Abstract
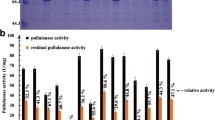
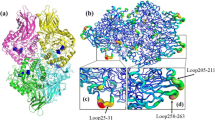
Based on conserved sites and homology modeling analysis, the residue Phe581 in the Klebsiella variicola SHN-1 pullulanase was selected as the potential thermostability-related site and its role on thermostability and activity was investigated by site-saturated mutagenesis. Compared with the wild-type pullulanase, the optimum temperature of the mutants including F581L, F581Q, F581R, F581T, F581V, and F581Y was increased from 53 to 56 °C, and correspondingly the half lives of these mutants at 55 °C were increased by 4.20, 3.70, 1.90, 7.16, 3.01, and 1.75 min, respectively. By modeling the structure of the pullulanase, formation of more hydrogen bonds by single-site substitution was supposed to be responsible for the improvement of thermostability. Of these mutants, furthermore, F581L and F581V exhibited higher values of V max and k cat/K m, compared with the wild-type enzyme. Therefore, the residue Phe581 was identified as an important site relevant to the activity and thermostability of the pullulanase of K. variicola, and by mutation at this single site, the mutated enzymes with enhanced thermostability and catalytic efficiency were achieved consequently.




Similar content being viewed by others
References
Wallenfels, K., Bender, H., & Rached, J. (1966). Pullulanase from Aerobacter aerogenes; production in a cell-bound state. Purification and properties of the enzyme. Biochemical and Biophysical Research Communications, 22(3), 254–261.
Roy, I., & Gupta, M. N. (2004). Hydrolysis of starch by a mixture of glucoamylase and pullulanase entrapped individually in calcium alginate beads. Enzyme and Microbial Technology, 34(1), 26–32.
Duan, X., Chen, J., & Wu, J. (2013). Improving the thermostability and catalytic efficiency of Bacillus deramificans pullulanase by site-directed mutagenesis. Applied and Environmental Microbiology, 79(13), 4072–4077.
Bertoldo, C., Armbrecht, M., Becker, F., Schäfer, T., Antranikian, G., & Liebl, W. (2004). Cloning, sequencing, and characterization of a heat-and alkali-stable type I pullulanase from Anaerobranca gottschalkii. Applied and Environmental Microbiology, 70(6), 3407–3416.
Schülein, M., & Højer-Pedersen, B. (1984). Characterization of a new class of thermophilic pullulanases from Bacillus acidopullulyticus. Annals of the New York Academy of Sciences, 434(1), 271–274.
Takasaki, Y. (1987). Pullulanase-amylase complex enzyme from Bacillus subtilis. Agricultural and Biological Chemistry, 51(1), 9–16.
Plant, A. R., Morgan, H. W., & Daniel, R. M. (1986). A highly stable pullulanase from Thermus aquaticus YT-1. Enzyme and Microbial Technology, 8(11), 668–672.
Duffner, F., Bertoldo, C., Andersen, J. T., Wagner, K., & Antranikian, G. (2000). A new thermoactive pullulanase from Desulfurococcus mucosus: cloning, sequencing, purification, and characterization of the recombinant enzyme after expression in Bacillus subtilis. Journal of Bacteriology, 182(22), 6331–6338.
Zeng, Q., Du, H., Wang, J., Wei, D., Wang, X., Li, Y., et al. (2009). Reversal of coenzyme specificity and improvement of catalytic efficiency of Pichia stipitis xylose reductase by rational site-directed mutagenesis. Biotechnology Letters, 31(7), 1025–1029.
Deng, Z., Yang, H., Li, J., Shin, H. D., Du, G., Liu, L., et al. (2014). Structure-based engineering of alkaline alpha-amylase from alkaliphilic Alkalimonas amylolytica for improved thermostability. Applied Microbiology and Biotechnology, 98(9), 3997–4007.
Liao, Y., Li, C., Chen, H., Wu, Q., Shan, Z., & Han, X. (2013). Site-directed mutagenesis improves the thermostability and catalytic efficiency of Aspergillus niger N25 phytase mutated by I44E and T252R. Applied Biochemistry and Biotechnology, 171(4), 900–915.
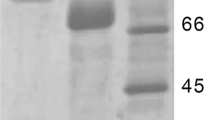
Chen, W. B., Nie, Y., & Xu, Y. (2013). Signal peptide-independent secretory expression and characterization of pullulanase from a newly isolated Klebsiella variicola SHN-1 in Escherichia coli. Applied Biochemistry and Biotechnology, 169(1), 41–54.
Yamashita, M., Kinoshita, T., Ihara, M., Mikawa, T., & Murooka, Y. (1994). Random mutagenesis of pullulanase from Klebsiella aerogenes for studies of the structure and function of the enzyme. Journal of Biochemistry, 116(6), 1233–1240.
Miyazaki, K., & Takenouchi, M. (2002). Creating random mutagenesis libraries using megaprimer PCR of whole plasmid. Biotechniques, 33(5), 1033–1038.
Eom, G. T., Lee, S. H., Song, B. K., Chung, K. W., Kim, Y. W., & Song, J. K. (2013). High-level extracellular production and characterization of Candida antarctica lipase B in Pichia pastoris. Journal of Bioscience and Bioengineering, 116(2), 165–170.
Hii, L., Rosfarizan, M., Ling, T., & Ariff, A. (2012). Statistical optimization of pullulanase production by Raoultella planticola DSMZ 4617 using sago starch as carbon and peptone as nitrogen sources. Food and Bioprocess Technology, 5(2), 729–737.
Malle, D., Itoh, T., Hashimoto, W., Murata, K., Utsumi, S., & Mikami, B. (2006). Overexpression, purification and preliminary X-ray analysis of pullulanase from Bacillus subtilis strain 168. Acta Crystallographica Section F: Structural Biology and Crystallization Communications, 62(Pt 4), 381–384.
Zhou, C., Xue, Y., & Ma, Y. (2010). Enhancing the thermostability of alpha-glucosidase from Thermoanaerobacter tengcongensis MB4 by single proline substitution. Journal of Bioscience and Bioengineering, 110(1), 12–17.
Kang, J., Park, K.-M., Choi, K.-H., Park, C.-S., Kim, G.-E., Kim, D., et al. (2011). Molecular cloning and biochemical characterization of a heat-stable type I pullulanase from Thermotoga neapolitana. Enzyme and Microbial Technology, 48(3), 260–266.
Vieille, C., & Gregory Zeikus, J. (1996). Thermozymes: identifying molecular determinants of protein structural and functional stability. Trends in Biotechnology, 14(6), 183–190.
Voigt, C. A., Mayo, S. L., Arnold, F. H., & Wang, Z.-G. (2001). Computational method to reduce the search space for directed protein evolution. Proceedings of the National Academy of Sciences, 98(7), 3778–3783.
Vogt, G., Woell, S., & Argos, P. (1997). Protein thermal stability, hydrogen bonds, and ion pairs. Journal of Molecular Biology, 269(4), 631–643.
Zhang, W., Mullaney, E. J., & Lei, X. G. (2007). Adopting selected hydrogen bonding and ionic interactions from Aspergillus fumigatus phytase structure improves the thermostability of Aspergillus niger PhyA phytase. Applied and Environmental Microbiology, 73(9), 3069–3076.
Yutani, K., Ogasahara, K., Tsujita, T., & Sugino, Y. (1987). Dependence of conformational stability on hydrophobicity of the amino acid residue in a series of variant proteins substituted at a unique position of tryptophan synthase alpha subunit. Proceedings of the National Academy of Sciences, 84(13), 4441–4444.
Tian, Y., Peng, R., Xu, J., Zhao, W., Gao, F., Fu, X., et al. (2010). Mutations in two amino acids in phyI1s from Aspergillus niger 113 improve its phytase activity. World Journal of Microbiology and Biotechnology, 26(5), 903–907.
Zhang, Y. H. P., Himmel, M. E., & Mielenz, J. R. (2006). Outlook for cellulase improvement: screening and selection strategies. Biotechnology Advances, 24(5), 452–481.
van der Veen, B. A., Potocki-Véronèse, G., Albenne, C., Joucla, G., Monsan, P., & Remaud-Simeon, M. (2004). Combinatorial engineering to enhance amylosucrase performance: construction, selection, and screening of variant libraries for increased activity. FEBS Letters, 560(1), 91–97.
Ben Ali, M., Khemakhem, B., Robert, X., Haser, R., & Bejar, S. (2006). Thermostability enhancement and change in starch hydrolysis profile of the maltohexaose-forming amylase of Bacillus stearothermophilus US100 strain. Biochemical Journal, 394(1), 51–56.
Acknowledgments
Financial supports from the National Key Basic Research and Development Program of China (973 Program) (2011CB710800), the National High Technology Research and Development Program of China (863 Program) (2012AA022207), the National Natural Science Foundation of China (NSFC) (21376107 and 21336009), the 111 Project (111-2-06), the High-end Foreign Experts Recruitment Program (GDW20133200113), the Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions, and the Jiangsu province “Collaborative Innovation Center for Advanced Industrial Fermentation” industry development program are greatly appreciated.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 3525 kb)
Rights and permissions
About this article
Cite this article
Mu, G.C., Nie, Y., Mu, X.Q. et al. Single Amino Acid Substitution in the Pullulanase of Klebsiella variicola for Enhancing Thermostability and Catalytic Efficiency. Appl Biochem Biotechnol 176, 1736–1745 (2015). https://doi.org/10.1007/s12010-015-1675-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-015-1675-2