Abstract
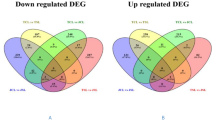
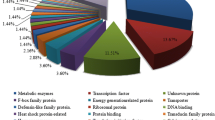
Pakchoi is an important vegetable grown in worldwide that prefers a cool climate, and its growth is severely affected by high temperatures in the summer. Little is known for the molecular regulatory mechanisms of high-temperature stress in pakchoi. The limited availability of gene sequence has greatly affected molecular breeding and functional genomic analysis. In this study, we conducted comprehensive analyses for heat treatment in pakchoi by RNA-Seq. Finally, 64.29 million clean reads containing 32,666 unigenes with an N50 length of 1405 bp, and a total length of 33.39 Mb were identified. Overall, 11,024 SSRs located in 8404 unigenes were obtained; these findings will be very useful for molecular assisted breeding. Totally, 1220 differentially expressed genes (DEGs) were detected between the high-temperature group (TH) and the control (CK), among which 699 DEGs were up-regulated and 521 were down-regulated. The enrichment analysis indicated that 12 GO subcategories and 9 KEGG pathways were enriched significantly (P value <0.05). Finally, six DEGs were validated by qRT-PCR, and the results further verified the reliability of RNA-Seq. Herein, we present the first comprehensive characterization of heat-treated pakchoi using transcriptome analysis. Several important genes that respond to high temperature were identified, and their roles in heat stress responses are discussed. In conclusion, our study represents a fully characterised pakchoi transcriptome. In addition, it provides the important resources for future genomic and genetic studies of pakchoi under heat stress.






Similar content being viewed by others
References
Anders S, Huber W (2010) Differential expression analysis for sequence count data. Genome Biol 11:R106
Bowman MJ, Park W, Bauer PJ, Udall JA, Page JT, Raney J, Scheffler BE, Jones DC, Campbell BT (2013) RNA-Seq transcriptome profiling of upland cotton (Gossypium hirsutum L.) root tissue under water-deficit stress. PLoS One 8:e82634
Carbonell-Bejerano P, Santa Maria E, Torres-Perez R, Royo C, Lijavetzky D, Bravo G, Aguirreolea J, Sanchez-Diaz M, Antolin MC, Martinez-Zapater JM (2013) Thermotolerance responses in ripening berries of Vitis vinifera L. cv Muscat Hamburg. Plant Cell Physiol 54:1200–1216
Chen B, Walser JC, Rodgers TH, Sobota RS, Burke MK, Rose MR, Feder ME (2007) Abundant, diverse, and consequential P elements segregate in promoters of small heat-shock genes in Drosophila populations. J Evol Biol 20:2056–2066
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
da Hao C, Ge G, Xiao P, Zhang Y, Yang L (2011) The first insight into the tissue specific taxus transcriptome via Illumina second generation sequencing. PLoS One 6:e21220
Fullwood MJ, Wei CL, Liu ET, Ruan Y (2009) Next-generation DNA sequencing of paired-end tags (PET) for transcriptome and genome analyses. Genome Res 19:521–532
Gotz S, Garcia-Gomez JM, Terol J, Williams TD, Nagaraj SH, Nueda MJ, Robles M, Talon M, Dopazo J, Conesa A (2008) High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res 36:3420–3435
Gracey AY, Fraser EJ, Li W, Fang Y, Taylor RR, Rogers J, Brass A, Cossins AR (2004) Coping with cold: an integrative, multitissue analysis of the transcriptome of a poikilothermic vertebrate. Proc Natl Acad Sci USA 101:16970–16975
Gulick PJ, Drouin S, Yu Z, Danyluk J, Poisson G, Monroy AF, Sarhan F (2005) Transcriptome comparison of winter and spring wheat responding to low temperature. Genome 48:913–923
Guy CL, Li QB (1998) The organization and evolution of the spinach stress 70 molecular chaperone gene family. Plant Cell 10:539–556
He CY, Cui K, Zhang JG, Duan AG, Zeng YF (2013) Next-generation sequencing-based mRNA and microRNA expression profiling analysis revealed pathways involved in the rapid growth of developing culms in Moso bamboo. BMC Plant Biol 13:119
Hiz MC, Canher B, Niron H, Turet M (2014) Transcriptome analysis of salt tolerant common bean (Phaseolus vulgaris L.) under saline conditions. PLoS One 9:e92598
Hsieh EJ, Cheng MC, Lin TP (2013) Functional characterization of an abiotic stress-inducible transcription factor AtERF53 in Arabidopsis thaliana. Plant Mol Biol 82:223–237
Hu J, You F, Wang Q, Weng S, Liu H, Wang L, Zhang PJ, Tan X (2014) Transcriptional responses of olive flounder (Paralichthys olivaceus) to low temperature. PLoS One 9:e108582
Kang L, Chen B, Wei JN, Liu TX (2009) Roles of thermal adaptation and chemical ecology in Liriomyza distribution and control. Annu Rev Entomol 54:127–145
Li QB, Haskell DW, Guy CL (1999) Coordinate and non-coordinate expression of the stress 70 family and other molecular chaperones at high and low temperature in spinach and tomato. Plant Mol Biol 39:21–34
Lin BL, Wang JS, Liu HC, Chen RW, Meyer Y, Barakat A, Delseny M (2001) Genomic analysis of the Hsp70 superfamily in Arabidopsis thaliana. Cell Stress Chaperones 6:201–208
Liu S, Kuang H, Lai Z (2014) Transcriptome analysis by Illumina high-throughout paired-end sequencing reveals the complexity of differential gene expression during in vitro plantlet growth and flowering in Amaranthus tricolor L. PLoS One 9:e100919
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA, Berka J, Braverman MS, Chen YJ, Chen Z, Dewell SB, Du L, Fierro JM, Gomes XV, Godwin BC, He W, Helgesen S, Ho CH, Irzyk GP, Jando SC, Alenquer ML, Jarvie TP, Jirage KB, Kim JB, Knight JR, Lanza JR, Leamon JH, Lefkowitz SM, Lei M, Li J, Lohman KL, Lu H, Makhijani VB, McDade KE, McKenna MP, Myers EW, Nickerson E, Nobile JR, Plant R, Puc BP, Ronan MT, Roth GT, Sarkis GJ, Simons JF, Simpson JW, Srinivasan M, Tartaro KR, Tomasz A, Vogt KA, Volkmer GA, Wang SH, Wang Y, Weiner MP, Yu P, Begley RF, Rothberg JM (2005) Genome sequencing in microfabricated high-density picolitre reactors. Nature 437:376–380
Mishkind M, Vermeer JE, Darwish E, Munnik T (2009) Heat stress activates phospholipase D and triggers PIP accumulation at the plasma membrane and nucleus. Plant J 60:10–21
Montero-Barrientos M, Hermosa R, Cardoza RE, Gutierrez S, Nicolas C, Monte E (2010) Transgenic expression of the Trichoderma harzianum hsp70 gene increases Arabidopsis resistance to heat and other abiotic stresses. J Plant Physiol 167:659–665
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Natarajan P, Parani M (2011) De novo assembly and transcriptome analysis of five major tissues of Jatropha curcas L. using GS FLX titanium platform of 454 pyrosequencing. BMC Genom 12:191
Oono Y, Yazawa T, Kawahara Y, Kanamori H, Kobayashi F, Sasaki H, Mori S, Wu J, Handa H, Itoh T, Matsumoto T (2014) Genome-wide transcriptome analysis reveals that cadmium stress signaling controls the expression of genes in drought stress signal pathways in rice. PLoS One 9:e96946
Pan J, Tristram-Nagle S, Kucerka N, Nagle JF (2008) Temperature dependence of structure, bending rigidity, and bilayer interactions of dioleoylphosphatidylcholine bilayers. Biophys J 94:117–124
Rocheta M, Becker JD, Coito JL, Carvalho L, Amancio S (2014) Heat and water stress induce unique transcriptional signatures of heat-shock proteins and transcription factors in grapevine. Funct Integr Genomics 14:135–148
Saidi Y, Finka A, Muriset M, Bromberg Z, Weiss YG, Maathuis FJ, Goloubinoff P (2009) The heat shock response in moss plants is regulated by specific calcium-permeable channels in the plasma membrane. Plant Cell 21:2829–2843
Scott GR, Johnston IA (2012) Temperature during embryonic development has persistent effects on thermal acclimation capacity in zebrafish. Proc Natl Acad Sci USA 109:14247–14252
Simon SA, Advani S, McIntosh TJ (1995) Temperature dependence of the repulsive pressure between phosphatidylcholine bilayers. Biophys J 69:1473–1483
Sokabe T, Tsujiuchi S, Kadowaki T, Tominaga M (2008) Drosophila painless is a Ca2+-requiring channel activated by noxious heat. J Neurosci 28:9929–9938
Somero GN (1995) Proteins and temperature. Annu Rev Physiol 57:43–68
Song X, Ge T, Li Y, Hou X (2015) Genome-wide identification of SSR and SNP markers from the non-heading Chinese cabbage for comparative genomic analyses. BMC Genom 16:328
Sung DY, Vierling E, Guy CL (2001) Comprehensive expression profile analysis of the Arabidopsis Hsp70 gene family. Plant Physiol 126:789–800
Swindell WR, Huebner M, Weber AP (2007) Plastic and adaptive gene expression patterns associated with temperature stress in Arabidopsis thaliana. Heredity (Edinb) 99:143–150
Ueda MJ, Ito T, Okada TS, Ohnishi SI (1976) A correlation between membrane fluidity and the critical temperature for cell adhesion. J Cell Biol 71:670–674
Van Verk MC, Hickman R, Pieterse CM, Van Wees SC (2013) RNA-Seq: revelation of the messengers. Trends Plant Sci 18:175–179
Walser JC, Chen B, Feder ME (2006) Heat-shock promoters: targets for evolution by P transposable elements in Drosophila. PLoS Genet 2:e165
Wei W, Qi X, Wang L, Zhang Y, Hua W, Li D, Lv H, Zhang X (2011) Characterization of the sesame (Sesamum indicum L.) global transcriptome using Illumina paired-end sequencing and development of EST-SSR markers. BMC Genom 12:451
Wray GA (2007) The evolutionary significance of cis-regulatory mutations. Nat Rev Genet 8:206–216
Xiao B, Coste B, Mathur J, Patapoutian A (2011) Temperature-dependent STIM1 activation induces Ca2+ influx and modulates gene expression. Nat Chem Biol 7:351–358
Xiong Y, Li QB, Kang BH, Chourey P (2011) Discovery of genes expressed in basal endosperm transfer cells in maize using 454 transcriptome sequencing. Plant Mol Biol Rep 29:835–847
Xu J, Xue C, Xue D, Zhao J, Gai J, Guo N, Xing H (2013) Overexpression of GmHsp90 s, a heat shock protein 90 (Hsp90) gene family cloning from soybean, decrease damage of abiotic stresses in Arabidopsis thaliana. PLoS One 8:e69810
Yang N, Yue XL, Chen XL, Wu GF, Zhang TG, An LZ (2012) Molecular cloning and partial characterization of a novel phospholipase D gene from Chorispora bungeana. Plant Cell Tissue Organ Culture (PCTOC) 108:201–212
Zhang G, Guo G, Hu X, Zhang Y, Li Q, Li R, Zhuang R, Lu Z, He Z, Fang X, Chen L, Tian W, Tao Y, Kristiansen K, Zhang X, Li S, Yang H, Wang J, Wang J (2010) Deep RNA sequencing at single base-pair resolution reveals high complexity of the rice transcriptome. Genome Res 20:646–654
Zhang X, Li J, Liu A, Zou J, Zhou X, Xiang J, Rerksiri W, Peng Y, Xiong X, Chen X (2012) Expression profile in rice panicle: insights into heat response mechanism at reproductive stage. PLoS One 7:e49652
Acknowledgments
This study was funded by the National Key Science and Technology Supporting Program of China (Grant No. 2013BAD01B04-11) and by the Jiangsu agricultural science and technology innovation fund of China [Grant No. CX (13) 2004]. We also thank Genepioneer Biotechnologies for their help with the bioinformatics analyses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
Not relevant.
Additional information
Communicated by W. Zhou.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, H., Chen, L., Song, B. et al. De novo transcriptome sequencing of pakchoi (Brassica rapa L. chinensis) reveals the key genes related to the response of heat stress. Acta Physiol Plant 38, 252 (2016). https://doi.org/10.1007/s11738-016-2269-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-016-2269-5