Summary
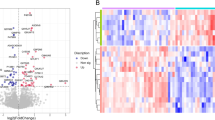
To identify acute renal allograft rejection biomarkers in human serum, two-dimensional differential in-gel electrophoresis (2-D DIGE) and reversed phase high-performance liquid chromatography (RP-HPLC) followed by electrospray ionization mass spectrometry (ESI-MS) were used. Serum samples from renal allograft patients and normal volunteers were divided into three groups: acute rejection (AR), stable renal function (SRF) and normal volunteer (N). Serum samples were firstly processed using Multiple Affinity Removal Column to selectively remove the highest abundance proteins. Differentially expressed proteins were analyzed using 2-D DIGE. These differential protein spots were excised, digested by trypsin, and identified by RP-HPLC-ESI/MS. Twenty-two differentially expressed proteins were identified in serum from AR group. These proteins included complement C9 precursor, apolipoprotein A-IV precursor, vitamin D-binding protein precursor, beta-2-glycoprotein 1 precursor, etc. Vitamin D-binding protein, one of these proteins, was confirmed by ELISA in the independent set of serum samples. In conclusion, the differentially expressed proteins as serum biomarker candidates may provide the basis of acute rejection noninvasive diagnosis. Confirmed vitamin D-binding protein may be one of serum biomarkers of acute rejection. Furthermore, it may provide great insights into understanding the mechanisms and potential treatment strategy of acute rejection.
Similar content being viewed by others
References
Hariharan S, Johnson CP, Bresnahan BA et al. Improved graft survival after renal transplantation in the United States, 1998–1996. N Engl J Med, 2000, 342(9):605–612
McLaren AJ, Fuggle SV, Welsh KI et al. Chronic allograft failure in human renal transplantation: a multivariate risk factor analysis. Ann Surg, 2000,232(1):98–103
Benfield MR, Herrin J, Feld L et al. Safety of kidney biopsy in pediatric transplantation: a report of the controlled clinical trials in pediatric transplantation trial of induction therapy study group. Transplantation, 1999,67(4):544–547
Traum AZ, Schachter AD. Transplantation proteomics. Pediatr Transplant, 2005,9(6):700–711
Solez K, Colvin RB, Racusen LC, et al. Banff’ 05 Meeting Report: differential diagnosis of chronic allograft injury and elimination of chronic allograft nephropathy (’CAN’). Am J Transplant, 2007,7(3):518–526
Peng J, Elias JE, Thoreen CC, et al. Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for large-scale protein analysis: the yeast proteome. J Proteome Res, 2003,2(1): 43–50
Anderson NL, Polanski M, Pieper R, et al. The human plasma proteome: a nonredundant list developed by combination of four separate sources. Mol Cell Proteomics, 2004, 3(4):311–326
Huang HL, Stasyk T, Morandell S, et al. Biomarker discovery in breast cancer serum using 2-D differential gel electrophoresis/ MALDI-TOF/TOF and data validation by routine clinical assays. Electrophoresis, 2006,27(8):1641–1650
Schaub S, Wilkins JA, Nickerson P. Proteomics and renal transplantation: searching for novel biomarkers and therapeutic targets. Contrib Nephrol, 2008,160:65–75
Clarke W. Proteomic research in renal transplantation. Ther Drug Monit, 2006, 28(1):19–22
Schaub S, Wilkins JA, Antonovici M, et al. Proteomic-based identification of cleaved urinarybeta2-microglobulin as a potential marker for acute tubular injury in renal allografts. Am J Transplant, 2005,5(4 Pt 1):729–738
Wittke S, Haubitz M, Walden M, et al. Detection of acute tubulointerstitial rejection by proteomic analysis of urinary samples in renal transplant recipients. Am J Transplant, 2005, 5(10):2479–2488
El Essawy B, Otu HH, Choy B, et al. Proteomic analysis of the allograft response. Transplantation, 2006,82(2): 267–274
Voshol H, Brendlen N, Müller D, et al. Evaluation of biomarker discovery approaches to detect protein biomarkers of acute renal allograft rejection. J Proteome Res, 2005,4(4):1192–1199
Yu KH, Rustgi AK, Blair IA. Characterization of proteins in human pancreatic cancer serum using differential gel electrophoresis and tandem mass spectrometry. J Proteome Res, 2005,4(5):1742–1751
Schiødt FV, Rossaro L, Stravitz RT, et al. Gc-globulin and prognosis in acute liver failure. Liver Transpl, 2005,11(10):1223–1227
Speeckaert MM, Glorieux GL, Vanholder R, et al. Vitamin D binding protein and the need for vitamin D in hemodialysis patients. J Ren Nutr, 2008,18(5):400–407
Zella LA, Shevde NK, Hollis BW, et al. Vitamin D-binding protein influences total circulating levels of 1,25-dihydroxyvitamin D3 but does not directly modulate the bioactive levels of the hormone in vivo. Endocrinology, 2008,149(7):3656–3667
Christakos S, Dhawan P, Liu Y, et al. New insights into the mechanisms of vitamin D action. J Cell Biochem, 2003,88(4):695–705
Mathieu C, Jafari M. Immunomodulation by 1,25-dihydroxyvitamin D3: therapeutic implications in hemodialysis and renal transplantation. Clin Nephrol, 2006,66(4):275–283
Koga Y, Naraparaju VR, Yamamoto N. Antitumor effect of vitamin D-binding protein-derived macrophage activating factor on Ehrlich ascites tumor-bearing mice. Proc Soc Exp Biol Med, 1999,220(1):20–26
Yamamoto N, Suyama H, Ushijima N. Immunotherapy of metastatic breast cancer patients with vitamin D-binding protein-derived macrophage activating factor (GcMAF). Int J Cancer, 2008, 122(2):461–467
Yamamoto N, Suyama H. Immunotherapy for prostate cancer with gc protein-derived macrophage-activating factor, GcMAF. Transl Oncol, 2008,1(2):65–72
Nickeleit V, Andreoni K. Inflammatory cells in renal allografts. Front Biosci, 2008,13:6202–6213
Qi F, Adair A, Ferenbach D, et al. Depletion of cells of monocyte lineage prevents loss of renal microvasculature in murine kidney transplantation. Transplantation, 2008,86(9):1267–1274
Author information
Authors and Affiliations
Additional information
This project was supported by a grant from National Basic Research 973 Program of China (No. 2009CB522407).
Rights and permissions
About this article
Cite this article
Gao, Y., Wu, K., Xu, Y. et al. Characterization of acute renal allograft rejection by human serum proteomic analysis. J. Huazhong Univ. Sci. Technol. [Med. Sci.] 29, 585–591 (2009). https://doi.org/10.1007/s11596-009-0511-8
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11596-009-0511-8