Abstract
Purpose
Combination of various intraoperative imaging modalities potentially can reduce error of brain shift estimation during neurosurgical operations. In the present work, a new combination of surface imaging and Doppler US images is proposed to calculate the displacements of cortical surface and deformation of internal vessels in order to estimate the targeted brain shift using a Finite Element Model (FEM). Registration error in each step and the overall performance of the method are evaluated.
Methods
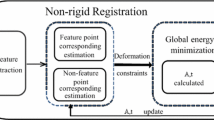
The preoperative steps include constructing a FEM from MR images and extracting vascular tree from MR Angiography (MRA). As the first intraoperative step, after the craniotomy and with the dura opened, a designed checkerboard pattern is projected on the cortex surface and projected landmarks are scanned and captured by a stereo camera (Int J Imaging Syst Technol 23(4):294–303, 2013. doi: 10.1002/ima.22064). This 3D point cloud should be registered to boundary nodes of FEM in the region of interest. For this purpose, we developed a new non-rigid registration method, called finite element drift that is more compatible with the underlying nature of deformed object. The presented algorithm outperforms other methods such as coherent point drift when the deformation is local or non-coherent. After registration, the acquired displacement vectors are used as boundary conditions for FE model. As the second step, by tracking a 2D Doppler ultrasound probe swept on the parenchyma, a 3D image of deformed vascular tree is constructed. Elastic registration of this vascular point cloud to the corresponding preoperative data results the second series of displacement vector applicable to closest internal nodes of FEM. After running FE analysis, the displacement of all nodes is calculated. The brain shift is then estimated as displacement of nodes in boundary of a deep target, e.g., a tumor. We used intraoperative MR (iMR) images as the references for measuring the performance of the brain shift estimator. In the present study, two set of tests were performed using: (a) a deformable brain phantom with surface data and (b) an alive brain of an approximately big dog with surface data and US Doppler images. In our designed phantom, small tubes connected to an inflatable balloon were considered as displaceable targets and in the animal model, the target was modeled by a cyst which was created by an injection.
Results
In the phantom study, the registration error for the surface points before FE analysis and for the target points after running FE model were \({<}0.76\) and 1.4 mm, respectively. In a real condition of operating room for animal model, the registration error was about 1 mm for the surface, 1.9 mm for the vascular tree and 1.55 mm for the target points.
Conclusions
The proposed projected surface imaging in conjunction with the Doppler US data combined in a powerful biomechanical model can result an acceptable performance in calculation of deformation during surgical navigation. However, the projected landmark method is sensitive to ambient light and surface conditions and the Doppler ultrasound suffers from noise and 3D image construction problems, the combination of these two methods applied on a FEM has an eligible performance.









Similar content being viewed by others
Notes
Parseh Intelligent Surgical Systems Parsiss Company, Tehran, Iran. www.parsiss.com.
References
Ahmadian A, Dadashi Serej N, Karimifard S, Farnia P (2013) An efficient method for estimation of soft tissue deformation based on intra-operative stereo image features and point-based registration. Int J Imaging Syst Technol 23(4):294–303. doi:10.1002/ima.22064
Nimsky C, Ganslandt O, Cerny S, Hastreiter P, Greiner G, Fahlbusch R (2000) Quantification of, visualization of, and compensation for brain shift using intraoperative magnetic resonance imaging. Neurosurgery 47(5):1070–1080
Nabavi A, Black P, Gering D, Westin C, Mehta V, Pergolizz R Jr, Ferrant M, Warfield S, Hata N, Schwartz R, Wells W, Kikinis R, Jolesz F (2001) Serial intraoperative magnetic resonance imaging of brain shift. Neurosurgery 48(4):787–797
Reinges M, Nguyen H, Krings T, Hutter B, Rohde V, Gilsbach J (2004) Course of brain shift during microsurgical resection of supratentorial cerebral lesions: limits of conventional neuronavigation. Acta Neurochir 146(4):369–377
Trantakis C, Tittgemeyer M, Schneider J, Lindner D, Winkler D, Strauss G, Meixensberger J (2003) Investigation of time-dependency of intracranial brain shift and its relation to the extent of tumor removal using intra-operative MRI. Neurol Res 25(1):9–12
Soza G, Hastreiter P, Vega F, Rezk-Salama C, Bauer M, Nimsky C, Greiner G (2003) Non-linear intraoperative correction of brain shift with 1.5 T data. Bildverarbeitung für die Medizin. Springer, Berlin. doi:10.1007/978-3-642-18993-7_5
Clatz O, Delingette H, Talos I, Golby A, Kikinis R, Jolesz F, Ayache N, Warfield S (2005) Robust non-rigid registration to capture brain shift from intra-operative MRI. IEEE Trans Med Imaging 24(11):1417–1427
Hu J, Jin X, Lee J, Zhang L, Chaudhary V, Guthikonda M, Yang K, King A (2007) Intraoperative brain shift prediction using a 3D inhomogeneous patient specific finite element model. J Neurosurg 106:164–169
Hata N, Nabavi A, Wells W III, Warfield S, Kikinis R, Black P, Jolesz F (2000) Three-dimensional optical flow method for measurement of volumetric brain deformation from intraoperative MR images. J Comput Assist Tomogr 24(4):531–538
Shattuck D, Leahy R (2002) Brain suite: an automated cortical surface identification tool. Med Image Anal 6(2):129–142
Ferrant M, Nabavi A, Macq B, Black P, Jolesz F, Kikinis R, Warfield S (2002) Serial registration of intraoperative MR images of the brain. Med Image Anal 6(4):337–359
Kyriacou S, Davatzikos C, Zinreich S, Bryan R (1999) Nonlinear elastic registration of brain images with tumor pathology using a biomechanical model. IEEE Trans Med Imaging 18(7):580–592
Miga M, Sinha T, Cash D, Galloway R, Weil R (2003) Cortical surface registration for image-guided neurosurgery using laser-range scanning. IEEE Trans Med Imaging 22(8):973–985
Comeau R, Sadikot A, Fenster A, Peters T (2000) Intraoperative ultrasound for guidance and tissue shift correction in image-guided neurosurgery. Med Phys 27:787–800
Reinertsen I, Lindseth F, Unsqaard G, Collins D (2007) Clinical validation of vessel-based registration for correction of brain-shift. Med Image Anal 11(6):673–684
Bucki M, Palombi O, Bailet M, Payan Y (2012) Doppler ultrasound driven biomedical model of the brain for intraoperative brain-shift compensation: a proof of concept in clinical conditions. In: Payan Y (ed) Soft tissue biomechanical modeling for computer assisted surgery. Studies in mechanobiology, tissue engineering and biomaterials series, vol 11. Springer, pp 135–165. doi:10.1007/978-3-642-29014-5
Hjelmeland A, Lathia J, Sathornsumetee S, Rich J (2011) Twisted tango: brain tumor neurovascular interactions. Nat Neurosci 14(11):1375–1381
Joldes G, Wittek A, Couton M, Warfield S, Miller K (2009) Real-time prediction of brain shift using nonlinear finite element algorithms. In: Yang G, Hawkes D, Rueckert D, Noble A, Taylor C (eds) Medical image computing and computer-assisted intervention (MICCAI 2009). Lecture notes in computer science, vol 5762. pp 300–307. doi:10.1007/978-3-642-04271-3_37
Mendoza C, Laugier C (2003) Tissue Cutting using finite elements and force feedback. In: Ayache N, Delingette H (eds) Surgery simulation and soft tissue modeling. Lecture notes in computer science, vol 2673, pp 175–182. doi:10.1007/3-540-45015-7_17
Dadashi Serej N, Ahmadian A, Mohagheghi S, Sadrehosseini SM (2014) A projected landmark method for reduction of registration error in image-guided surgery systems. Int J Comput Assist Radiol Surg 10(5):541–554. doi:10.1007/s11548-014-1075-z
Farnia P, Ahmadian A, Khoshnevisan A, Jaberzadeh A, Dadashi Serej N, Kazerooni AF (2011) An efficient point based registration of intra-operative ultrasound images with MR images for computation of brain shift; A phantom study. Proceedings of IEEE conference on engineering in medicine and biology (EMBS 2011) pp 8074–8077. doi:10.1109/IEMBS.2011.6091991
Free and open source software package for visualization and medical image computing. Brigham and Women’s Hospital, Harvard Medical School. http://www.slicer.org/
Free software for extracting triangulated iso-surfaces from a block of data. Machine Intelligence Laboratory, Department of Engineering, Cambridge University. http://mi.eng.cam.ac.uk/~gmt11/software/isosurf/isosurf.html
Free software for 3D finite element grid generation. Dept. of Electrical Engineering and Computer Science, University of Liege. http://www.geuz.org/gmsh/
Chui H, Rangarajan A (2003) A new point matching algorithm for non-rigid registration. Comput Vis Image Underst 89(2/3):114–141
Myronenko A, Song X (2010) Point set registration: coherent point drift. IEEE Trans Pattern Anal Machine Intell 32(12):2262–2275
Lewin JS, Nour SG, Meyers ML, Metzger AK, Maciunas RJ, Wendt M, Duerk JL, Oppelt A, Selman WR (2007) Intraoperative MRI with a rotating, tiltable surgical table: a time use study and clinical results in 122 patients. AJR Am J Roentgenol 189(5):1096–1103. doi:10.2214/AJR.06.1247
Acknowledgments
This Project was supported by the Research Center for Biomedical Technology and Robotics (RCBTR), and Rajaei Cardiovascular, Medical and Research Center, and Parseh Intelligent Surgical Systems (Parsiss Company), Tehran, Iran.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mohammadi, A., Ahmadian, A., Azar, A.D. et al. Estimation of intraoperative brain shift by combination of stereovision and doppler ultrasound: phantom and animal model study. Int J CARS 10, 1753–1764 (2015). https://doi.org/10.1007/s11548-015-1216-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-015-1216-z