Abstract
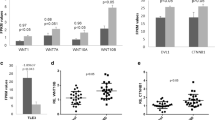
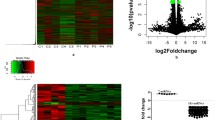
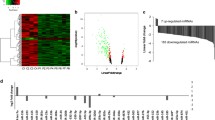
While Post Traumatic Stress Disorder (PTSD) is associated with immune dysfunction, the underlying mechanisms remain unclear. Studies suggest a role for involvement of epigenetic mechanisms and microRNAs (miRNAs). Here, we examined genome-wide histone and DNA methylation in the peripheral blood mononuclear cells (PBMCs) in PTSD. We noted significant differences in histone H3 trimethylation at K4, K9, K27 and K36 sites in PTSD when compared to control. While overall DNA methylation level did not differ significantly between control and PTSD, the promoters of several individual genes (e.g., Interferon gamma (IFNG) and Interleukin (IL)-12B) were differentially methylated. ChIP-seq data revealed that the promoter of IFNG and TBX-21 was associated with the activation marker H3K4me3 in PTSD. The transcript levels of both IFNG and TBX-21 were higher in PTSD correlating well with the altered methylation patterns. Furthermore, PTSD patients showed increased expression of IL-12 in their PBMCs. Analysis of both histone and DNA methylation markers suggested that the expression of IL-12 was also possibly activated through epigenetic modification. Knockdown of lysine (K)-specific demethylase 5B (KDM5B), or inhibition of DNA (Cytosine-5-)-methyltransferase 1 (DNMT1) caused up-regulation of IL-12. Furthermore, the expression of these cytokines was also regulated by miRNAs. Our miRNA microarray identified many downregulated miRNAs in PTSD that are predicted to target IFNG and IL-12. Consequently, we showed that up-regulation of hsa-miR-193a-5p could decrease the expression of IL-12. Overall, the current study demonstrated that the elevated expression of pro-inflammatory cytokines in PTSD patients might be regulated by multiple epigenetic mechanisms and miRNAs.






Similar content being viewed by others
References
Ambros V (2004) The functions of animal microRNAs. Nature 431(7006):350–355. doi:10.1038/nature02871
Bannister AJ, Schneider R, Myers FA, Thorne AW, Crane-Robinson C, Kouzarides T (2005) Spatial distribution of di- and tri-methyl lysine 36 of histone H3 at active genes. J Biol Chem 280:17732–17736. doi:10.1074/jbc.M500796200
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297. doi:10.1016/S0092-8674(04)00045-5
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, Fry B, Meissner A, Wernig M, Plath K, Jaenisch R, Wagschal A, Feil R, Schreiber SL, Lander ES (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125:315–326. doi:10.1016/j.cell.2006.02.041
Blankenberg D, Von Kuster G, Coraor N, Ananda G, Lazarus R, Mangan M, Nekrutenko A, Taylor J (2010) Galaxy: a web-based genome analysis tool for experimentalists. Curr Protoc Mol Biol Chapter 19: Unit 19 10 11–21
Christman JK (2002) 5-Azacytidine and 5-aza-2′-deoxycytidine as inhibitors of DNA methylation: mechanistic studies and their implications for cancer therapy. Oncogene 21(35):5483–5495. doi:10.1038/sj.onc.1205699
Collier SP, Henderson MA, Tossberg JT, Aune TM (2014) Regulation of the Th1 genomic locus from Ifng through Tmevpg1 by T-bet. J Immunol 193(8):3959–3965. doi:10.4049/jimmunol.1401099
Djuretic IM, Levanon D, Negreanu V, Groner Y, Rao A, Ansel KM (2007) Transcription factors T-bet and Runx3 cooperate to activate Ifng and silence Il4 in T helper type 1 cells. Nat Immunol 8(2):145–153. doi:10.1038/ni1424
Flintoft L (2013) Non-coding RNA: structure and function for lncRNAs. Nat Rev Genet 14:598. doi:10.1038/nrg3561
Frayne SM, Chiu VY, Iqbal S, Berg EA, Laungani KJ, Cronkite RC, Pavao J, Kimerling R (2011) Medical care needs of returning veterans with PTSD: their other burden. J Gen Intern Med 26(1):33–39. doi:10.1007/s11606-010-1497-4
Gill JM, Saligan L, Woods S, Page G (2009) PTSD is associated with an excess of inflammatory immune activities. Perspect Psychiatr Care 45:262–277. doi:10.1111/j.1744-6163.2009.00229.x
Greer EL, Shi Y (2012) Histone methylation: a dynamic mark in health, disease and inheritance. Nat Rev Genet 13:343–357. doi:10.1038/nrg3173
He L, Hannon GJ (2004) MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531. doi:10.1038/nrg1379
Hoge EA, Brandstetter K, Moshier S, Pollack MH, Wong KK, Simon NM (2009) Broad spectrum of cytokine abnormalities in panic disorder and posttraumatic stress disorder. Depress Anxiet 26:447–455. doi:10.1002/da.20564
Hunter RG, McCarthy KJ, Milne TA, Pfaff DW, McEwen BS (2009) Regulation of hippocampal H3 histone methylation by acute and chronic stress. Proc Natl Acad Sci U S A 106:20912–20917. doi:10.1073/pnas.0911143106
Kessler RC, Chiu WT, Demler O, Merikangas KR, Walters EE (2005) Prevalence, severity, and comorbidity of 12-month DSM-IV disorders in the National Comorbidity Survey Replication. Arch Gen Psychiatry 62:617–627. doi:10.1001/archpsyc.62.6.617
Klein BJ, Piao L, Xi Y, Rincon-Arano H, Rothbart SB, Peng D, Wen H, Larson C, Zhang X, Zheng X, Cortazar MA, Peña PV, Mangan A, Bentley DL, Strahl BD, Groudine M, Li W, Shi X, Kutateladze TG (2014) The histone-H3K4-specific demethylase KDM5B binds to its substrate and product through distinct PHD fingers. Cell Rep 6(2):325–335. doi:10.1016/j.celrep.2013.12.021
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10(3):R25. doi:10.1186/gb-2009-10-3-r25
Lehnertz B, Ueda Y, Derijck AA, Braunschweig U, Perez-Burgos L, Kubicek S, Chen T, Li E, Jenuwein T, Peters AH (2003) Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr Biol 13:1192–1200. doi:10.1016/S0960-9822(03)00432-9
Li LC, Dahiya R (2002) MethPrimer: designing primers for methylation PCRs. Bioinformatics 18(11):1427–1431. doi:10.1093/bioinformatics/18.11.1427
Lienhard M, Grimm C, Morkel M, Herwig R, Chavez L (2014) MEDIPS: genome-wide differential coverage analysis of sequencing data derived from DNA enrichment experiments. Bioinformatics 30:284–286. doi:10.1093/bioinformatics/btt650
Mikkelsen TS, Ku M, Jaffe DB, Issac B, Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP, Lee W, Mendenhall E, O’Donovan A, Presser A, Russ C, Xie X, Meissner A, Wernig M, Jaenisch R, Nusbaum C, Lander ES, Bernstein BE (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448:553–560. doi:10.1038/nature06008
Muers M (2013) Gene expression: disentangling DNA methylation. Nat Rev Genet 14:519. doi:10.1038/nrg3535
Puddu P, Fantuzzi L, Borghi P, Varano B, Rainaldi G, Guillemard E, Malorni W, Nicaise P, Wolf SF, Belardelli F, Gessani S (1997) IL-12 induces IFN-gamma expression and secretion in mouse peritoneal macrophages. J Immunol 159:3490–3497
Roh TY, Cuddapah S, Cui K, Zhao K (2006) The genomic landscape of histone modifications in human T cells. Proc Natl Acad Sci U S A 103:15782–15787. doi:10.1073/pnas.0607617103
Roh TY, Wei G, Farrell CM, Zhao K (2007) Genome-wide prediction of conserved and nonconserved enhancers by histone acetylation patterns. Genome Res 17:74–81. doi:10.1101/gr.5767907
Rohleder N, Karl A (2006) Role of endocrine and inflammatory alterations in comorbid somatic diseases of post-traumatic stress disorder. Minerva Endocrinol 31:273–288
Ross-Innes CS, Stark R, Teschendorff AE, Holmes KA, Ali HR, Dunning MJ, Brown GD, Gojis O, Ellis IO, Green AR, Ali S, Chin SF, Palmieri C, Caldas C, Carroll JS (2012) Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481:389–393. doi:10.1038/nature10730
Rothbart SB, Strahl BD (2014) Interpreting the language of histone and DNA modifications. Biochim Biophys Acta 1839(8):627–643. doi:10.1016/j.bbagrm.2014.03.001
Rusiecki JA, Byrne C, Galdzicki Z, Srikantan V, Chen L, Poulin M, Yan L, Baccarelli A (2013) PTSD and DNA methylation in select immune function gene promoter regions: a repeated measures case–control study of U.S. Military Service Members. Front Psychiatr 4:56. doi:10.3389/fpsyt.2013.00056
Schroder K, Hertzog PJ, Ravasi T, Hume DA (2004) Interferon-gamma: an overview of signals, mechanisms and functions. J Leukoc Biol 75(2):163–189. doi:10.1189/jlb.0603252
Shin H, Liu T, Manrai AK, Liu XS (2009) CEAS: cis-regulatory element annotation system. Bioinformatics 25:2605–2606. doi:10.1093/bioinformatics/btp479
Smith AK, Conneely KN, Kilaru V, Mercer KB, Weiss TE, Bradley B, Tang Y, Gillespie CF, Cubells JF, Ressler KJ (2011) Differential immune system DNA methylation and cytokine regulation in post-traumatic stress disorder. Am J Med Genet B Neuropsychiatr Genet 156B:700–708. doi:10.1002/ajmg.b.31212
Sun H, Kennedy PJ, Nestler EJ (2013) Epigenetics of the depressed brain: role of histone acetylation and methylation. Neuropsychopharmacology 38:124–137. doi:10.1038/npp.2012.73
Sutherland AG, Alexander DA, Hutchison JD (2003) Disturbance of pro-inflammatory cytokines in post-traumatic psychopathology. Cytokine 24:219–225. doi:10.1016/j.cyto.2003.09.004
Thomas JL, Wilk JE, Riviere LA, McGurk D, Castro CA, Hoge CW (2010) Prevalence of mental health problems and functional impairment among active component and National Guard soldiers 3 and 12 months following combat in Iraq. Arch Gen Psychiatry 67:614–623. doi:10.1001/archgenpsychiatry.2010.54
Uddin M, Aiello AE, Wildman DE, Koenen KC, Pawelec G, de Los Santos R, Goldmann E, Galea S (2010) Epigenetic and immune function profiles associated with posttraumatic stress disorder. Proc Natl Acad Sci U S A 107:9470–9475. doi:10.1073/pnas.0910794107
Vakoc CR, Mandat SA, Olenchock BA, Blobel GA (2005) Histone H3 lysine 9 methylation and HP1gamma are associated with transcription elongation through mammalian chromatin. Mol Cell 19:381–391. doi:10.1016/j.molcel.2005.06.011
Vire E, Brenner C, Deplus R, Blanchon L, Fraga M, Didelot C, Morey L, Van Eynde A, Bernard D, Vanderwinden JM, Bollen M, Esteller M, Di Croce L, de Launoit Y, Fuks F (2006) The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439:871–874. doi:10.1038/nature04431
von Kanel R, Hepp U, Kraemer B, Traber R, Keel M, Mica L, Schnyder U (2007) Evidence for low-grade systemic proinflammatory activity in patients with posttraumatic stress disorder. J Psychiatr Res 41:744–752. doi:10.1016/j.jpsychires.2006.06.009
Wei G, Wei L, Zhu J, Zang C, Hu-Li J, Yao Z, Cui K, Kanno Y, Roh TY, Watford WT, Schones DE, Peng W, Sun HW, Paul WE, O’Shea JJ, Zhao K (2009) Global mapping of H3K4me3 and H3K27me3 reveals specificity and plasticity in lineage fate determination of differentiating CD4+ T cells. Immunity 30:155–167. doi:10.1016/j.immuni.2008.12.009
Zang C, Schones DE, Zeng C, Cui K, Zhao K, Peng W (2009) A clustering approach for identification of enriched domains from histone modification ChIP-Seq data. Bioinformatics 25:1952–1958. doi:10.1093/bioinformatics/btp340
Zhou VW, Goren A, Bernstein BE (2011) Charting histone modifications and the functional organization of mammalian genomes. Nat Rev Genet 12:7–18. doi:10.1038/nrg2905
Zhou J, Nagarkatti P, Zhong Y, Ginsberg JP, Singh NP, Zhang J, Nagarkatti M (2014) Dysregulation in microRNA expression is associated with alterations in immune functions in combat veterans with post-traumatic stress disorder. PLoS One 9:e94075. doi:10.1371/journal.pone.0094075. eCollection 2014
Acknowledgments
This study is supported in part by National Institutes of Health grants P01AT003961, R01AT006888, R01ES019313, R01MH094755, and P20GM103641 as well as by Veterans Affairs Merit Award BX001357 to PN and MN.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical Approval
All procedures performed in studies involving human participants were approved by the Institutional Review Board.
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Marpe Bam and Xiaoming Yang contributed equally to this work.
Rights and permissions
About this article
Cite this article
Bam, M., Yang, X., Zhou, J. et al. Evidence for Epigenetic Regulation of Pro-Inflammatory Cytokines, Interleukin-12 and Interferon Gamma, in Peripheral Blood Mononuclear Cells from PTSD Patients. J Neuroimmune Pharmacol 11, 168–181 (2016). https://doi.org/10.1007/s11481-015-9643-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11481-015-9643-8