Abstract
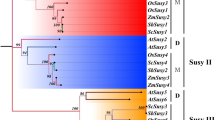
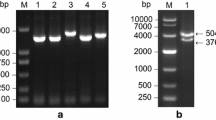
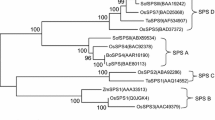
Sucrose synthase is one of the key enzymes involved in sucrose metabolism. In order to increase the sucrose content at the molecular level, it is very necessary to understand the biological function of sucrose synthase gene in sugarcane. In this study, homology cloning and rapid amplification of cDNA ends technology were used to obtain the 2898 bp full-length cDNA of ScSuSy4 in sugarcane, which contains an opening reading frame of 2430 bp (GenBank: KM598653). This gene encodes a protein of 809 amino acids with a theoretical molecular weight of 92.31 kDa and an isoelectric point of 6.30. The amino acid sequence homologies between ScSuSy4 and SbSus4, ZmSus3 and OsSus4 from Group II were 99.5, 98.3, and 97.2 %, respectively. Expression analysis showed that at the elongation stage of low-sugar and medium-sugar sugarcane species, the expression level of ScSuSy4 in immature stems was higher than that in mature leaves, whereas in all three high-sugar species, ScSuSy4 gene expression levels were higher in mature leaves than that in immature stems. At the early stage of sucrose accumulation, expression levels of ScSuSy4 in all species were lower in immature stems than that in mature leaves. At different stages of the same species, regardless in immature stems or in mature leaves, ScSuSy4 gene expression reached the highest level at the elongation stage, dropped to the lowest level at early stage of sucrose accumulation, and increased again at late stage of sucrose accumulation. It is putative that ScSuSy4 gene might be involved in the regulation of sucrose distribution via different functions at various stages. This study provides a foundation for further dissection of the biological function of this gene.





Similar content being viewed by others
References
Baroja-Fernández, E., F.J. Muñoz, M. Montero, E. Etxeberria, M.T. Sesma, M. Ovecka, A. Bahaji, I. Ezquer, J. Li, S. Prat, and J. Pozueta-Romero. 2009. Enhancing sucrose synthase activity in transgenic potato (Solanum tuberosum L.) tubers results in increased levels of starch, ADPglucose and UDPglucose and total yield. Plant and Cell Physiology 50(9): 1651–1662.
Bieniawska, Z., D.H. Paul Barratt, A.P. Garlick, V. Thole, N.J. Kruger, C. Martin, R. Zrenner, and A.M. Smith. 2007. Analysis of the sucrose synthase gene family in Arabidopsis. Plant Journal 49: 810–828.
Chen, A., S. He, F. Li, Z. Li, M. Ding, Q. Liu, and J. Rong. 2012. Analyses of the sucrose synthase gene family in cotton: structure, phylogeny and expression patterns. BMC Plant Biology 12: 85.
Coleman, H.D., J. Yan, and S.D. Mansfield. 2009. Sucrose synthase affects carbon partitioning to increase cellulose production and altered cell wall ultrastructure. Proceedings of the National Academy of Sciences 106(31): 13118–13123.
Duncan, K.A., S.C. Hardin, and S.C. Huber. 2006. The three maize sucrose synthase isoforms differ in distribution, localization, and phosphorylation. Plant Cell Physiology 47(7): 959–971.
Hirose, T., G.N. Scofield, and T. Terao. 2008. An expression analysis profile for the entire sucrose synthase gene family in rice. Plant Science 174: 534–543.
Jiang, Q., J. Hou, C. Hao, L. Wang, H. Ge, Y. Dong, and X. Zhang. 2011. The wheat (T. aestivum) sucrose synthase 2 gene (TaSus2) active in endosperm development is associated with yield traits. Functional and Integrative Genomics 11: 49–61.
Komatsu, A., T. Moriguchi, K. Koyama, M. Omura, and T. Akiyama. 2002. Analysis of sucrose synthase genes in citrus suggests different roles and phylogenetic relationship. Journal of Experimental Botany 53: 61–71.
Lei, M.H., B.Y. Ye, B.M. Wang, Z.X. Huang, H. Zhang, L.P. Xu, Y.Q. Chen, and R.K. Chen. 2008. Cloning of sucrose synthase gene from sugarcane (Saccharum officinarum L.). Chinese Journal of Applied and Environmental Biology 14(2): 177–179.
Li, H.P., H.M. Li, and S.M. Pan. 2013a. Cloning and sequence analysis of a chewing cane sucrose synthase gene (sosus1) and investigation of expression profiling. Biotechnology Bulletin 11: 58–62.
Li, J., E. Baroja-Fernández, A. Bahaji, F.J. Muñoz, M. Ovecka, M. Montero, M.T. Sesma, N. Alonso-Casajús, G. Almagro, A.M. Sánchez-López, M. Hidalgo, M. Zamarbide, and J. Pozueta-Romero. 2013b. Enhancing sucrose synthase activity results in increased levels of starch and ADP-glucose in maize (Zea mays L.) seed endosperms. Plant and Cell Physiology 54(2): 282–294.
Lingle, S.E., and J.M. Dyer. 2001. Cloning and expression of sucrose synthase-1 cDNA from sugarcane. Journal of Plant Physiology 158: 129–131.
Ruan, Y.L., D.J. Llewellyn, and R.T. Furbank. 2003. Suppression of sucrose synthase gene expression represses cotton fiber cell initiation, elongation, and seed development. Plant Cell 15: 952–964.
Tang, G.Q., and A. Sturm. 1999. Antisense repression of sucrose synthase in carrot (Daucus carota L.) affects growth rather than sucrose partitioning. Plant Molecular Biology 41(4): 465–479.
Wang, H., X. Sui, J. Guo, Z. Wang, J. Cheng, S. Ma, X. Li, and Z. Zhang. 2014. Antisense suppression of cucumber (Cucumis sativus L.) sucrose synthase 3 (CsSUS3) reduces hypoxic stress tolerance. Plant, Cell and Environment 37(3): 795–810.
Xiao, X., C. Tang, Y. Fang, M. Yang, B. Zhou, J. Qi, and Y. Zhang. 2014. Structure and expression profile of the sucrose synthase gene family in the rubber tree: indicative of roles in stress response and sucrose utilization in the laticifers. FEBS Journal 281(1): 291–305.
Xu, S.M., E. Brill, D.J. Llewellyn, R.T. Furbank, and Y.L. Ruan. 2012. Overexpression of a potato sucrose synthase gene in cotton accelerates leaf expansion, reduces seed abortion, and enhances fiber production. Molecular Plant 2: 430–441.
Yang, J., J. Zhang, Z. Wang, G. Xu, and Q. Zhu. 2004. Activities of key enzymes in sucrose-to-starch conversion in wheat grains subjected to water deficit during grain filling. Plant Physiology 135: 1621–1629.
Zhang, D., B. Xu, X. Yang, Z. Zhang, and B. Li. 2011. The sucrose synthase gene family in Populus: structure, expression, and evolution. Tree Genetics and Genomes 7(3): 443–456.
Zhang, J., J. Arro, Y. Chen, and R. Ming. 2013. Haplotype analysis of sucrose synthase gene family in three Saccharum species. BMC Genomics 14: 314.
Zou, C., C. Lu, H. Shang, X. Jing, H. Cheng, Y. Zhang, and G. Song. 2013. Genome-wide analysis of the Sus gene family in cotton. Journal of Integrative Plant Biology 55(7): 643–653.
Acknowledgments
This work was supported by National High-tech R&D Program (863 Program) project (2013AA102604), Natural Science Foundation of China (31160301), National Program for International Scientific Exchange projects (2013DFA31600), Guangxi Funds for Bagui Scholars, 948 Project of the Ministry of Agriculture, China (2013-S13), Natural Science Foundation of Guangxi (2011GXNSFF018002), and Fundamental Research Fund of Guangxi Academy of Agriculture Sciences (Guinongke 2014JZ02, Guinongke 2013YM41, Guinongke 2012YZ11).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Zhong-Liang Chen and Yi-Yun Gui have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Chen, ZL., Gui, YY., Qin, CX. et al. Isolation and Expression Analysis of Sucrose Synthase Gene (ScSuSy4) from Sugarcane. Sugar Tech 18, 134–140 (2016). https://doi.org/10.1007/s12355-015-0372-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-015-0372-3