Abstract
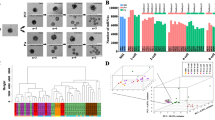
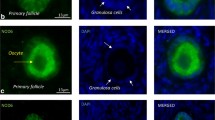
Several germ cell-specific transcription factors essential for ovarian formation and folliculogenesis have been identified and studied. However, their function during early embryo development has been poorly explored. In this study, we investigated the role of mixed-lineage leukemia protein 2 (MLL2) in the development of porcine preimplantation embryos. To explore the function of MLL2 in porcine embryo development, expression and localization of MLL2 were assessed by qRT-PCR and immunofluorescence assays. Results showed that expression of MLL2 was significantly reduced after the four-cell stage. However, immunofluorescence results showed that MLL2 only localized in the nucleus of blastocysts, revealing a potential role of MLL2 in regulating the gene expression in the blastocyst stage. Knockdown of Mll2 by double-stranded RNA (dsRNA) caused a reduction in MLL2 signal in porcine blastocyst cells and in blastocyst formation. No significant differences in the cleavage and morula stages were observed. The mechanism of MLL2 regulation in blastocysts was assessed by assaying the trimethylation of histone 3 at lysine 4 (H3K4m3). MLL2 knockdown significantly reduced H3K4m3 in the nucleus and further reduced expression of Sox2 and Magoh genes. In conclusion, MLL2 is essential for porcine embryo development by the regulation of methylation of H3K4 in vitro.






Similar content being viewed by others
References
Andreu-Vieyra CV, Chen R, Agno JE, Glaser S, Anastassiadis K, Stewart AF, Matzuk MM (2010) MLL2 is required in oocytes for bulk histone 3 lysine 4 trimethylation and transcriptional silencing. PLoS Biol 8
Denissov S, Hofemeister H, Marks H, Kranz A, Ciotta G, Singh S, Anastassiadis K, Stunnenberg HG, Stewart AF (2014) Mll2 is required for H3K4 trimethylation on bivalent promoters in embryonic stem cells, whereas Mll1 is redundant. Development 141:526–537
Glaser S, Lubitz S, Loveland KL, Ohbo K, Robb L, Schwenk F, Seibler J, Roellig D, Kranz A, Anastassiadis K, Stewart AF (2009) The histone 3 lysine 4 methyltransferase, Mll2, is only required briefly in development and spermatogenesis. Epigenetics Chromatin 2:5
Glaser S, Schaft J, Lubitz S, Vintersten K, van der Hoeven F, Tufteland KR, Aasland R, Anastassiadis K, Ang SL, Stewart AF (2006) Multiple epigenetic maintenance factors implicated by the loss of Mll2 in mouse development. Development 133:1423–1432
Keramari M, Razavi J, Ingman KA, Patsch C, Edenhofer F, Ward CM, Kimber SJ (2010) Sox2 is essential for formation of trophectoderm in the preimplantation embryo. PLoS One 5:e13952
Ladopoulos V, Hofemeister H, Hoogenkamp M, Riggs AD, Stewart AF, Bonifer C (2013) The histone methyltransferase KMT2B is required for RNA polymerase II association and protection from DNA methylation at the MagohB CpG island promoter. Mol Cell Biol 33:1383–1393
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408
Lorthongpanich C, Solter D, Lim CY (2010) Nuclear reprogramming in zygotes. Int J Dev Biol 54:1631–1640
Milne TA, Briggs SD, Brock HW, Martin ME, Gibbs D, Allis CD, Hess JL (2002) MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol Cell 10:1107–1117
Rao RC, Dou Y (2015) Hijacked in cancer: the KMT2 (MLL) family of methyltransferases. Nat Rev Cancer 15:334–346
Roguev A, Schaft D, Shevchenko A, Pijnappel WW, Wilm M, Aasland R, Stewart AF (2001) The Saccharomyces cerevisiae Set1 complex includes an Ash2 homologue and methylates histone 3 lysine 4. EMBO J 20:7137–7148
Ruthenburg AJ, Allis CD, Wysocka J (2007) Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol Cell 25:15–30
Schmidt R, Plath K (2012) The roles of the reprogramming factors Oct4, Sox2 and Klf4 in resetting the somatic cell epigenome during induced pluripotent stem cell generation. Genome Biol 13:251
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH Image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Smallwood SA, Kelsey G (2012) De novo DNA methylation: a germ cell perspective. Trends Genet : TIG 28:33–42
Wei Z, Yang Y, Zhang P, Andrianakos R, Hasegawa K, Lyu J, Chen X, Bai G, Liu C, Pera M, Lu W (2009) Klf4 interacts directly with Oct4 and Sox2 to promote reprogramming. Stem Cells 27:2969–2978
Yang X, Smith SL, Tian XC, Lewin HA, Renard JP, Wakayama T (2007) Nuclear reprogramming of cloned embryos and its implications for therapeutic cloning. Nat Genet 39:295–302
Yokoyama A, Wang Z, Wysocka J, Sanyal M, Aufiero DJ, Kitabayashi I, Herr W, Cleary ML (2004) Leukemia proto-oncoprotein MLL forms a SET1-like histone methyltransferase complex with menin to regulate Hox gene expression. Mol Cell Biol 24:5639–5649
Zhang A, Xu B, Sun Y, Lu X, Gu R, Wu L, Feng Y, Xu C (2012) Dynamic changes of histone H3 trimethylated at positions K4 and K27 in human oocytes and preimplantation embryos. Fertil Steril 98:1009–1016
Zhang S, Cui W (2014) Sox2, a key factor in the regulation of pluripotency and neural differentiation. World J Stem Cells 6:305–311
Zhao MH, Kim NH, Cui XS (2016) GlutaMAX prolongs the shelf life of the culture medium for porcine parthenotes. Theriogenology 85:368–375
Zhao MH, Kwon JW, Liang S, Kim SH, Li YH, Oh JS, Kim NH, Cui XS (2014) Zinc regulates meiotic resumption in porcine oocytes via a protein kinase C-related pathway. PLoS One 9:e102097
Acknowledgments
This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (No. 2015R1D1A1A01057629), and the Next-Generation BioGreen 21 Program (PJ011126), Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Editor: Tetsuji Okamoto
Rights and permissions
About this article
Cite this article
Zhao, MH., Liang, S., Kim, NH. et al. MLL2 is essential for porcine embryo development in vitro. In Vitro Cell.Dev.Biol.-Animal 52, 699–704 (2016). https://doi.org/10.1007/s11626-016-0017-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11626-016-0017-1