Abstract
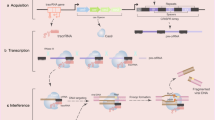
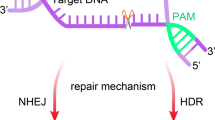
Human pluripotent stem cells have been much anticipated as a powerful system to study developmental events, model genetic disorders, and serve as a source of autologous cells for cell therapy in genetic disorders. Precise genetic manipulation is crucial to all these applications, and many recent advances have been made in site specific nuclease systems like zinc finger nucleases, TALENs, and CRISPR/Cas. In this review, we address the importance of site-specific genome modification and how this technology can be applied to manipulate human pluripotent stem cells.
Similar content being viewed by others
References
Bibikova M, Beumer K, Trautman J K, Carroll D (2003). Enhancing gene targeting with designed zinc finger nucleases. Science, 300(5620): 764
Bibikova M, Golic M, Golic K G, Carroll D (2002). Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics, 161(3): 1169–1175
Bogdanove A J, Voytas D F (2011). TAL effectors: customizable proteins for DNA targeting. Science, 333(6051): 1843–1846
Brunet E, Simsek D, Tomishima M, DeKelver R, Choi V M, Gregory P, Urnov F, Weinstock D M, Jasin M (2009). Chromosomal translocations induced at specified loci in human stem cells. Proc Natl Acad Sci USA, 106(26): 10620–10625
Bultmann S, Morbitzer R, Schmidt C S, Thanisch K, Spada F, Elsaesser J, Lahaye T, Leonhardt H (2012). Targeted transcriptional activation of silent oct4 pluripotency gene by combining designer TALEs and inhibition of epigenetic modifiers. Nucleic Acids Res, 40(12): 5368–5377
Carroll D (2011). Genome engineering with zinc-finger nucleases. Genetics, 188(4): 773–782
Cermak T, Doyle E L, Christian M, Wang L, Zhang Y, Schmidt C, Baller J A, Somia N V, Bogdanove A J, Voytas D F (2011). Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res, 39(12): e82
Chaikind B, Kilambi K P, Gray J J, Ostermeier M (2012). Targeted DNA methylation using an artificially bisected M.HhaI fused to zinc fingers. PLoS ONE, 7(9): e44852
Chang C J, Bouhassira E E (2012). Zinc-finger nuclease-mediated correction of α-thalassemia in iPS cells. Blood, 120(19): 3906–3914
Cho S W, Kim S, Kim J M, Kim J S (2013). Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat Biotechnol, 31(3): 230–232
Christian M, Cermak T, Doyle E L, Schmidt C, Zhang F, Hummel A, Bogdanove A J, Voytas D F (2010). Targeting DNA double-strand breaks with TAL effector nucleases. Genetics, 186(2): 757–761
Cong L, Ran F A, Cox D, Lin S, Barretto R, Habib N, Hsu P D, Wu X, Jiang W, Marraffini L A, Zhang F (2013). Multiplex genome engineering using CRISPR/Cas systems. Science, 339(6121): 819–823
Ding Q, Lee Y K, Schaefer E A, Peters D T, Veres A, Kim K, Kuperwasser N, Motola D L, Meissner T B, Hendriks W T, Trevisan M, Gupta R M, Moisan A, Banks E, Friesen M, Schinzel R T, Xia F, Tang A, Xia Y, Figueroa E, Wann A, Ahfeldt T, Daheron L, Zhang F, Rubin L L, Peng L F, Chung R T, Musunuru K, Cowan C A (2013). A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell, 12(2): 238–251
Doetschman T, Gregg R G, Maeda N, Hooper M L, Melton D W, Thompson S, Smithies O (1987). Targetted correction of a mutant HPRT gene in mouse embryonic stem cells. Nature, 330(6148): 576–578
Doyle J P, Dougherty J D, Heiman M, Schmidt E F, Stevens T R, Ma G, Bupp S, Shrestha P, Shah R D, Doughty M L, Gong S, Greengard P, Heintz N (2008). Application of a translational profiling approach for the comparative analysis of CNS cell types. Cell, 135(4): 749–762
Durai S, Mani M, Kandavelou K, Wu J, Porteus M H, Chandrasegaran S (2005). Zinc finger nucleases: custom-designed molecular scissors for genome engineering of plant and mammalian cells. Nucleic Acids Res, 33(18): 5978–5990
Heiman M, Schaefer A, Gong S, Peterson J D, Day M, Ramsey K E, Suárez-Fariñas M, Schwarz C, Stephan D A, Surmeier D J, Greengard P, Heintz N (2008). A translational profiling approach for the molecular characterization of CNS cell types. Cell, 135(4): 738–748
Hockemeyer D, Soldner F, Beard C, Gao Q, Mitalipova M, DeKelver R C, Katibah G E, Amora R, Boydston E A, Zeitler B, Meng X, Miller J C, Zhang L, Rebar E J, Gregory P D, Urnov F D, Jaenisch R (2009). Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat Biotechnol, 27(9): 851–857
Hockemeyer D, Wang H, Kiani S, Lai C S, Gao Q, Cassady J P, Cost G J, Zhang L, Santiago Y, Miller J C, Zeitler B, Cherone J M, Meng X, Hinkley S J, Rebar E J, Gregory P D, Urnov F D, Jaenisch R (2011). Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol, 29(8): 731–734
Hwang W Y, Fu Y, Reyon D, Maeder M L, Tsai S Q, Sander J D, Peterson R T, Yeh J R, Joung J K (2013). Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol, 31(3): 227–229
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna J A, Charpentier E (2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science, 337(6096): 816–821
Kim Y, Kweon J, Kim A, Chon J K, Yoo J Y, Kim H J, Kim S, Lee C, Jeong E, Chung E, Kim D, Lee M S, Go E M, Song H J, Kim H, Cho N, Bang D, Kim S, Kim J S (2013). A library of TAL effector nucleases spanning the human genome. Nat Biotechnol, 31(3): 251–258
Lengner C J, Gimelbrant A A, Erwin J A, Cheng A W, Guenther M G, Welstead G G, Alagappan R, Frampton G M, Xu P, Muffat J, Santagata S, Powers D, Barrett C B, Young R A, Lee J T, Jaenisch R, Mitalipova M (2010). Derivation of pre-X inactivation human embryonic stem cells under physiological oxygen concentrations. Cell, 141(5): 872–883
Liu H, Ye Z, Kim Y, Sharkis S, Jang Y Y (2010). Generation of endoderm-derived human induced pluripotent stem cells from primary hepatocytes. Hepatology, 51(5): 1810–1819
Maeder M L, Thibodeau-Beganny S, Osiak A, Wright D A, Anthony R M, Eichtinger M, Jiang T, Foley J E, Winfrey R J, Townsend J A, Unger-Wallace E, Sander J D, Müller-Lerch F, Fu F, Pearlberg J, Göbel C, Dassie J P, Pruett-Miller S M, Porteus M H, Sgroi D C, Iafrate A J, Dobbs D, McCray P B Jr, Cathomen T, Voytas D F, Joung J K (2008). Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol Cell, 31(2): 294–301
Mali P, Aach J, Lee J H, Levner D, Nip L, Church G M (2013). Barcoding cells using cell-surface programmable DNA-binding domains. Nat Methods, 10(5): 403–406
Mali P, Yang L, Esvelt K M, Aach J, Guell M, DiCarlo J E, Norville J E, Church G M (2013). RNA-guided human genome engineering via Cas9. Science, 339(6121): 823–826
Miller J C, Tan S, Qiao G, Barlow K A, Wang J, Xia D F, Meng X, Paschon D E, Leung E, Hinkley S J, Dulay G P, Hua K L, Ankoudinova I, Cost G J, Urnov F D, Zhang H S, Holmes M C, Zhang L, Gregory P D, Rebar E J (2011). A TALE nuclease architecture for efficient genome editing. Nat Biotechnol, 29(2): 143–148
Mussolino C, Cathomen T (2011). On target? Tracing zinc-fingernuclease specificity. Nat Methods, 8(9): 725–726
Nishiyama A, Xin L, Sharov A A, Thomas M, Mowrer G, Meyers E, Piao Y, Mehta S, Yee S, Nakatake Y, Stagg C, Sharova L, Correa-Cerro L S, Bassey U, Hoang H, Kim E, Tapnio R, Qian Y, Dudekula D, Zalzman M, Li M, Falco G, Yang H T, Lee S L, Monti M, Stanghellini I, Islam M N, Nagaraja R, Goldberg I, Wang W, Longo D L, Schlessinger D, Ko M S (2009). Uncovering early response of gene regulatory networks in ESCs by systematic induction of transcription factors. Cell Stem Cell, 5(4): 420–433
Piganeau M, Ghezraoui H, De Cian A, Guittat L, Tomishima M, Perrouault L, René O, Katibah G E, Zhang L, Holmes M C, Doyon Y, Concordet J P, Giovannangeli C, Jasin M, Brunet E (2013). Cancer translocations in human cells induced by zinc finger and TALE nucleases. Genome Res, 23(7): 1182–1193
Porteus M H, Baltimore D (2003). Chimeric nucleases stimulate gene targeting in human cells. Science, 300(5620): 763
Qi L S, Larson MH, Gilbert L A, Doudna J A, Weissman J S, Arkin A P, Lim W A (2013). Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 152(5): 1173–1183
Sermon K D, Simon C, Braude P, Viville S, Borstlap J, Veiga A (2009). Creation of a registry for human embryonic stem cells carrying an inherited defect: joint collaboration between ESHRE and hESCreg. Hum Reprod, 24(7): 1556–1560
Smih F, Rouet P, Romanienko P J, Jasin M (1995). Double-strand breaks at the target locus stimulate gene targeting in embryonic stem cells. Nucleic Acids Res, 23(24): 5012–5019
Soldner F, Laganière J, Cheng A W, Hockemeyer D, Gao Q, Alagappan R, Khurana V, Golbe L I, Myers R H, Lindquist S, Zhang L, Guschin D, Fong L K, Vu B J, Meng X, Urnov F D, Rebar E J, Gregory P D, Zhang H S, Jaenisch R (2011). Generation of isogenic pluripotent stem cells differing exclusively at two early onset Parkinson point mutations. Cell, 146(2): 318–331
Stephenson E L, Mason C, Braude P R (2009). Preimplantation genetic diagnosis as a source of human embryonic stem cells for disease research and drug discovery. BJOG, 116(2): 158–165
Sussman D, Chadsey M, Fauce S, Engel A, Bruett A, Monnat R Jr, Stoddard B L, Seligman LM(2004). Isolation and characterization of new homing endonuclease specificities at individual target site positions. J Mol Biol, 342(1): 31–41
Tachibana M, Amato P, Sparman M, Gutierrez N M, Tippner-Hedges R, Ma H, Kang E, Fulati A, Lee H S, Sritanaudomchai H, Masterson K, Larson J, Eaton D, Sadler-Fredd K, Battaglia D, Lee D, Wu D, Jensen J, Patton P, Gokhale S, Stouffer R L, Wolf D, Mitalipov S (2013). Human embryonic stem cells derived by somatic cell nuclear transfer. Cell, 153(6): 1228–1238
Tomishima M J, Hadjantonakis A K, Gong S, Studer L (2007). Production of green fluorescent protein transgenic embryonic stem cells using the GENSAT bacterial artificial chromosome library. Stem Cells, 25(1): 39–45
Urnov F D, Rebar E J, Holmes M C, Zhang H S, Gregory P D (2010). Genome editing with engineered zinc finger nucleases. Nat Rev Genet, 11(9): 636–646
Wang H, Yang H, Shivalila C S, Dawlaty M M, Cheng A W, Zhang F, Jaenisch R (2013). One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell, 153(4): 910–918
Wang Y, Zheng C G, Jiang Y, Zhang J, Chen J, Yao C, Zhao Q, Liu S, Chen K, Du J, Yang Z, Gao S (2012). Genetic correction of β-thalassemia patient-specific iPS cells and its use in improving hemoglobin production in irradiated SCID mice. Cell Res, 22(4): 637–648
Xiao A, Wang Z, Hu Y, Wu Y, Luo Z, Yang Z, Zu Y, Li W, Huang P, Tong X, Zhu Z, Lin S, Zhang B (2013). Chromosomal deletions and inversions mediated by TALENs and CRISPR/Cas in zebrafish. Nucleic Acids Res, doi: 10.1093/nar/gkt464
Yamanaka S, Blau H M (2010). Nuclear reprogramming to a pluripotent state by three approaches. Nature, 465(7299): 704–712
Yu J, Thomson J A (2008). Pluripotent stem cell lines. Genes Dev, 22(15): 1987–1997
Yusa K, Rashid S T, Strick-Marchand H, Varela I, Liu P Q, Paschon D E, Miranda E, Ordóñez A, Hannan N R, Rouhani F J, Darche S, Alexander G, Marciniak S J, Fusaki N, Hasegawa M, Holmes M C, Di Santo J P, Lomas D A, Bradley A, Vallier L (2011). Targeted gene correction of α1-antitrypsin deficiency in induced pluripotent stem cells. Nature, 478(7369): 391–394
Zou J, Maeder M L, Mali P, Pruett-Miller S M, Thibodeau-Beganny S, Chou B K, Chen G, Ye Z, Park I H, Daley G Q, Porteus MH, Joung J K, Cheng L (2009). Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell, 5(1): 97–110
Zou J, Mali P, Huang X, Dowey S N, Cheng L (2011). Site-specific gene correction of a point mutation in human iPS cells derived from an adult patient with sickle cell disease. Blood, 118(17): 4599–4608
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mukherjee-Clavin, B., Tomishima, M. & Lee, G. Current approaches for efficient genetic editing in human pluripotent stem cells. Front. Biol. 8, 461–467 (2013). https://doi.org/10.1007/s11515-013-1275-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11515-013-1275-x