Abstract
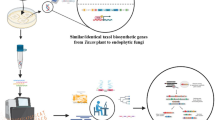
As a well-known industrial fungus for cellulase production, the strain RUT-C30 of Trichoderma reesei was selected to produce the feruloyl esterase A (FAEA) by a random integration protocol. The strong promoter of cellobiohydrolase 1 (cbh1) gene was used to drive the expression of FAEA. Using double-joint PCR protocol, Pcbh1-faeA-TtrpC expression cassette was successfully constructed and co-transformed into RUT C30 strain of T. reesei. One transformant with high feruloyl esterase yield (3.44 ± 0.16 IU/mL) was obtained through plate screening and named TrfaeA1. Sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) analysis of fermentation supernatant from transformant TrfaeA1 showed a distinct protein band appearing at the position of about 34 kDa, indicating that faeA gene has been successfully expressed in T. reesei. Compared with that in original RUT C30 strain, β-glucosidase production in transformant TrfaeA1 was significantly increased by about 86.4%, reaching 63.2 IU/mL due to the random insertion of faeA. Moreover, the total secretion protein and filter paper activities of the transformant TrfaeA1 were also improved by up to 5.5 and 4.3%, respectively. The present results indicated that the random insertion strategy could be an effective and feasible method to improve and optimize the cellulase system of filamentous fungi.





Similar content being viewed by others
References
Peterson, R., & Nevalarinen, H. (2012). Trichoderma reesei RUT C30—thirty years of strain improvement. Microbiology, 158, 58–68.
Schmoll, M., & Kubicek, C. P. (2003). Regulation of Trichoderma cellulase formation: lessons in molecular biology from an industrial fungus. Acta Microbiologica et Immunologica Hungarica, 50, 125–145.
Le Crom, S., Schackwitz, W., Pennacchio, L., Magnuson, J. K., Culley, D. E., Collett, J. R., et al. (2009). Tracking the roots of cellulase hyperproduction by the fungus Trichoderma reesei using massively parallel DNA sequencing. Proceedings of the National Academy of Sciences of the United States of America, 106, 16151–16156.
Bhatia, Y., Mishra, S., & Bisaria, V. S. (2002). Microbial beta-glucosidases: cloning, properties, and applications. Critical Reviews in Biotechnology, 22, 375–407.
Lynd, L. R., Weimer, P. J., van Zyl, W. H., & Pretorius, I. S. (2002). Microbial cellulose utilization: fundamentals and biotechnology. Microbiology and Molecular Biology Reviews, 66, 506–577.
Zhang, Y. H., & Lynd, L. R. (2004). Toward an aggregated understanding of enzymatic hydrolysis of cellulose: noncomplexed cellulase systems. Biotechnology and Bioengineering, 88, 797–824.
Saloheimo, M., Kuja-Panula, J., Ylösmäki, E., et al. (2002). Enzymatic properties and intracellular localization of the novel Trichoderma reesei β-glucosidase BGLII (Cel1A). Applied and Environmental Microbiology, 68, 4546–4553.
Zhang JW, Zhong YH, Zhao XN, Wang TH. (2010). Development of the cellulolytic fungus Trichoderma reesei strain with enhanced beta-glucosidase and filter paper activity using strong artifical cellobiohydrolase 1 promoter. Bioresour Technol, 101, 9815–9818.
Mikiko, N., Takanori, F., Yosuke, S., et al. (2012). A new Zn(II)2Cys(6)-type transcription factor BglR regulates β-glucosidase expression in Trichoderma reesei. Fungal Genetics and Biology, 49, 388–397.
Santhanam, P. (2012). Random insertional mutagenesis in fungal genomes to identify virulence factors. Methods in Molecular Biology, 835, 509–517.
Cai, Z., Li, G., Lin, C., et al. (2013). Identifying pathogenicity genes in the rubber tree anthracnose fungus Colletotrichum gloeosporioides through random insertional mutagenesis. Microbiological Research, 168, 340–350.
Ma, L., Zhang, J., Zou, G., Wang, C., & Zhou, Z. (2011). Improvement of cellulase activity in Trichoderma reesei by heterologous expression of a β-glucosidase gene from Penicillium decumbens. Enzyme Microb Tech, 49, 366–371.
Nakazawa, H., Kawai, T., et al. (2011). Construction of a recombinant Trichoderma reesei strain expressing Aspergillus aculeatus β-glucosidase 1 for efficient biomass conversion. Biotechnology and Bioengineering, 109, 92–99.
Dashtban, M., & Qin, W. (2012). Overexpression of an exotic thermotolerant β-glucosidase in Trichoderma reesei and its significant increase in cellulolytic activity and saccharification of barley straw. Microbial Cell Factories, 11, 63.
Xue, X. (2016). Revisiting overexpression of a heterologous β-glucosidase in Trichoderma reesei: fusion expression of the Neosartorya fischeri Bgl3A to cbh1 enhances the overall as well as individual cellulase activities. Microbial Cell Factories, 15, 1–13.
Wong, D. W. (2006). Feruloyl esterase: a key enzyme in biomass degradation. Applied Biochemistry and Biotechnology, 133, 87–112.
Kroon, P. A., Garcia-Conesa, M. T., Fillinghaim, I. J., et al. (1999). Release of ferulic acid dehydrodimers from plant cell walls by feruloyl esterases. Journal of the Science of Food and Agriculture, 79, 428–434.
Penttilä, M., Nevalainen, H., Rättö, M., Salminen, E., & Knowles, J. (1987). A versatile transformation system for the cellulolytic filamentous fungus Trichoderma reesei. Gene, 61, 155–164.
Yu, J. H., Hamari, Z., Han, K. H., et al. (2004). Double-joint PCR: a PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genetics and Biology, 41, 973–981.
Levasseur, A., Pagès, S., Fierobe, H.-P., et al. (2004). Design and production in Aspergillus niger of a chimeric protein associating a fungal feruloyl esterase and a clostridial dockerin domain. Applied and Environmental Microbiology, 70, 6984–6991.
Hegde, S., Srinivas, P., & Muralikrishna, G. (2009). Single-step synthesis of 4-nitrophenyl ferulate for spectrophotometric assay of feruloyl esterases. Analytical Biochemistry, 387, 128–129.
Laemmli, U. K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227, 680–685.
Murray, P., Aro, N., Collins, C., et al. (2004). Expression in Trichoderma reesei and characterisation of a thermostable family β-glucosidase from the moderately thermophilic fungus Talaromyces emersonii. Protein Expression Purification, 38, 248–257.
Faulds, C. B. (2010). What can feruloyl esterases do for us? Phytochemistry Reviews, 9, 121–132.
de Vries, R. P., Michelsen, B., Poulsen, C. H., et al. (1997). The faeA genes from Aspergillus niger and Aspergillus tubingensis encode ferulic acid esterases involved in degradation of complex cell wall polysaccharides. Applied and Environmental Microbiology, 63, 4638–4644.
Record, E., Asther, M., Sigoillot, C., et al. (2003). Overproduction of the Aspergillus niger, feruloyl esterase for pulp bleaching application. Applied Microbiology and Biotechnology, 62, 349–355.
Verdoes, J. C., van Diepeningen, A. D., Punt, P. J., et al. (1994). Evaluation of molecular and genetic approaches to generate glucoamylase overproducing strains of Aspergillus niger. Journal of Biotechnology, 36, 165–175.
Acknowledgements
The authors are grateful for the financial support from National Natural Science Foundation of China (Grant No. 31570118); S&T Plan Projects of Shandong Provincial Education Department (J10LC08); and Shandong Provincial Natural Science Foundation, China (Grant No. ZR2015CM029).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hou, Y., Pan, Y., Yan, M. et al. Influence of Randomly Inserted Feruloyl Esterase A on β-Glucosidase Activity in Trichoderma reesei . Appl Biochem Biotechnol 183, 254–264 (2017). https://doi.org/10.1007/s12010-017-2442-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-017-2442-3