Abstract
The Notch pathway is a core cell–cell signaling system in Metazoa which plays a key role in development and adult homeostasis. Whereas most Notch structural biology research has focused on the negative regulatory region and the intracellular domain, relatively little structural information is available for the extracellular part of human Notch-1 (hN-1) which mediates ligand recognition. This region consists of 36 epidermal growth factor-like (EGF) domains, many of which contain a calcium-binding consensus sequence. The calcium-binding site in each case is located within the N-terminal portion of the domain, and is associated with both intra- and inter-domain rigidity. The absence of calcium-binding sites in EGF6, EGF10 and EGF22, however, suggests that these domains might represent regions of flexibility in the receptor which could influence the cell-surface architecture (usually depicted as an extended rod projecting from the cell surface). To probe this, we have purified a quadruple-domain construct from hN-1, in which the non-calcium-binding EGF6 is flanked by EGF4–5 and EGF7. Here, we report 1H, 13C and 15N resonance assignments for this four-domain 157 amino acid construct. The assignments presented here are the prerequisite for a detailed study of the structure and dynamics of this region of the Notch receptor.
Similar content being viewed by others
Biological context
The Notch signaling pathway is highly conserved in the animal kingdom and is responsible for cell–cell communication. The Notch signal is activated when ligands on a signal-sending cell bind to the Notch receptor on a signal-receiving cell, resulting in regulated proteolysis and release of the intracellular domain of Notch (NICD). This protein fragment translocates to the nucleus and following interaction with the transcription factor CSL and a coactivator Mastermind regulates transcription of a subset of genes. Canonical signaling via ligand-mediated activation of the Notch receptor influences many aspects of cell growth, differentiation and death which are vital for embryonic development and maintenance of the adult organism. Dysregulation of the pathway results in both inherited and acquired diseases including cancer. The outcome of the Notch signal is often context specific (Bray 2006; Guruharsha et al. 2012; Kopan and Ilagan 2009). For example, Notch signaling can be oncogenic, as in hematopoietic malignancies, but can also have tumour suppressor functions, such as in solid tumours (Koch and Radtke 2007). In addition, Notch is most likely a cancer stem cell factor (Pannuti et al. 2010).
The downstream consequences of Notch activation have been studied widely, and high resolution structures of the NRR and NICD region (Fig. 1) have facilitated the understanding of receptor activation and signaling (for review see Chillakuri et al. 2012). However, the molecular mechanisms behind receptor/ligand recognition and regulation by O-glycosylation are not yet understood. The membrane distal region of Notch (Fig. 1) consists of 36 EGF-like domains organized as multiple tandem repeats. A critical role for EGF11 and EGF12 in ligand binding was identified by deletion studies which showed these domains were sufficient to confer ligand binding in cell aggregation assays (Rebay et al. 1991). More recently Drosophila Notch EGF8 has been shown to contain an additional binding site for the ligand Serrate (Yamamoto et al. 2012). The high resolution X-ray structure of EGF11–13 showed rigid interdomain contacts (with twist and tilt angles of 119° ± 5° and 17° ± 4° respectively (Cordle et al. 2008), in good agreement with an earlier NMR structure (Hambleton et al. 2004) and a near linear organisation. Other multiple tandem repeats of Notch EGF domains which bind calcium are likely to have similar extended structures since residues involved in interface stabilisation are highly conserved. Subsequent site-directed mutagenesis has mapped the ligand-binding site to the central β-hairpin of hN-1 EGF12 (Whiteman et al. 2013), which is post-translationally modified by O-glycosylation (Taylor et al. 2014).
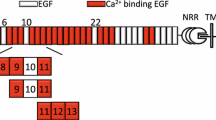
Modular organization of the extracellular domain of the human Notch-1 receptor. The negative regulatory region and the transmembrane domain (TM) are indicated. Individual domains belonging to the Notch intracellular domain are not indicated separately. Calcium-binding EGF domains are indicated in red. Schematic pictures of the constructs EGF4–7, EGF4–6, EGF5–7 and EGF4–5 used in the NMR study are shown. Each EGF domain contains six highly-conserved cysteine residues paired in a 1–3, 2–4, 5–6 arrangement to stabilize domain structure. There are six residues between the 6th cysteine of an EGF domain and the 1st cysteine of the following domain in EGF4–7. Since EGF5 and EGF7 have a calcium-binding consensus sequence, the linker length for EGF4/EGF5 and EGF6/EGF7 comprises two residues; the other four residues are constrained by calcium binding. In the case of EGF5/EGF6 the conformation of the six amino acids is unknown and will be the subject of a future structural study
Sequence inspection demonstrates that a number of EGF-like domains of hNotch-1 do not bind Ca2+; this includes EGF6, EGF10 and EGF22. This could result in very different interfaces which would alter the overall architecture of the receptor, such as is seen in other EGF-containing proteins (Brandstetter et al. 1995; Kettle et al. 1999; Lee et al. 2004). This information is crucial to understand the molecular mechanisms underlying Notch activation and regulation. We have therefore begun a study of these three regions by NMR to understand their structure and dynamics in solution. In this report we present 1H, 13C and 15N assignments for the quadruple EGF-like domain construct EGF4–7 expressed in E. coli.
Methods and experiments
Protein expression, purification and refolding
Protein expression and purification was carried out as described (Knott et al. 1996; Whiteman et al. 2014). The protein fragment, consisting of EGF4 to EGF7 of human Notch-1, was expressed in E. coli BL21 cells transformed with a pQE30 (Qiagen) based protein expression construct and a pREP4 plasmid for control of expression via the Lac Repressor. The expression vector contained an N-terminal His6 tag for purification, a factor Xa protease recognition site (IEGR) for later removal of the His6 tag, an SA spacer, followed by residue Q140 of EGF4; the last residue in the construct was E294. For the constructs EGF4–6, EGF5–7 and EGF4–5, the first and last residues were Q140/E256, D178/E294 and Q140/R217, respectively; all constructs have an SA spacer at their N-terminus following factor Xa cleavage of the His6 tag.
15N or 15N/13C-double-labeled protein was produced by growing cells in M9 medium containing 0.1 % (w/v) 15NH4Cl and 0.5 % (w/v) unlabeled glucose, or 0.1 % (w/v) 15NH4Cl and 0.5 % (w/v) 13C-glucose (Cambridge Isotope Laboratories), 100 μg/ml ampicillin and 25 μg/ml kanamycin, as described previously (Muranyi et al. 2004). 100 ml of starter culture, grown in labelled medium at 37 °C for ~18 h, was added to 0.5 l of medium. Cells were grown to OD600 ~0.8 and expression was induced with isopropyl-β-D-thiogalactopyranoside (IPTG) at a final concentration of 100 μM. Cells were incubated at 18 °C for at least 6 h. Cells were spun down at 5,000 rpm for 10 min and lysed with 6 M guanidine-HCl, 50 mM sodium phosphate, 5 mM β-mercaptoethanol for 1 h at room temperature (RT). DNA was sheared by sonication for 1 min at 20 Watt (Jencons Ultrasonic Processor) and the cell lysate was spun down at 40,000 rpm for 45 min (Beckman L7–55). The supernatant was loaded onto a Ni2+ chelating Sepharose column (GE Healthcare) and eluted with buffer containing 50 mM EDTA and 100 mM Tris pH 8.3. The protein was reduced for 1 h at RT by addition of dithiothreitol (DTT) to a final concentration of 0.1 M. The pH was then lowered to pH ~ 2, by addition of concentrated HCl, and the solution was dialysed against 0.1 % (v/v) trifluoroacetic acid (TFA) overnight. The solution was filtered through 0.2 μM filters (Millex-GP) and the soluble fraction purified by reverse-phase HPLC using a Beckman Gold system with reverse phase C8 column. Purified, reduced protein was refolded under the following conditions: 100 mM Tris–HCl pH 8.3, 3 mM l-cysteine, 0.3 mM l-cystine, 0.2 mg/ml protein at 37 °C for ~48 h. The refolding mixture was acidified to pH ~ 2 and dialysed against 0.1 % (v/v) TFA for at least 5 h. Protein was then concentrated by ultrafiltration and purified by HPLC. After lyophilisation, the His6 tag was removed by incubation overnight with one unit of bovine factor Xa (DENZYME) per mg of protein in 50 mM Tris–HCl pH 7.5, 0.1 M NaCl, 1 mM CaCl2 at 37 °C. Protein was further purified by cation-exchange fast liquid protein chromatography and HPLC. The final product was analysed by SDS-PAGE in the presence and absence of 5 % (v/v) 2-mercaptoethanol. Electrospray ionisation mass spectrometry was performed to check the correct mass of the constructs (16603 Dalton for the unlabeled EGF4–7 construct).
NMR spectroscopy
Ca2+ binding in EGF4–7 was followed by 1D NMR for a sample in 100 % D2O, 5 mM Tris, 150 mM NaCl, pH 7.5 and a protein concentration of ~0.5 mM. Protein concentrations for resonance assignment experiments were 1 mM for 15N-single-labelled, and 0.5 mM for 15N/13C-double-labelled material, in 95 % H2O/5 % D2O (v/v) at pH 6.1 and a Ca2+ concentration of 30 mM. All NMR experiments were carried out at 298 K on home-built spectrometers with triple-resonance probes and GE/Omega data acquisition systems operating at 1H-operating frequencies of 500, 600, 750 and 950 MHz. In addition, a Bruker Avance 500 MHz spectrometer with a Cryoplatform, equipped with a TCI CryoProbe, was used.
Assignment of the spectrum of EGF4–7 was carried out using a dissection approach; NMR spectra were collected for the full-length EGF4–7 protein but also for a series of shorter constructs EGF4–6, EGF5–7 and EGF4–5 (Fig. 1). Initially, sequential backbone assignments were obtained with data from 3D 15N-edited TOCSY-HSQC and NOESY-HSQC experiments (datasets collected for EGF4–7, EGF4–5 and EGF4–6). Sequential 1H and 15N assignments were confirmed and extended to 13C with 3D HNCA (for EGF4–7, EGF4–6 and EGF5–7), HNCO (for EGF4–7 and EGF5–7), HN(CA)CO (for EGF5–7), CBCANH (for EGF4–7 and EGF5–7), CBCA(CO)NH (for EGF4–7, EGF4–6 and EGF5–7) and HBHA(CO)NH (for EGF4–6 and EGF5–7) experiments with 15N/13C-double labeled material. Side-chain assignments were obtained using these experiments, supplemented with 3D HCCH-TOCSY (for EGF4–6 and EGF5–7), 3D 13C-edited NOESY-HSQC (for EGF4–6 and EGF5–7) and 2D 1H-1H NOESY (for all constructs). NMRPipe was used for NMR data processing (Delaglio et al. 1995) and CcpNmr Analysis for subsequent analysis (Vranken et al. 2005).
Assignments and data deposition
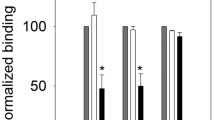
The correct fold of the EGF4–7 construct was confirmed using Ca2+ titrations monitored by 1D NMR (Fig. 2). The chemical shifts of resonances from the consensus aromatic residue located between the 3rd and 4th cysteines of a calcium-binding EGF domain and from the aromatic ‘packing’ residue located between the 5th and 6th cysteine of the preceding domain are known to be sensitive to calcium binding (Knott et al. 1996). Changes in the aromatic region of the spectrum around 6.7–6.8, 7.1 and 7.3 ppm are observed upon addition of 250–500 μM Ca2+ (Fig. 2); peaks in these regions correspond to Y279, the consensus aromatic in EGF7, and F249, the packing aromatic in EGF6. At higher concentrations of added Ca2+, peak shifts are observed around 7.7–7.8 ppm; peaks in this region correspond to H196, which is located near to the calcium-binding site in EGF5, and H171, which is next to the packing aromatic (F170) in EGF4. These data demonstrate calcium binding in EGF5 and EGF7 and formation of interfaces of these domains with EGF4 and EGF6, respectively.
Ca2+ titration of EGF4–7 in 100 % D2O, 5 mM Tris, 150 mM NaCl, pH 7.5. The concentration of added CaCl2 is indicated on the left side of the 1D spectra. Black arrows highlight the chemical shift changes arising from Y279 and F249, which are indicative of calcium binding in EGF7 and the formation of a packing interface with EGF6. At higher concentrations of added calcium, chemical shift changes in the histidine region of the spectrum arise from H196 and H171 are indicative of calcium binding in EGF5 and the formation of a packing interface with EGF4
The spectrum of EGF4–7 was assigned using a dissection approach with a series of shorter constructs EGF4–6, EGF5–7 and EGF4–5 (Fig. 1). In general, peaks in the HSQC spectrum of EGF4–7 could be assigned to a specific EGF domain by comparison with spectra of the smaller constructs. Some construct-dependent chemical shift differences were observed in the proximity (~2–3 residues) of the N- and C-termini and in calcium-binding sites. For example, differences are observed for V199 and G200, located near to the calcium-binding site in EGF5, in the EGF4–7 and EGF5–7 constructs; in the former these residues are involved in a packing interaction with F170 located in EGF4 while in the later construct this interaction is missing due to the absence of EGF4.
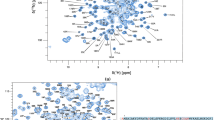
Resonances from residues Q140 to E294 were assigned for EGF4–7; Fig. 3 shows a 1H-15N HSQC spectrum of EGF4–7. All non-proline backbone 1HN and 15N resonances were assigned except N180, E181 and C182 at the beginning of EGF5; these residues also gave broad resonances in the shorter EGF4–5 construct that otherwise showed a very high signal-to-noise ratio. Further assignments were 93.1 % of Hα, 93.5 % of Hβ, 55.9 % of Hγ, 58.4 % of Hδ and 39.0 % of Hε, 91.6 % of 13C’, 95.5 % of 13Cα, 94.9 % of 13Cβ, 50.0 % of 13Cγ, 31.8 % of 13Cδ. Most missing assignments corresponded to residues at the C-terminus of EGF5 and at the N-terminus of EGF6, where residues showed line broadening. The chemical shift assignments for EGF4–7 have been deposited in the BioMagResBank (http://www.bmrb.wisc.edu) under the accession number 25172.
References
Brandstetter H, Bauer M, Huber R, Lollar P, Bode W (1995) X-ray structure of clotting factor IXa: active site and module structure related to xase activity and hemophilia b. Proc Natl Acad Sci USA 92(21):9796–9800
Bray SJ (2006) Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol 7(9):678–689
Chillakuri CR, Sheppard D, Lea SM, Handford PA (2012) Notch receptor-ligand binding and activation: insights from molecular studies. Semin Cell Dev Biol 23(4):421–428
Cordle J, Johnson S, Tay JZ, Roversi P, Wilkin MB, de Madrid BH, Shimizu H, Jensen S, Whiteman P, Jin B, Redfield C, Baron M, Lea SM, Handford PA (2008) A conserved face of the Jagged/Serrate DSL domain is involved in notch trans-activation and cis-inhibition. Nat Struct Mol Biol 15(8):849–857
Delaglio F, Grzesiek S, Vuister G, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6(3):277–293
Guruharsha KG, Kankel MW, Artavanis-Tsakonas S (2012) The notch signalling system: recent insights into the complexity of a conserved pathway. Nat Rev Genet 13(9):654–666
Hambleton S, Valeyev NV, Muranyi A, Knott AV, Werner JM, McMichael AJ, Handford PA, Downing AK (2004) Structural and functional properties of the human notch-1 ligand binding region. Structure 12(12):2173–2183
Kettle S, Yuan X, Grundy G, Knott V, Downing AK, Handford PA (1999) Defective calcium binding to fibrillin-1: consequence of an N2144S change for fibrillin-1 structure and function. J Mol Biol 285(3):1277–1287
Knott V, Downing AK, Cardy CM, Handford PA (1996) Calcium binding properties of an epidermal growth factor-like domain pair from human fibrillin-1. J Mol Biol 255(1):22–27
Koch U, Radtke F (2007) Notch and cancer: a double-edged sword. Cell Mol Life Sci 64(21):2746–2762
Kopan R, Ilagan MX (2009) The canonical notch signaling pathway: unfolding the activation mechanism. Cell 137(2):216–233
Lee SSJ, Knott V, Jovanovic J, Harlos K, Grimes JM, Choulier L, Mardon HJ, Stuart DI, Handford PA (2004) Structure of the integrin binding fragment from fibrillin-1 gives new insights into microfibril organization. Structure 12(4):717–729
Muranyi A, Hambleton S, Knott V, McMichael A, Handford PA, Downing AK (2004) 1H, 13C, and 15N resonance assignments of human Notch-1 calcium binding EGF domains 11-13. J Biomol NMR 29(3):443–444
Pannuti A, Foreman K, Rizzo P, Osipo C, Golde T, Osborne B, Miele L (2010) Targeting Notch to target cancer stem cells. Clin Cancer Res 16(12):3141–3152
Rebay I, Fleming RJ, Fehon RG, Cherbas L, Cherbas P, Artavanis-Tsakonas S (1991) Specific EGF repeats of Notch mediate interactions with Delta and Serrate: implications for Notch as a multifunctional receptor. Cell 67(4):687–699
Taylor P, Takeuchi H, Sheppard D, Chillakuri C, Lea SM, Haltiwanger RS, Handford PA (2014) Fringe-mediated extension of O-linked fucose in the ligand-binding region of Notch1 increases binding to mammalian Notch ligands. Proc Natl Acad Sci USA 111(20):7290–7295
Vranken W, Boucher W, Stevens T, Fogh R, Pajon A, Llinas M, Ulrich E, Markley J, Ionides J, Laue E (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59(4):687–696
Whiteman P, de Madrid BH, Taylor P, Li D, Heslop R, Viticheep N, Tan JZ, Shimizu H, Callaghan J, Masiero M, Li JL, Banham AH, Harris AL, Lea SM, Redfield C, Baron M, Handford PA (2013) Molecular basis for Jagged-1/Serrate ligand recognition by the Notch receptor. J Biol Chem 288(10):7305–7312
Whiteman P, Redfield C, Handford PA (2014) Bacterial expression and in vitro refolding of limited fragments of the Notch receptor and its ligands. Methods Mol Biol 1187:193–208
Yamamoto S, Charng WL, Rana NA, Kakuda S, Jaiswal M, Bayat V, Xiong B, Zhang K, Sandoval H, David G, Wang H, Haltiwanger RS, Bellen HJ (2012) A mutation in EGF repeat-8 of Notch discriminates between Serrate/Jagged and Delta family ligands. Science 338(6111):1229–1232
Acknowledgments
P.A.H. and C.R. acknowledge a Cancer Research UK-funded PhD studentship which supported P.C.W.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Weisshuhn, P.C., Handford, P.A. & Redfield, C. 1H, 13C and 15N assignments of EGF domains 4 to 7 of human Notch-1. Biomol NMR Assign 9, 275–279 (2015). https://doi.org/10.1007/s12104-014-9591-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-014-9591-x