Abstract
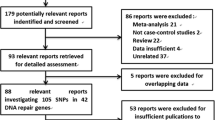
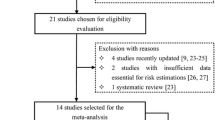
Genetic variants found in DNA repair genes (ERCC1, rs3212986; ERCC2, rs13181; ERCC4, rs1800067; ERCC5, rs17655; XRCC1, rs1799782, rs25487, rs25489; XRCC3, rs861539) have been reported to have an ambivalent association with the development of glioma. In the present study, a meta-analysis was conducted to confirm the relationship, taking heterogeneity of population into consideration. We analyzed 21 articles of 6 genes along with 8 single nucleotide polymorphisms (SNPs) (24,078 cases and 30,926 healthy individuals), which assessed the relationship between nucleotide excision, base excision, double-strand break repair gene, and the development of glioma under five models. All statistical analysis was implemented by the software of R 3.2.1, and the relationships between key polymorphic loci in DNA repair genes and glioma were quantified by the pooled odds ratio (OR) and 95 % confidential intervals. Overall, the synthesized evidence demonstrated that the SNP of rs13181 and rs1799782 significantly increased the risk of glioma whereas SNP of rs1800067 were significantly associated with a decrease in the risk of glioma. Additionally, subgroup analyses of 8 SNPs by ethnicity indicated that the mutation of rs13181, rs1800067 were apparently protective factors of glioma among Asians, while the mutation of rs13181 was a risk factors of glioma in Caucasians. Furthermore, the mutation of rs1799782 significantly raises the risk of glioma for Asian. Our study suggested that rs13181*C and rs1799782*A are risk alleles for glioma; rs1800067*A are beneficial alleles for decreased susceptibility to glioma. Future studies with large sample size and other races are strongly recommended to confirm the results from this study.





Similar content being viewed by others
Abbreviations
- SNP:
-
Single nucleotide polymorphism
- BER:
-
Base excision repair
- NER:
-
Nucleotide excision repair
- MMR:
-
Mismatch repair
- DSBR:
-
Double-strand break repair
- XRCC :
-
X-ray cross-complementing group
- ERCC1 :
-
Excision repair cross-complementing group 1
- NOS:
-
Newcastle-Ottawa Scale
References
Ricard D, Idbaih A, Ducray F, Lahutte M, Hoang-Xuan K, Delattre JY (2012) Primary brain tumours in adults. Lancet 379(9830):1984–96. doi:10.1016/S0140-6736(11)61346-9
Rousseau A, Mokhtari K, Duyckaerts C (2008) The 2007 WHO classification of tumors of the central nervous system—what has changed? Curr Opin Neurol 21(6):720–7. doi:10.1097/WCO.0b013e328312c3a7
Chen J, McKay RM, Parada LF (2012) Malignant glioma: lessons from genomics, mouse models, and stem cells. Cell 149(1):36–47. doi:10.1016/j.cell.2012.03.009
Ohgaki H, Kleihues P (2005) Epidemiology and etiology of gliomas. Acta Neuropathol 109(1):93–108. doi:10.1007/s00401-005-0991-y
Wen PY, Kesari S (2008) Malignant gliomas in adults. N Engl J Med 359(5):492–507. doi:10.1056/NEJMra0708126
Schwartzbaum JA, Fisher JL, Aldape KD, Wrensch M (2006) Epidemiology and molecular pathology of glioma. Nat clin pract Neurol 2(9):494–503. doi:10.1038/ncpneuro0289, quiz 491 p following 516
Prasad G, Haas-Kogan DA (2009) Radiation-induced gliomas. Expert Rev Neurother 9(10):1511–7. doi:10.1002/pmic.200800802
Bondy M, Wiencke J, Wrensch M, Kyritsis AP (1994) Genetics of primary brain tumors: a review. J Neurooncol 18(1):69–81
Little MP, de Vathaire F, Shamsaldin A, Oberlin O, Campbell S, Grimaud E, Chavaudra J, Haylock RG et al (1998) Risks of brain tumour following treatment for cancer in childhood: modification by genetic factors, radiotherapy and chemotherapy. International journal of cancer Journal international du cancer 78(3):269–75. doi:10.1002/(SICI)1097-0215(19981029)78:3<269::AID-IJC1>3.0.CO;2-T
Bondy ML, Wang LE, El-Zein R, de Andrade M, Selvan MS, Bruner JM, Levin VA, Alfred Yung WK et al (2001) Gamma-radiation sensitivity and risk of glioma. J Natl Cancer Inst 93(20):1553–7
Friedberg EC (2003) DNA damage and repair. Nature 421(6921):436–40. doi:10.1038/nature01408
Rajaraman P, Hutchinson A, Wichner S, Black PM, Fine HA, Loeffler JS, Selker RG, Shapiro WR et al (2010) DNA repair gene polymorphisms and risk of adult meningioma, glioma, and acoustic neuroma. Neuro Oncol 12(1):37–48. doi:10.1093/neuonc/nop012
Deeks JJ, Dinnes J, D’Amico R, Sowden AJ, Sakarovitch C, Song F, Petticrew M, Altman DG (2003) Evaluating non-randomised intervention studies. Health Technol Assess 7(27):1–173, iii-x
Higgins JP, Thompson SG, Deeks JJ, Altman DG (2003) Measuring inconsistency in meta-analyses. BMJ 327(7414):557–60. doi:10.1136/bmj.327.7414.557
Higgins JP, Thompson SG (2002) Quantifying heterogeneity in a meta-analysis. Stat Med 21(11):1539–58. doi:10.1002/sim.1186
Wang LE, Bondy ML, Shen H, El-Zein R, Aldape K, Cao Y, Pudavalli V, Levin VA et al (2004) Polymorphisms of DNA repair genes and risk of glioma. Cancer Res 64(16):5560–3. doi:10.1158/0008-5472.can-03-2181
Wrensch M, Kelsey KT, Liu M, Miike R, Moghadassi M, Sison JD, Aldape K, McMillan A et al (2005) ERCC1 and ERCC2 polymorphisms and adult glioma. Neuro Oncol 7(4):495–507. doi:10.1215/S1152851705000037
Felini MJ, Olshan AF, Schroeder JC, North KE, Carozza SE, Kelsey KT, Liu M, Rice T et al (2007) DNA repair polymorphisms XRCC1 and MGMT and risk of adult gliomas. Neuroepidemiology 29(1–2):55–8. doi:10.1159/000108919
Cengiz SL, Acar H, Inan Z, Yavuz S, Baysefer A (2008) Deoxy-ribonucleic acid repair genes XRCC1 and XPD polymorphisms and brain tumor risk. Neurosciences (Riyadh) 13(3):227–32
McKean-Cowdin R, Barnholtz-Sloan J, Inskip PD, Ruder AM, Butler M, Rajaraman P, Razavi P, Patoka J et al (2009) Associations between polymorphisms in DNA repair genes and glioblastoma. Cancer Epidemiol Biomark Prev: a Publication of the American Association for Cancer Research, Cosponsored by the American Society of Preventive Oncology 18(4):1118–26. doi:10.1158/1055-9965.EPI-08-1078
Zhou K, Liu Y, Zhang H, Liu H, Fan W, Zhong Y, Xu Z, Jin L et al (2009) XRCC3 haplotypes and risk of gliomas in a Chinese population: a hospital-based case–control study. Int J Cancer 124(12):2948–53. doi:10.1002/ijc.24307, 2009/03/31 edn
Yosunkaya E, Kucukyuruk B, Onaran I, Gurel CB, Uzan M, Kanigur-Sultuybek G (2010) Glioma risk associates with polymorphisms of DNA repair genes, XRCC1 and PARP1. Br J Neurosurg 24(5):561–5. doi:10.3109/02688697.2010.489655
Custodio AC, Almeida LO, Pinto GR, Santos MJ, Almeida JR, Clara CA, Rey JA, Casartelli C (2011) Analysis of the polymorphisms XRCC1Arg194Trp and XRCC1Arg399Gln in gliomas. Genet Mol Res: GMR 10(2):1120–9. doi:10.4238/vol10-2gmr1125
Hu XB, Feng Z, Fan YC, Xiong ZY, Huang QW (2011) Polymorphisms in DNA repair gene XRCC1 and increased genetic susceptibility to glioma. Asian Pac J Cancer Prev 12(11):2981–4
Zhou L-Q (2011) Polymorphisms of DNA repair gene XRCC1 and risk of glioma: a case–control study in Southern China. Asian Pac J Cancer Prev 12(10):2547–50
Chen D-Q, Yao D-X, Zhao H-Y, Yang S-J (2012) DNA repair gene ERCC1 and XPD polymorphisms predict glioma susceptibility and prognosis. Asian Pac J Cancer Prev 13(6):2791–4. doi:10.7314/apjcp.2012.13.6.2791
Custodio AC, Almeida LO, Pinto GR, Santos MJ, Almeida JR, Clara CA, Rey JA, Casartelli C (2012) Variation in DNA repair gene XRCC3 affects susceptibility to astrocytomas and glioblastomas. Genet mol res : GMR 11(1):332–9. doi:10.4238/2012.February.10.4
Liu H-B, Peng Y-P, Dou C-W, Su X-L, Gao N-K, Tian F-M, Bai J (2012) Comprehensive study on associations between nine SNPs and glioma risk. Asian Pac J Cancer Prev 13(10):4905–8. doi:10.7314/apjcp.2012.13.10.4905
Wang D, Hu Y, Gong H, Li J, Ren Y, Li G, Liu A (2012) Genetic polymorphisms in the DNA repair gene XRCC1 and susceptibility to glioma in a Han population in northeastern China: a case–control study. Gene 509(2):223–7. doi:10.1016/j.gene.2012.08.023
Pan WR, Li G, Guan JH (2013) Polymorphisms in DNA repair genes and susceptibility to glioma in a Chinese population. Int J Mol Sci 14(2):3314–24. doi:10.3390/ijms14023314
Wang XF, Liu S, Shao ZK (2013) Effects of polymorphisms in nucleotide excision repair genes on glioma risk in a Chinese population. Gene 529(2):317–20. doi:10.1016/j.gene.2013.07.025
Zhao P, Zou P, Zhao L, Yan W, Kang C, Jiang T, You Y (2013) Genetic polymorphisms of DNA double-strand break repair pathway genes and glioma susceptibility. BMC Cancer 13:234. doi:10.1186/1471-2407-13-234
Gao K, Mu SQ, Wu ZX (2014) Investigation of the effects of single-nucleotide polymorphisms in DNA repair genes on the risk of glioma. Genet Mol Res: GMR 13(1):1203–11. doi:10.4238/2014.February.27.5
Li J, Qu Q, Qu J, Luo WM, Wang SY, He YZ, Luo QS, Xu YX et al (2014) Association between XRCC1 polymorphisms and glioma risk among Chinese population. Med Oncol (Northwood, London, England) 31(10):186. doi:10.1007/s12032-014-0186-2
Xu G, Wang M, Xie W, Bai X (2014) DNA repair gene XRCC3 Thr241Met polymorphism and susceptibility to glioma: a case–control study. Oncology letters 8(2):864–8. doi:10.3892/ol.2014.2192
Rajaraman P, Melin BS, Wang Z, McKean-Cowdin R, Michaud DS, Wang SS, Bondy M, Houlston R et al (2012) Genome-wide association study of glioma and meta-analysis. Hum Genet 131(12):1877–88. doi:10.1007/s00439-012-1212-0
Bondy ML, Scheurer ME, Malmer B, Barnholtz-Sloan JS, Davis FG, Il’yasova D, Kruchko C, McCarthy BJ et al (2008) Brain tumor epidemiology: consensus from the Brain Tumor Epidemiology Consortium. Cancer 113(7 Suppl):1953–68. doi:10.1002/cncr.23741
Zhang N, Lin LY, Zhu LL, Wu F, Wen H, Pan D, Huang YC, Chen DQ (2012) ERCC1 polymorphisms and risk of adult glioma in a Chinese population: a hospital-based case–control study. Cancer Invest 30(3):199–202. doi:10.3109/07357907.2011.651233
Adel Fahmideh M, Schwartzbaum J, Frumento P, Feychting M (2014) Association between DNA repair gene polymorphisms and risk of glioma: a systematic review and meta-analysis. Neuro Oncol 16(6):807–14. doi:10.1093/neuonc/nou003
Sung P, Bailly V, Weber C, Thompson LH, Prakash L, Prakash S (1993) Human xeroderma pigmentosum group D gene encodes a DNA helicase. Nature 365(6449):852–5. doi:10.1038/365852a0
de Boer J, Hoeijmakers JH (2000) Nucleotide excision repair and human syndromes. Carcinogenesis 21(3):453–60
Chen P, Wiencke J, Aldape K, Kesler-Diaz A, Miike R, Kelsey K, Lee M, Liu J et al (2000) Association of an ERCC1 polymorphism with adult-onset glioma. Cancer Epidemiol Biomarkers Prev: a Publication of the American Association for Cancer Research, Cosponsored by the American Society of Preventive Oncology 9(8):843–7
Caggana M, Kilgallen J, Conroy JM, Wiencke JK, Kelsey KT, Miike R, Chen P, Wrensch MR (2001) Associations between ERCC2 polymorphisms and gliomas. Cancer Epidemiol Biomarkers Prev: a Publication of the American Association for Cancer Research, Cosponsored by the American Society of Preventive Oncology 10(4):355–60
Cui QK, Zhu JX, Liu WD, Wang YH, Wang ZG (2014) Association of ERCC1 rs3212986 & ERCC2 rs13181 polymorphisms with the risk of glioma. Pakistan j med sci 30(6):1409–1414. doi:10.12669/pjms.306.5221
Yuan Q, Liu JW, Xing CZ, Yuan Y (2014) Associations of ERCC4 rs1800067 polymorphism with cancer risk: an updated meta-analysis. Asian Pac J Cancer Prev: APJCP 15(18):7639–44
Geng P, Ou J, Li J, Liao Y, Wang N, Xie G, Sa R, Liu C, Xiang L, Liang H (2015) A Comprehensive analysis of influence ERCC polymorphisms confer on the development of brain tumors. Molecular Neurobiology. doi:10.1007/s12035-015-9371-3
Liu HB, Peng YP, Dou CW, Su XL, Gao NK, Tian FM, Bai J (2012) Comprehensive study on associations between nine SNPs and glioma risk. Asian Pac J Cancer Prev 13(10):4905–8
Hanssen-Bauer A, Solvang-Garten K, Akbari M, Otterlei M (2012) X-ray repair cross complementing protein 1 in base excision repair. Int J Mol Sci 13(12):17210–29. doi:10.3390/ijms131217210
Lee Y, Katyal S, Li Y, El-Khamisy SF, Russell HR, Caldecott KW, McKinnon PJ (2009) The genesis of cerebellar interneurons and the prevention of neural DNA damage require XRCC1. Nat Neurosci 12(8):973–80. doi:10.1038/nn.2375
Zhao B, Ye J, Li B, Ma Q, Su G, Han R (2013) DNA repair gene XRCC3 Thr241Met polymorphism and glioma risk: a meta-analysis. International Journal of Clinical and Experimental Medicine 6(6):438–43
Source of Support
This study was funded by the National Natural Science Foundation of China (no. 81201671 and 81372793) and Foundation of Science and Technology Department of Jilin Province (no. 20140414049GH and 20140101114JC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors declared no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
Forest plot for a ERCC1 rs3212986 and b ERCC5 rs17655 under the allelic model (GIF 29 kb)
Fig. S2
Forest plot for XRCC1 rs25487 under the allelic model (GIF 23 kb)
Fig. S3
Forest plot for a XRCC1 rs25489 and b XRCC3 rs861539 (GIF 37 kb)
Fig. S4
Forest plot for cumulative meta-analysis of a rs3212986, b rs13181, c rs1800067, and d rs17655 under the allelic model (GIF 26 kb)
Fig. S5
Forest plot for cumulative meta-analysis of a rs1799782, b rs25487, c rs25489, and d rs861539 under the allelic model (GIF 47 kb)
Table S1
Sensitivity analysis results of rs1799782, rs25487, rs25489, and rs861539 SNPs (DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Qi, L., Yu, Hq., Zhang, Y. et al. A Comprehensive Meta-analysis of Genetic Associations Between Key Polymorphic Loci in DNA Repair Genes and Glioma Risk. Mol Neurobiol 54, 1314–1325 (2017). https://doi.org/10.1007/s12035-016-9725-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-016-9725-5