plant evolution. Proceedings of the National Academy of Sciences of the United States of America 90: 363–367.
Daniels, J., and B.T. Roach. 1987. Taxonomy and evolution. In Sugarcane improvement through breeding, ed. D.J. Heinz, 7–84. Amsterdam: Elsevier.
Devarumath, R., S. Kalwade, P. Bundock, F.G. Eliott, and R. Henry. 2013. Independent target region amplification polymorphism and single-nucleotide polymorphism marker utility in genetic evaluation of sugarcane genotypes. Plant Breeding 132: 736–747.
D’Hont, A., Y.H. Lu, P. Feldmann, and J.C. Glaszmann. 1993. Cytoplasmic diversity in sugarcane revealed by heterologous probes. Sugar Cane 1: 12–15.
Irvine, J.E. 1999. Saccharum species as horticultural classes. Theoretical and Applied Genetics 98: 186–194.
Liu, P., Y. Que, and Y.-B. Pan. 2011. Highly polymorphic microsatellite DNA markers for sugarcane germplasm evaluation and variety identity testing. Sugar Tech 13(2): 129–136.
Lu, Y.H., A. D’Hont, D.I.T. Walker, P. Feldman, P.S. Rao, and J.C. Glassmann. 1994. Relationships among ancestral species of sugarcane revealed by RFLP using single-copy maize nuclear probes. Euphytica 78: 7–18.
Mangelsdorf, A.J. 1983. Cytoplasmic diversity in relation to pest and pathogens. Sugarcane Breeders Newsletter 45: 45–49.
Marshall, H.D., C. Newton, and K. Ritland. 2001. Sequence-repeat polymorphisms exhibit the signature of recombination in lodgepole pine chloroplast DNA. Molecular Biology and Evolution 18: 2136–2138.
Nair, N.V., S. Nair, T.V. Sreenivasan, and M. Mohan. 1999. Analysis of genetic diversity and phylogeny in Saccharum and related genera using RAPD markers. Genetic Resources and Crop Evolution 46: 73–79.
Nair, N.V. 1999. Production and cyto-morphological analysis of intergeneric hybrids of Sorghum x Saccharum. Euphytica 108: 187–191.
Powell, W., G.C. Machray, and J. Provan. 1996. Polymorphism revealed by simple sequence repeats. Trends in Plant Science 1: 215–222.
Rajendrakumar, P., A.K. Biswal, S.M. Balachandran, K. Srinivasarao, and R.M. Sundaram. 2007. Simple sequence repeats in organellar genomes of rice: Frequency and distribution in genic and intergenic regions. Bioinformatics 23: 1–4.
Selvi, A., N.V. Nair, N. Balasundaram, and T. Mohapatra. 2003. Evaluation of maize microsatellite markers for genetic diversity analysis and fingerprinting in sugar cane. Genome 46: 394–403.
Selvi, A., N.V. Nair, J.L. Noyer, N.K. Singh, N. Balasundaram, K.C. Bansal, K.R. Koundal, and T. Mohapatra. 2006. AFLP analysis of the phenetic organization and genetic diversity in the sugarcane complex, Saccharum and Erianthum. Genetic Resources and Crop Evolution 53: 831–842.
Selvi, A., O. Garsmeur, and A. D’Hont. 2008. Phylogenetic analysis of sugarcane. In: Deputation report of the Department of Biotechnology overseas associateship, submitted to the Department of Biotechnology, India, p. 3.
Singh, R.K., S.K. Mishra, S.P. Singh, N. Mishra, and M.L. Sharma. 2010. Evaluation of microsatellite markers for genetic diversity analysis among sugarcane species and commercial hybrids. Australian Journal of Crop Science 4: 115–124.
Sobral, B.W.S., D.P.V. Braga, E.S. LaHood, and P. Keim. 1994. Phylogenetic analysis of chloroplast restriction enzyme site mutations in the Saccharinae Griseb, subtribe of the Andropogoneae. Theoretical and Applied Genetics 87: 843–853.
Suman, A., C.A. Kimbeng, S.J. Edme, and J. Vermis. 2008. Sequence related amplified polymorphism (SRAP) markers for accessing genetic relationship and diversity in sugarcane germplasm collections. Plant Genetic Resources 6: 222–231.
Takahashi, S., T. Furukawa, T. Asano, and Y. Terajima. 2005. Very close relationship of the chloroplast genomes among Saccharum species. Theoretical and Applied Genetics 110: 1523–1529.
Tambarussi, E.V., D.M. Melotto-Passarin, S.G. Gonzalez, J.B. Brigati, F.A. de Jesus, A.L. Barbosa, K. Dressano, and H. Carrer. 2009. In silico analysis of simple sequence repeats from chloroplast genomes of Solanaceae species. Crop Breeding and Applied Biotechnology 9: 344–352.
Walbot, V. 1998. Preparation of DNA from rice seedlings. Rice Genetics Newsletters 5: 149–151.
Zhu, J.R., H. Zhou, Y. Pan, and X. Lu. 2014. Genetic variability among the chloroplast genomes of sugarcane (Saccharum spp.) and its wild progenitor species Saccharum spontaneum L. Genetics and Molecular Research 13(2): 3037–3047.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Raj, P., Selvi, A., Prathima, P.T. et al. Analysis of Genetic Diversity of Saccharum Complex Using Chloroplast Microsatellite Markers. Sugar Tech 18, 141–148 (2016). https://doi.org/10.1007/s12355-015-0382-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-015-0382-1