Abstract
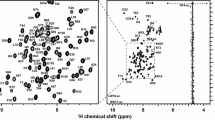
Homeodomain-containing transcription factors including Hox proteins play fundamental roles in the regulation of different cellular and developmental processes. These proteins all contain a homeodomain, which is a 60-amino acid DNA binding domain encoded by a 180-base pair DNA sequence. Homeodomains also serve as protein interaction targets to regulate the functions of these transcription factors or other proteins. The cell cycle regulator Geminin interacts with Hox homeodomains to inhibit the transcription activities of Hox proteins and enrolls Hox proteins in the cell proliferation process. Here we report complete chemical shift assignments of the homeodomain of Hoxc9 (Hoxc9-HD) in complex with the homeodomain binding region of Geminin (Gem-HBR), which were determined by triple resonance NMR experiments. These resonance assignments provide a basis for the structure determination of the Hoxc9-HD/Gem-HBR complex and for the further study of homeodomains in complex with other regulatory protein partners (BMRB deposits with accession number 17407).


Similar content being viewed by others
References
Bell SP, Dutta A (2002) DNA replication in eukaryotic cells. Annu Rev Biochem 71:333–374
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRpipe—a multidimensional spectral processing system based on Unix pipes. J Biomol NMR 6(3):277–293
Gehring WJ, Affolter M, Burglin T (1994) Homeodomain proteins. Annu Rev Biochem 63:487–526
Goddard TD, Kneller DG (2002) SPARKY 3. University of California, San Francisco
Luo L, Yang X, Takihara Y, Knoetgen H, Kessel M (2004) The cell-cycle regulator Geminin inhibits Hox function through direct and polycomb-mediated interactions. Nature 427(6976):749–753
Ohno Y, Yasunaga S, Ohtsubo M, Mori S, Tsumura M, Okada S, Ohta T, Ohtani K, Kobayashi M, Takihara Y (2010) Hoxb4 transduction down-regulates Geminin protein, providing hematopoietic stem and progenitor cells with proliferation potential. Proc Natl Acad Sci USA 107(50):21529–21534
Rezsohazy R (2014) Non-transcriptional interactions of Hox proteins: inventory, facts, and future directions. Dev Dyn 243(1):117–131
Sattler M, Schleucher J, Griesinger C (1999) Heteronuclear multidimensional NMR experiments for the structure determination of proteins in solution employing pulsed field gradients. Prog Nucl Magn Reson Spectrosc 34(2):93–158
Wishart DS, Sykes BD (1994) The C-13 Chemical-Shift Index—a simple method for the identification of protein secondary structure using C-13 chemical-shift data. J Biomol NMR 4(2):171–180
Zhou B, Liu C, Xu Z, Zhu G (2012) Structural basis for homeodomain recognition by the cell-cycle regulator Geminin. Proc Natl Acad Sci USA 109(23):8931–8936
Zhu G, Bax A (1992) Improved linear prediction of truncated damped sinusoids using modified backward–forward linear prediction. J Magn Reson 100:202–207
Acknowledgments
We thank Mr. R. Feng for setting up the NMR experiments; C. Liang for the Geminin cDNA. This work was supported by grants from Hong Kong Research Grants Council (RGC663911) and AoE/M-06/08.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhou, B., Liu, C. & Zhu, G. 1H, 15N and 13C chemical shift assignments of the homeodomain of Hoxc9 in complex with the cell cycle regulator Geminin. Biomol NMR Assign 9, 165–168 (2015). https://doi.org/10.1007/s12104-014-9566-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-014-9566-y