Abstract
This study aims to explore the potential mechanism of glioma through bioinformatic approaches. The gene expression profile (GSE4290) of glioma tumor and non-tumor samples was downloaded from Gene Expression Omnibus database. A total of 180 samples were available, including 23 non-tumor and 157 tumor samples. Then the raw data were preprocessed using robust multiarray analysis, and 8,890 differentially expressed genes (DEGs) were identified by using t-test (false discovery rate < 0.0005). Furthermore, 16 known glioma related genes were abstracted from Genetic Association Database. After mapping 8,890 DEGs and 16 known glioma related genes to Human Protein Reference Database, a glioma associated protein-protein interaction network (GAPN) was constructed. In addition, 51 sub-networks in GAPN were screened out through Molecular Complex Detection (score ≥ 1), and sub-network 1 was found to have the closest interaction (score = 3). What’ more, for the top 10 sub-networks, Gene Ontology (GO) enrichment analysis (p value < 0.05) was performed, and DEGs involved in sub-network 1 and 2, such as BRMS1L and CCNA1, were predicted to regulate cell growth, cell cycle, and DNA replication via interacting with known glioma related genes. Finally, the overlaps of DEGs and human essential, housekeeping, tissue-specific genes were calculated (p value = 1.0, 1.0, and 0.00014, respectively) and visualized by Venn Diagram package in R. About 61 % of human tissue-specific genes were DEGs as well. This research shed new light on the pathogenesis of glioma based on DEGs and GAPN, and our findings might provide potential targets for clinical glioma treatment.





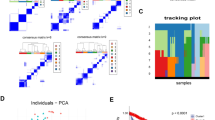
Similar content being viewed by others
Abbreviations
- DEGs:
-
Differentially expressed genes
- GAPN:
-
Glioma associated protein-protein interaction network
- GO:
-
Gene Ontology
- NCBI:
-
National Center for Biotechnology Information
- GEO:
-
Gene Expression Omnibus
- RMA:
-
Robust multiarray analysis
- Gene ID:
-
Gene identifier
- FDR:
-
False discovery rate
- GAD:
-
Genetic Association Database
- HPRD:
-
Human Protein Reference Database
- PPIs:
-
Protein-protein interactions
- MCODE:
-
Molecular Complex Detection
- BINGO:
-
Biological Networks Gene Ontology tool
- BRMS1L:
-
Breast cancer metastasis suppressor 1-like proteins
- BRMS1:
-
Breast cancer metastasis suppressor 1
- CCNA1:
-
Cyclin A1
- MCM4:
-
Minichromosome maintenance complex component 4
- MCM6:
-
Minichromosome maintenance complex component 6
References
Parney IF (2012) Basic concepts in glioma immunology. Glioma. Springer New York,42–52
Knizhnik AV, Roos WP, Nikolova T et al (2013) Survival and death strategies in glioma cells: autophagy, senescence and apoptosis triggered by a single type of temozolomide-induced DNA damage. PLoS One 8:e55665
Wang Y, Jiang T (2013) Understanding high grade glioma: molecular mechanism, therapy and comprehensive management. Cancer Lett 331: 139–146.
Heim S, Mitelman F (2011) Cancer cytogenetics: chromosomal and molecular genetic abberations of tumor cells. John Wiley & Sons
Furnari FB, Fenton T, Bachoo RM et al (2007) Malignant astrocytic glioma: genetics, biology, and paths to treatment. Genes Dev 21:2683–2710
Kapoor GS, O’rourke DM (2003) Mitogenic signaling cascades in glial tumors. Neurosurgery 52:1425–1435
Milinkovic V, Bankovic J, Rakic M et al (2012) Genomic instability and p53 alterations in patients with malignant glioma. Exp Mol Pathol 93:200–206
Bansal K, Liang ML, Rutka JT (2006) Molecular biology of human gliomas. Technol Cancer Res Treat 5:185–194
Sun L, Hui A-M, Su Q et al (2006) Neuronal and glioma-derived stem cell factor induces angiogenesis within the brain. Cancer Cell 9:287–300
Barrett T, Troup DB, Wilhite SE et al (2007) NCBI GEO: mining tens of millions of expression profiles–database and tools update. Nucleic Acids Res 35:D760–765
Barrett T, Troup DB, Wilhite SE et al (2011) NCBI GEO: archive for functional genomics data sets—10 years on. Nucleic Acids Res 39:D1005–D1010
Mccall MN, Bolstad BM, Irizarry RA (2010) Frozen robust multiarray analysis (fRMA). Biostatistics 11:242–253
Gautier L, Irizarry R, Cope L, and Bolstad B (2012) Description of Affy
Falcon S, Gentleman R (2007) Using GOstats to test gene lists for GO term association. Bioinformatics 23:257–258
Abbas A, Kong X-B, Liu Z, Jing B-Y, Gao X (2013) Automatic peak selection by a Benjamini-Hochberg-based algorithm. PLoS One 8:e53112
Becker KG, Barnes KC, Bright TJ, Wang SA (2004) The genetic association database. Nat Genet 36:431–432
Peri S, Navarro JD, Amanchy R et al (2003) Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res 13:2363–2371
Kohl M, Wiese S, Warscheid B (2011) Cytoscape: software for visualization and analysis of biological networks, in data mining in proteomics. Humana Press. 291–303
Assenov Y, Ramirez F, Schelhorn SE, Lengauer T, Albrecht M (2008) Computing topological parameters of biological networks. Bioinformatics 24:282–284
Zhou T, Yan G, Wang B-H (2005) Maximal planar networks with large clustering coefficient and power-law degree distribution. Phys Rev E 71:046141
Bader GD, Hogue CW (2003) An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinforma 4:2
Mi G, Di Y, Emerson S, Cumbie JS, Chang JH (2012) Length bias correction in gene ontology enrichment analysis using logistic regression. PLoS One 7:e46128
Maere S, Heymans K, Kuiper M (2005) BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21:3448–3449
Li X, Li C, Shang D et al (2011) The implications of relationships between human diseases and metabolic subpathways. PLoS One 6:e21131
Chen H, Boutros P (2011) VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinforma 12:35
Hurst DR, Mehta A, Moore BP et al (2006) Breast cancer metastasis suppressor 1 (BRMS1) is stabilized by the Hsp90 chaperone. Biochem Biophys Res Commun 348:1429–1435
Mei P, Bai J, Shi M et al (2014) BRMS1 suppresses glioma progression by regulating invasion, migration and adhesion of glioma cells. PLoS One 9:e98544
Hurst DR, Welch DR (2011) Unraveling the enigmatic complexities of BRMS1-mediated metastasis suppression. FEBS Lett 585:3185–3190
Nikolaev AY, Papanikolaou NA, Li M, Qin J, Gu W (2004) Identification of a novel BRMS1-homologue protein p40 as a component of the mSin3A/p33<sup>ING1b</sup>/HDAC1 deacetylase complex. Biochem Biophys Res Commun 323:1216–1222
Liu D, Liao C, Wolgemuth DJ (2000) A role for cyclin A1 in the activation of MPF and G2–M transition during meiosis of male germ cells in mice. Dev Biol 224:388–400
Ji T, Liu D, Shao W, Yang W, Wu H, Bian X (2012) Decreased expression of LATS1 is correlated with the progression and prognosis of glioma. J Exp Clin Cancer Res 31:67
Watanabe T, Yokoo H, Yokoo M, Yonekawa Y, Kleihues P, Ohgaki H (2001) Concurrent inactivation of RB1 and TP53 pathways in anaplastic oligodendrogliomas. J Neuropathol Exp Neurol 60:1181–1189
Kanter DM, Bruck I, Kaplan DL (2008) Mcm subunits can assemble into two different active unwinding complexes. J Biol Chem 283:31172–31182
Van Den Boom J, Wolter M, Kuick R et al (2003) Characterization of gene expression profiles associated with glioma progression using oligonucleotide-based microarray analysis and real-time reverse transcription-polymerase chain reaction. Am J Pathol 163:1033–1043
Dickerson JE, Zhu A, Robertson DL, Hentges KE (2011) Defining the role of essential genes in human disease. PLoS One 6:e27368
Tunbridge EM, Eastwood SL, Harrison PJ (2011) Changed relative to what? Housekeeping genes and normalization strategies in human brain gene expression studies. Biol Psychiatry 69:173–179
Xiao S-J, Zhang C, Zou Q, Ji Z-L (2010) TiSGeD: a database for tissue-specific genes. Bioinformatics 26:1273–1275
Gong X-J, Yu H, Yang C-B, Li Y-F (2012) Knowledge enrichment analysis for human tissue-specific genes uncover new biological insights. J Integr Bioinform 9:194
Conflict of interest
The authors have declared that no competing interests exist.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pan, W., Li, G., Yang, X. et al. Revealing the Potential Pathogenesis of Glioma by Utilizing a Glioma Associated Protein-Protein Interaction Network. Pathol. Oncol. Res. 21, 455–462 (2015). https://doi.org/10.1007/s12253-014-9848-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-014-9848-9