Abstract
Hirschsprung’s disease (HSCR) is a complex developmental defect characterized by the absence of enteric ganglia in the gastrointestinal tract. Although the genetic defect of enteric nervous system (ENS) was identified to play a critical role in the progress of HSCR, the systemic genetic dissection of HSCR still needs to be clarified. In this study, we firstly performed exome sequencing of two HSCR patients from a Han Chinese family, including the affected mother and son. After the initial quality filtering (coverage ≥ 5X and SNP quality score ≥ 40) of the raw data, we identified 13,948 and 13,856 single nucleotide variants (SNVs), respectively. We subsequently compared the SNVs against public databases (dbSNP130, HapMap, and 1000 Genome Project) and obtained a total of 15 novel nonsynonymous SNVs in 15 genes, which were shared between these two patients. Follow-up Sanger sequencing and bioinformatics analysis highlighted variant c.853G>A (p.E285K) in NRG3, a gene involved in the development of ENS. In the validation phase, we sequenced all nine exons of NRG3 in 96 additional sporadic HSCR cases and 110 healthy individuals and identified another nonsynonymous variant c.1329G>A (p.M443I) and two synonymous variants c.828G>A (p.T276T) and c.1365T>A (p.P455P) only in the cases. Our results indicated that NRG3 may be a susceptibility gene for HSCR in a Chinese population.



Similar content being viewed by others
References
Iwashita T, Kruger GM, Pardal R, Kiel MJ, Morrison SJ (2003) Hirschsprung disease is linked to defects in neural crest stem cell function. Science 301(5635):972–976
Kenny SE, Tam PK, Garcia-Barcelo M (2010) Hirschsprung’s disease. Semin Pediatr Surg 19(3):194–200
Amiel J, Sproat-Emison E, Garcia-Barcelo M, Lantieri F, Burzynski G, Borrego S, Pelet A, Arnold S, Miao X, Griseri P, Brooks AS, Antinolo G, de Pontual L, Clement-Ziza M, Munnich A, Kashuk C, West K, Wong KK, Lyonnet S, Chakravarti A, Tam PK, Ceccherini I, Hofstra RM, Fernandez R (2008) Hirschsprung disease, associated syndromes and genetics: a review. J Med Genet 45(1):1–14
Garcia-Barcelo MM, Tang CS, Ngan ES, Lui VC, Chen Y, So MT, Leon TY, Miao XP, Shum CK, Liu FQ, Yeung MY, Yuan ZW, Guo WH, Liu L, Sun XB, Huang LM, Tou JF, Song YQ, Chan D, Cheung KM, Wong KK, Cherny SS, Sham PC, Tam PK (2009) Genome-wide association study identifies NRG1 as a susceptibility locus for Hirschsprung’s disease. Proc Natl Acad Sci U S A 106(8):2694–2699
Tam PK, Garcia-Barcelo M (2009) Genetic basis of Hirschsprung’s disease. Pediatr Surg Int 25(7):543–558
Amiel J, Laudier B, Attie-Bitach T, Trang H, de Pontual L, Gener B, Trochet D, Etchevers H, Ray P, Simonneau M, Vekemans M, Munnich A, Gaultier C, Lyonnet S (2003) Polyalanine expansion and frameshift mutations of the paired-like homeobox gene PHOX2B in congenital central hypoventilation syndrome. Nat Genet 33(4):459–461
Wakamatsu N, Yamada Y, Yamada K, Ono T, Nomura N, Taniguchi H, Kitoh H, Mutoh N, Yamanaka T, Mushiake K, Kato K, Sonta S, Nagaya M (2001) Mutations in SIP1, encoding Smad interacting protein-1, cause a form of Hirschsprung disease. Nat Genet 27(4):369–370
Puffenberger EG, Hosoda K, Washington SS, Nakao K, deWit D, Yanagisawa M, Chakravart A (1994) A missense mutation of the endothelin-B receptor gene in multigenic Hirschsprung’s disease. Cell 79(7):1257–1266
Hofstra RM, Osinga J, Tan-Sindhunata G, Wu Y, Kamsteeg EJ, Stulp RP, van Ravenswaaij-Arts C, Majoor-Krakauer D, Angrist M, Chakravarti A, Meijers C, Buys CH (1996) A homozygous mutation in the endothelin-3 gene associated with a combined Waardenburg type 2 and Hirschsprung phenotype (Shah–Waardenburg syndrome). Nat Genet 12(4):445–447
Edery P, Lyonnet S, Mulligan LM, Pelet A, Dow E, Abel L, Holder S, Nihoul-Fekete C, Ponder BA, Munnich A (1994) Mutations of the RET proto-oncogene in Hirschsprung’s disease. Nature 367(6461):378–380
Garcia-Barcelo MM, Miao X, Lui VC, So MT, Ngan ES, Leon TY, Lau DK, Liu TT, Lao X, Guo W, Holden WT, Moore J, Tam PK (2007) Correlation between genetic variations in Hox clusters and Hirschsprung’s disease. Ann Hum Genet 71(Pt 4):526–536
Miao X, Garcia-Barcelo MM, So MT, Leon TY, Lau DK, Liu TT, Chan EK, Lan LC, Wong KK, Lui VC, Tam PK (2007) Role of RET and PHOX2B gene polymorphisms in risk of Hirschsprung’s disease in Chinese population. Gut 56(5):736
Miao X, Leon TY, Ngan ES, So MT, Yuan ZW, Lui VC, Chen Y, Wong KK, Tam PK, Garcia-Barcelo M (2010) Reduced RET expression in gut tissue of individuals carrying risk alleles of Hirschsprung’s disease. Hum Mol Genet 19(8):1461–1467
Emison ES, McCallion AS, Kashuk CS, Bush RT, Grice E, Lin S, Portnoy ME, Cutler DJ, Green ED, Chakravarti A (2005) A common sex-dependent mutation in a RET enhancer underlies Hirschsprung disease risk. Nature 434(7035):857–863
Tang CS, Sribudiani Y, Miao XP, de Vries AR, Burzynski G, So MT, Leon YY, Yip BH, Osinga J, Hui KJ, Verheij JB, Cherny SS, Tam PK, Sham PC, Hofstra RM, Garcia-Barcelo MM (2010) Fine mapping of the 9q31 Hirschsprung’s disease locus. Hum Genet 127(6):675–683
Natarajan D, Marcos-Gutierrez C, Pachnis V, de Graaff E (2002) Requirement of signalling by receptor tyrosine kinase RET for the directed migration of enteric nervous system progenitor cells during mammalian embryogenesis. Development 129(22):5151–5160
Emison ES, Garcia-Barcelo M, Grice EA, Lantieri F, Amiel J, Burzynski G, Fernandez RM, Hao L, Kashuk C, West K, Miao X, Tam PK, Griseri P, Ceccherini I, Pelet A, Jannot AS, de Pontual L, Henrion-Caude A, Lyonnet S, Verheij JB, Hofstra RM, Antinolo G, Borrego S, McCallion AS, Chakravarti A (2010) Differential contributions of rare and common, coding and noncoding Ret mutations to multifactorial Hirschsprung disease liability. Am J Hum Genet 87(1):60–74
Majewski J, Schwartzentruber J, Lalonde E, Montpetit A, Jabado N (2011) What can exome sequencing do for you? J Med Genet 48(9):580–589
Ng PC, Kirkness EF (2010) Whole genome sequencing. Methods Mol Biol 628:215–226
Ng SB, Bigham AW, Buckingham KJ, Hannibal MC, McMillin MJ, Gildersleeve HI, Beck AE, Tabor HK, Cooper GM, Mefford HC, Lee C, Turner EH, Smith JD, Rieder MJ, Yoshiura K, Matsumoto N, Ohta T, Niikawa N, Nickerson DA, Bamshad MJ, Shendure J (2010) Exome sequencing identifies MLL2 mutations as a cause of Kabuki syndrome. Nat Genet 42(9):790–793
Adie EA, Adams RR, Evans KL, Porteous DJ, Pickard BS (2005) Speeding disease gene discovery by sequence based candidate prioritization. BMC Bioinforma 6:55
Chen J, Bardes EE, Aronow BJ, Jegga AG (2009) ToppGene Suite for gene list enrichment analysis and candidate gene prioritization. Nucleic Acids Res 37(Web Server issue):W305–W311
Aerts S, Lambrechts D, Maity S, Van Loo P, Coessens B, De Smet F, Tranchevent LC, De Moor B, Marynen P, Hassan B, Carmeliet P, Moreau Y (2006) Gene prioritization through genomic data fusion. Nat Biotechnol 24(5):537–544
Dahlquist KD, Salomonis N, Vranizan K, Lawlor SC, Conklin BR (2002) GenMAPP, a new tool for viewing and analyzing microarray data on biological pathways. Nat Genet 31(1):19–20
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7(4):248–249
Britsch S, Goerich DE, Riethmacher D, Peirano RI, Rossner M, Nave KA, Birchmeier C, Wegner M (2001) The transcription factor Sox10 is a key regulator of peripheral glial development. Genes Dev 15(1):66–78
Crone SA, Negro A, Trumpp A, Giovannini M, Lee KF (2003) Colonic epithelial expression of ErbB2 is required for postnatal maintenance of the enteric nervous system. Neuron 37(1):29–40
Altshuler D, Daly MJ, Lander ES (2008) Genetic mapping in human disease. Science 322(5903):881–888
Redon R, Ishikawa S, Fitch KR, Feuk L, Perry GH, Andrews TD, Fiegler H, Shapero MH, Carson AR, Chen W, Cho EK, Dallaire S, Freeman JL, Gonzalez JR, Gratacos M, Huang J, Kalaitzopoulos D, Komura D, MacDonald JR, Marshall CR, Mei R, Montgomery L, Nishimura K, Okamura K, Shen F, Somerville MJ, Tchinda J, Valsesia A, Woodwark C, Yang F, Zhang J, Zerjal T, Armengol L, Conrad DF, Estivill X, Tyler-Smith C, Carter NP, Aburatani H, Lee C, Jones KW, Scherer SW, Hurles ME (2006) Global variation in copy number in the human genome. Nature 444(7118):444–454
Burns AJ, Pachnis V (2009) Development of the enteric nervous system: bringing together cells, signals and genes. Neurogastroenterol Motil 21(2):100–102
Lemke G (1996) Neuregulins in development. Mol Cell Neurosci 7(4):247–262
Howard B, Panchal H, McCarthy A, Ashworth A (2005) Identification of the scaramanga gene implicates Neuregulin3 in mammary gland specification. Genes Dev 19(17):2078–2090
Howard BA (2008) The role of NRG3 in mammary development. J Mammary Gland Biol Neoplasia 13(2):195–203
Gizatullin RZ, Muravenko OV, Al-Amin AN, Wang F, Protopopov AI, Kashuba VI, Zelenin AV, Zabarovsky ER (2000) Human NRG3 gene Map position 10q22-q23. Chromosome Res 8(6):560
Britsch S (2007) The neuregulin-I/ErbB signaling system in development and disease. Adv Anat Embryol Cell Biol 190:1–65
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signalling network. Nat Rev Mol Cell Biol 2(2):127–137
Zhang D, Sliwkowski MX, Mark M, Frantz G, Akita R, Sun Y, Hillan K, Crowley C, Brush J, Godowski PJ (1997) Neuregulin-3 (NRG3): a novel neural tissue-enriched protein that binds and activates ErbB4. Proc Natl Acad Sci U S A 94(18):9562–9567
Tang CS, Cheng G, So MT, Yip BH, Miao XP, Wong EH, Ngan ES, Lui VC, Song YQ, Chan D, Cheung K, Yuan ZW, Lei L, Chung PH, Liu XL, Wong KK, Marshall CR, Scherer SW, Cherny SS, Sham PC, Tam PK, Garcia-Barcelo MM (2012) Genome-wide copy number analysis uncovers a new HSCR gene: NRG3. PLoS Genet 8(5):e1002687
Erlich Y, Edvardson S, Hodges E, Zenvirt S, Thekkat P, Shaag A, Dor T, Hannon GJ, Elpeleg O (2011) Exome sequencing and disease-network analysis of a single family implicate a mutation in KIF1A in hereditary spastic paraparesis. Genome Res 21(5):658–664
Acknowledgments
This work was supported by the Program for New Century Excellent Talents in University NCET-10-0388, Natural Science Foundation of Hubei 2010CDB03203 and 2011CDB304, and the Fundamental Research Funds for the Central Universities HUST: No. 2010QN005. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Conflict of interest
The authors have declared no conflicts of interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Jun Yang, Shengyu Duan, and Rong Zhong contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
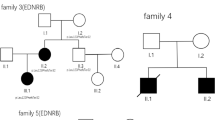
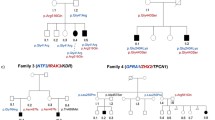
The pedigree of the affected Han Chinese family. Blue color represents HSCR patients. Patient II:3 is the mother and III:3 is the son (JPEG 23 kb)
Supplementary Fig. 2
The sequence diagram of the identified false positive variants in UNC5C and C22orf42 (JPEG 50 kb)
Supplementary Fig. 2b
(JPEG 42 kb)
ESM 1
(DOC 30 kb)
ESM 2
(DOCX 13 kb)
ESM 3
(DOCX 15 kb)
ESM 4
(PDF 128 kb)
ESM 5
(PDF 103 kb)
ESM 6
(PDF 134 kb)
ESM 7
(DOCX 13 kb)
ESM 8
(DOCX 13 kb)
Rights and permissions
About this article
Cite this article
Yang, J., Duan, S., Zhong, R. et al. Exome Sequencing Identified NRG3 as a Novel Susceptible Gene of Hirschsprung’s Disease in a Chinese Population. Mol Neurobiol 47, 957–966 (2013). https://doi.org/10.1007/s12035-012-8392-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-012-8392-4