Abstract
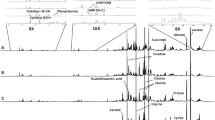
Melanoma is a malignant tumor of melanocytes. Although extensive investigations have been done to study metabolic changes in primary melanoma in vivo and in vitro, little effort has been devoted to metabolic profiling of metastatic tumors in organs other than lymph nodes. In this work, NMR-based metabolomics combined with multivariate data analysis is used to study metastatic B16-F10 melanoma in C57BL/6J mouse spleen. Principal component analysis, an unsupervised multivariate data analysis method, is used to detect possible outliers, while orthogonal projection to latent structure (OPLS), a supervised multivariate data analysis method, is employed to find important metabolites responsible for discriminating the control and the melanoma groups. Two different strategies, i.e. spectral binning and spectral deconvolution, are used to reduce the original spectral data before statistical analysis. Spectral deconvolution is found to be superior for identifying a set of discriminatory metabolites between the control and the melanoma groups, especially when the sample size is small. OPLS results show that the melanoma group can be well separated from its control group. It is found that taurine, glutamate, aspartate, O-phosphoethanolamine, niacinamide, ATP, lipids and glycerol derivatives are decreased statistically and significantly while alanine, malate, xanthine, histamine, dCTP, GTP, thymidine, 2′-deoxyguanosine are statistically and significantly elevated. These significantly changed metabolites are associated with multiple biological pathways and may be potential biomarkers for metastatic melanoma in spleen.




Similar content being viewed by others
References
Abaffy, T., Moller, M. G., Riemer, D. D., Milikowski, C., & DeFazio, R. A. (2013). Comparative analysis of volatile metabolomics signals from melanoma and benign skin: a pilot study. Metabolomics, 9(5), 998–1008. doi:10.1007/s11306-013-0523-z.
Akoh, C. C., & Min, D. B. (2008). Food Lipids: Chemistry, Nutrition, and Biotechnology (Food Science and Technology) (3rd ed.). Boca Raton: CRC Press.
Ali, K., Iqbal, M., Yuliana, N. D., Lee, Y. J., Park, S., Han, S., et al. (2013). Identification of bioactive metabolites against adenosine A1 receptor using NMR-based metabolomics. Metabolomics, 9(4), 778–785. doi:10.1007/s11306-013-0498-9.
Alizadeh, H., Howard, K., Mellon, J., Mayhew, E., Rusciano, D., & Niederkorn, J. Y. (2003). Reduction of liver metastasis of intraocular melanoma by interferon-beta gene transfer. Investigative Ophthalmology and Visual Science, 44(7), 3042–3051. doi:10.1167/Iovs.02-1147.
An, Y. P., Xu, W. X., Li, H. H., Lei, H. H., Zhang, L. M., Hao, F. H., et al. (2013). High-fat diet induces dynamic metabolic alterations in multiple biological matrices of rats. Journal of Proteome Research, 12(8), 3755–3768. doi:10.1021/Pr400398b.
Bailet, O., Fenouille, N., Abbe, P., Robert, G., Rocchi, S., Gonthier, N., et al. (2009). Spleen tyrosine kinase functions as a tumor suppressor in melanoma cells by inducing senescence-like growth arrest. Cancer Research, 69(7), 2748–2756.
Beckonert, O., Keun, H. C., Ebbels, T. M. D., Bundy, J. G., Holmes, E., Lindon, J. C., et al. (2007). Metabolic profiling, metabolomic and metabonomic procedures for NMR spectroscopy of urine, plasma, serum and tissue extracts. Nature Protocols, 2(11), 2692–2703.
Bernardo, S. G., Moskalenko, M., Pan, M., Shah, S., Sidhu, H. K., Sicular, S., et al. (2013). Elevated rates of transaminitis during ipilimumab therapy for metastatic melanoma. Melanoma Research, 23(1), 47–54. doi:10.1097/Cmr.0b013e32835c7e68.
Bourne, R. M., Stanwell, P., Stretch, J. R., Scolyer, R. A., Thompson, J. F., Mountford, C. E., et al. (2005). In vivo and ex vivo proton MR spectroscopy of primary and secondary melanoma. European Journal of Radiology, 53(3), 506–513.
Chiu, K. P., Ariyaratne, P., Xu, H., Tan, A., Ng, P., Liu, E. T. B., et al. (2007). Pathway aberrations of murine melanoma cells observed in paired-end diTag transcriptomes. BMC Cancer,. doi:10.1186/1471-2407-7-109.
Choi, K. Y., Chang, K., Pickel, J. M., Badger, J. D., & Roche, K. W. (2011). Expression of the metabotropic glutamate receptor 5 (mGluR5) induces melanoma in transgenic mice. Proceedings of the National Academy of Sciences of the USA, 108(37), 15219–15224. doi:10.1073/pnas.1107304108.
Cloarec, O., Dumas, M. E., Trygg, J., Craig, A., Barton, R. H., Lindon, J. C., et al. (2005). Evaluation of the orthogonal projection on latent structure model limitations caused by chemical shift variability and improved visualization of biomarker changes in H-1 NMR spectroscopic metabonomic studies. Analytical Chemistry, 77(2), 517–526.
Constantinou, M. A., Papakonstantinou, E., Spraul, M., Sevastiadou, S., Costalos, C., Koupparis, M. A., et al. (2005). H-1 NMR-based metabonomics for the diagnosis of inborn errors of metabolism in urine. Analytica Chimica Acta, 542(2), 169–177. doi:10.1016/j.aca.2005.03.059.
Craig, A., Cloareo, O., Holmes, E., Nicholson, J. K., & Lindon, J. C. (2006). Scaling and normalization effects in NMR spectroscopic metabonomic data sets. Analytical Chemistry, 78(7), 2262–2267.
Cybulska-Stopa, B., Skoczek, M., Ziobro, M., Switaj, T., Falkowski, S., Morysinski, T., et al. (2012). Results of systemic treatment of cutaneous melanoma in inoperable stage III and IV. Wspolczesna Onkologia (Contemporary Oncology), 16(6), 532–545. doi:10.5114/wo.2012.32487.
Davis, S. C., Clark, S., Hayes, J. R., Green, T. L., & Gruetter, C. A. (2011). Up-regulation of histidine decarboxylase expression and histamine content in B16F10 murine melanoma cells. Inflammation Research, 60(1), 55–61. doi:10.1007/s00011-010-0234-0.
Dong, F. C., Zhang, L. L., Hao, F. H., Tang, H. R., & Wang, Y. L. (2013). Systemic responses of mice to dextran sulfate sodium-induced acute ulcerative colitis using H-1 NMR spectroscopy. Journal of Proteome Research, 12(6), 2958–2966. doi:10.1021/Pr4002383.
Enot, D. P., & Draper, J. (2007). Statistical measures for validating plant genotype similarity assessments following multivariate analysis of metabolome fingerprint data. Metabolomics, 3(3), 349–355. doi:10.1007/s11306-007-0066-2.
Eriksson, L., Trygg, J., & Wold, S. (2008). CV-ANOVA for significance testing of PLS and OPLS (R) models. Journal of Chemometrics, 22(11–12), 594–600. doi:10.1002/Cem.1187.
Fages, A., Morvan, D., Schwartz, L., Steyaert, J. M., Stepien, G., & Demidem, A. (2010). Disturbance of metabolic pathways of glucose consumption by CENU treatment in B16 melanoma tumors: a NMR spectroscopy-based [1,2-13C]glucose fluxomics. Bulletin du Cancer, 97, S39–S40.
Feng, J. I., Isern, N. G., Burton, S. D., & Hu, J. Z. (2013). Studies of secondary melanoma on C57BL/6J mouse liver using 1H NMR metabolomics. Metabolites, 3(4), 1011–1035.
Filipp, F. V., Ratnikov, B., De Ingeniis, J., Smith, J. W., Osterman, A. L., & Scott, D. A. (2012). Glutamine-fueled mitochondrial metabolism is decoupled from glycolysis in melanoma. Pigment Cell and Melanoma Research, 25(6), 732–739. doi:10.1111/Pcmr.12000.
Frolkis, A., Knox, C., Lim, E., Jewison, T., Law, V., Hau, D. D., et al. (2010). SMPDB: The small molecule pathway database. Nucleic Acids Research, 38, D480–D487. doi:10.1093/Nar/Gkp1002.
Gheorgheosu, D., Dehelean, C., Cristea, M., & Muntean, D. (2011). Development of the B16 murine melanoma model. Annals of RSCB, 2, 148–152.
Gruetter, C. A., Davis, S. C., Clark, S. L., McKee, K. R., & Green, T. L. (2008). Comparative expression of histidine decarboxylase (HDC) protein in B16F10 melanoma cells and nontumorigenic melan-A melanocytes. FASEB Journal, 22, 898.
Gruetter, C. A., Davis, S. C., Hayes, J. R., Clark, S. L., & Green, T. L. (2009). Comparison of histamine content in B16F10 melanoma cells and non-tumorigenic melan-A melanocytes. FASEB Journal, 16(1), 105.
Guitera, P., Bourgeat, P., Stretch, J. R., Scolyer, R. A., Ourselin, S., Lean, C., et al. (2010). Diagnostic value of 8.5 T magnetic resonance spectroscopy of benign and malignant skin lesion biopsies. Melanoma Research, 20(4), 311–317.
Hatse, S., De Clercq, E., & Balzarini, J. (1999). Role of antimetabolites of purine and pyrimidine nucleotide metabolism in tumor cell differentiation. Biochemical Pharmacology, 58(4), 539–555. doi:10.1016/S0006-2952(99)00035-0.
Huang, R. Z., Gao, H. C., Ma, L. H., Wang, X., Jia, J. M., Wang, M. J., et al. (2014). Dynamic H-1 NMR-based extracellular metabonomic analysis of oligodendroglia cells infected with herpes simplex virus type 1. Metabolomics, 10(1), 33–41. doi:10.1007/s11306-013-0548-3.
Itzhaki, O., Greenberg, E., Shalmon, B., Kubi, A., Treves, A. J., Shapira-Frommer, R., et al. (2013). Nicotinamide inhibits vasculogenic mimicry, an alternative vascularization pathway observed in highly aggressive melanoma. PLoS ONE,. doi:10.1371/journal.pone.0057160.
Iverson, S. J., Lang, S. L. C., & Cooper, M. H. (2001). Comparison of the Bligh and Dyer and Folch methods for total lipid determination in a broad range of marine tissue. Lipids, 36(11), 1283–1287.
Jacobson, E. L., & Jacobson, M. K. (1993). A biomarker for the assessment of niacin nutriture as a potential preventive factor in carcinogenesis. Journal of Internal Medicine, 233(1), 59–62.
Jiang, L. M., Huang, J., Wang, Y. L., & Tang, H. R. (2012). Metabonomic analysis reveals the CCl4-induced systems alterations for multiple rat organs. Journal of Proteome Research, 11(7), 3848–3859.
Keun, H. C., Ebbels, T. M. D., Antti, H., Bollard, M. E., Beckonert, O., Schlotterbeck, G., et al. (2002). Analytical reproducibility in H-1 NMR-based metabonomic urinalysis. Chemical Research in Toxicology, 15(11), 1380–1386. doi:10.1021/Tx0255774.
Kizaki, H., Williams, J. C., Morris, H. P., & Weber, G. (1980). Increased cytidine 5′-triphosphate synthetase activity in rat and human tumors. Cancer Research, 40(11), 3921–3927.
Kristensen, M., Savorani, F., Ravn-Haren, G., Poulsen, M., Markowski, J., Larsen, F. H., et al. (2010). NMR and interval PLS as reliable methods for determination of cholesterol in rodent lipoprotein fractions. Metabolomics, 6(1), 129–136. doi:10.1007/s11306-009-0181-3.
Lee, H. J., Wall, B. A., Wangari-Talbot, J., Shin, S. S., Rosenberg, S., Chan, J. L. K., et al. (2011). Glutamatergic pathway targeting in melanoma: Single-agent and combinatorial therapies. Clinical Cancer Research, 17(22), 7080–7092.
Li, W., Slominski, R., & Slominski, A. T. (2009). High-resolution magic angle spinning nuclear magnetic resonance analysis of metabolic changes in melanoma cells after induction of melanogenesis. Analytical Biochemistry, 386(2), 282–284.
Lindon, J. C., Nicholson, J. K., & Everett, J. R. (1999). NMR spectroscopy of biofluids. Annual Reports on NMR Spectroscopy, 38(38), 1–88.
Lu, Y. H., Wang, C. M., Chen, Z. X., Zhao, H., Chen, J. Y., Liu, X. B., et al. (2012). Serum metabolomics for the diagnosis and classification of myasthenia gravis. Metabolomics, 8(4), 704–713. doi:10.1007/s11306-011-0364-6.
Maiese, K., Chong, Z. Z., Hou, J. L., & Shang, Y. C. (2009). The vitamin nicotinamide: Translating nutrition into clinical care. Molecules, 14(9), 3446–3485. doi:10.3390/molecules14093446.
Maire, C., Vercambre-Darras, S., Devos, P., D’Herbomez, M., Dubucquoi, S., & Mortier, L. (2013). Metastatic melanoma: spontaneous occurrence of auto antibodies is a good prognosis factor in a prospective cohort. Journal of the European Academy of Dermatology and Venereology, 27(1), 92–96.
Maniotis, A. J., Folberg, R., Hess, A., Seftor, E. A., Gardner, L. M. G., Pe’er, J., et al. (1999). Vascular channel formation by human melanoma cells in vivo and in vitro: Vasculogenic mimicry. American Journal of Pathology, 155(3), 739–752. doi:10.1016/S0002-9440(10)65173-5.
Martin, F. P. J., Dumas, M. E., Wang, Y. L., Legido-Quigley, C., Yap, I. K. S., Tang, H. R., et al. (2007a). A top-down systems biology view of microbiome-mammalian metabolic interactions in a mouse model. Molecular Systems Biology, 3, 112.
Martin, F. P. J., Wang, Y. L., Sprenger, N., Holmes, E., Lindon, J. C., Kochhar, S., et al. (2007b). Effects of probiotic Lactobacillus paracasei treatment on the host gut tissue metabolic profiles probed via magic–angle–spinning NMR spectroscopy. Journal of Proteome Research, 6(4), 1471–1481.
Massari, N. A., Medina, V. A., Cricco, G. P., Lamas, D. J. M., Sambuco, L., Pagotto, R., et al. (2013). Antitumor activity of histamine and clozapine in a mouse experimental model of human melanoma. Journal of Dermatological Science, 72(3), 252–262. doi:10.1016/j.jdermsci.2013.07.012.
Mizuarai, S., Miki, S., Araki, H., Takahashi, K., & Kotani, H. (2005). Identification of dicarboxylate carrier Slc25a10 as malate transporter in de novo fatty acid synthesis. Journal of Biological Chemistry, 280(37), 32434–32441. doi:10.1074/jbc.M503152200.
Morvan, D., Demidem, A., Guenin, S., & Madelmont, J. C. (2006). Methionine-dependence phenotype of tumors: Metabolite profiling in a melanoma model using l-[methyl-C-13]methionine and high-resolution magic angle spinning H-1-C-13 nuclear magnetic resonance spectroscopy. Magnetic Resonance in Medicine, 55(5), 984–996.
Morvan, D., Demidem, A., Papon, J., De Latour, M., & Madelmont, J. C. (2002). Melanoma tumors acquire a new phospholipid metabolism phenotype under cystemustine as revealed by high-resolution magic angle spinning proton nuclear magnetic resonance spectroscopy of intact tumor sampled. Cancer Research, 62(6), 1890–1897.
Morvan, D., Demidem, A., Papon, J., & Madelmont, J. C. (2003). Quantitative HRMAS proton total correlation spectroscopy applied to cultured melanoma cells treated by chloroethyl nitrosourea: Demonstration of phospholipid metabolism alterations. Magnetic Resonance in Medicine, 49(2), 241–248.
Mounet, F., Lemaire-Chamley, M., Maucourt, M., Cabasson, C., Giraudel, J. L., Deborde, C., et al. (2007). Quantitative metabolic profiles of tomato flesh and seeds during fruit development: Complementary analysis with ANN and PCA. Metabolomics, 3(3), 273–288. doi:10.1007/s11306-007-0059-1.
Namkoong, J., Shin, S. S., Lee, H. J., Marin, Y. E., Wall, B. A., Goydos, J. S., et al. (2007). Metabotropic glutamate receptor 1 and glutamate signaling in human melanoma. Cancer Research, 67(5), 2298–2305.
Nicholson, J. K., Foxall, P. J. D., Spraul, M., Farrant, R. D., & Lindon, J. C. (1995). 750-Mhz H-1 and H-1-C-13 NMR-spectroscopy of human blood-plasma. Analytical Chemistry, 67(5), 793–811.
Orlov, N. V., Weeraratna, A. T., Hewitt, S. M., Coletta, C. E., Delaney, J. D., Eckley, D., et al. (2012). Automatic detection of melanoma progression by histological analysis of secondary sites. Cytometry Part A, 81A(5), 364–373.
Patwardhan, G. A., & Liu, Y. Y. (2011). Sphingolipids and expression regulation of genes in cancer. Progress in Lipid Research, 50(1), 104–114. doi:10.1016/j.plipres.2010.10.003.
Pontarin, G., Ferraro, P., Hakansson, P., Thelander, L., Reichard, P., & Bianchi, V. (2007). p53R2-dependent ribonucleotide reduction provides deoxyribonucleotides in quiescent human fibroblasts in the absence of induced DNA damage. Journal of Biological Chemistry, 282(23), 16820–16828. doi:10.1074/jbc.M701310200.
Pos, Z., Safrany, G., Muller, K., Toth, R., Falus, A., & Hegyesi, H. (2005). Phenotypic profiling of engineered mouse melanomas with manipulated histamine production identifies histamine H2 receptor and Rho-C as histamine-regulated melanoma progression markers. Cancer Research, 65(10), 4458–4466. doi:10.1158/0008-5472.Can-05-0011.
Qiu, Y., Rajagopalan, D., Connor, S. C., Damian, D., Zhu, L., Handzel, A., et al. (2008). Multivariate classification analysis of metabolomic data for candidate biomarker discovery in type 2 diabetes mellitus. Metabolomics, 4(4), 337–346. doi:10.1007/s11306-008-0123-5.
Rocha, C. M., Carrola, J., Barros, A. S., Gil, A. M., Goodfellow, B. J., Carreira, I. M., et al. (2011). Metabolic signatures of lung cancer in biofluids: NMR-based metabonomics of blood plasma. Journal of Proteome Research, 10(9), 4314–4324. doi:10.1021/Pr200550p.
Roomi, M. W., Kalinovsky, T., Roomi, N. W., Ivanov, V., Rath, M., & Niedzwiecki, A. (2008). Suppression of growth and hepatic metastasis of murine B16FO melanoma cells by a novel nutrient mixture. Oncology Reports, 20(4), 809–817. doi:10.3892/or_00000078.
Rzeski, W., Turski, L., & Ikonomidou, C. (2001). Glutamate antagonists limit tumor growth. Proceedings of the National Academy of Sciences of the USA, 98(11), 6372–6377.
Sato, A., Yoshikawa, N., Kubo, E., Kakuda, M., Nishiuchi, A., Kimoto, Y., et al. (2013). Inhibitory Effect of Cordycepin on Experimental Hepatic Metastasis of B16-F0 Mouse Melanoma Cells. Vivo, 27(6), 729–732.
Sikora-Borgula, A., Slominska, M., Trzonkowski, P., Zielke, R., Mysliwski, A., Wegrzyn, G., et al. (2002). A role for the common GTP-binding protein in coupling of chromosome replication to cell growth and cell division. Biochemical and Biophysical Research Communications, 292(2), 333–338. doi:10.1006/bbrc.2002.6671.
Song, Z. Q., He, C. D., Liu, J., Sun, C. K., Lu, P., Li, L. L., et al. (2012). Blocking glutamate-mediated signalling inhibits human melanoma growth and migration. Experimental Dermatology, 21(12), 926–931. doi:10.1111/Exd.12048.
Stepulak, A., Sifringer, M., Rzeski, W., Endesfelder, S., Gratopp, A., Pohl, E. E., et al. (2005). NMDA antagonist inhibits the extracellular signal-regulated kinase pathway and suppresses cancer growth. Proceedings of the National Academy of Sciences of the USA, 102(43), 15605–15610.
Stretch, J. R., Somorjai, R., Bourne, R., Hsiao, E., Scolyer, R. A., Dolenko, B., et al. (2005). Melanoma metastases in regional lymph nodes are accurately detected by proton magnetic resonance spectroscopy of fine-needle aspirate biopsy samples. Annals of Surgical Oncology, 12(11), 943–949.
Szymanska, E., Saccenti, E., Smilde, A. K., & Westerhuis, J. A. (2012). Double-check: validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics, 8(1), S3–S16. doi:10.1007/s11306-011-0330-3.
Triba, M. N., Starzec, A., Bouchemal, N., Guenin, E., Perret, G. Y., & Le Moyec, L. (2010). Metabolomic profiling with NMR discriminates between biphosphonate and doxorubicin effects on B16 melanoma cells. NMR in Biomedicine, 23(9), 1009–1016.
Trygg, J., & Wold, S. (2002). Orthogonal projections to latent structures (O-PLS). Journal of Chemometrics, 16(3), 119–128. doi:10.1002/Cem.695.
Washburn, K. W. (1989). A modification of the Folch method of lipid extraction for poultry. Poultry Science, 68(11), 1425–1427.
Weljie, A. M., Dowlatabadi, R., Miller, B. J., Vogel, H. J., & Jirik, F. R. (2007). An inflammatory arthritis-associated metabolite biomarker pattern revealed by H-1 NMR Spectroscopy. Journal of Proteome Research, 6(9), 3456–3464. doi:10.1021/Pr070123j.
Wiklund, S., Johansson, E., Sjostrom, L., Mellerowicz, E. J., Edlund, U., Shockcor, J. P., et al. (2008). Visualization of GC/TOF-MS-based metabolomics data for identification of biochemically interesting compounds using OPLS class models. Analytical Chemistry, 80(1), 115–122. doi:10.1021/Ac0713510.
Wu, B. J., Li, W. P., Qian, C., Ding, W., Zhou, Z. W., & Jiang, H. (2013). Increased serum level of thymidine kinase 1 correlates with metastatic site in patients with malignant melanoma. Tumor Biology, 34(2), 643–648. doi:10.1007/s13277-012-0591-0.
Xia, J. M., Wu, X. J., & Yuan, Y. J. (2007). Integration of wavelet transform with PCA and ANN for metabolomics data-mining. Metabolomics, 3(4), 531–537. doi:10.1007/s11306-007-0090-2.
Yu, J., & Kim, A. K. (2009). Effect of taurine on antioxidant enzyme system in B16F10 melanoma cells. Taurine, 7(643), 491–499. doi:10.1007/978-0-387-75681-3_51.
Acknowledgments
Research reported in this publication was supported by the National Institute of Environmental Health Sciences of the National Institute of Health (NIH) under Award Numbers R01ES022176, and NIH/R01EB013231. All of the NMR experiments were performed in the Environmental Molecular Sciences Laboratory, a national scientific user facility sponsored by the DOE’s Office of Biological and Environmental Research, and located at PNNL. PNNL is a multi-program national laboratory operated for the DOE by Battelle Memorial Institute under Contract DE-AC06-76RLO 1830. And finally we thank Dr. Xihai Wang for his contributions in pathology and animal work.
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, X., Hu, M., Feng, J. et al. 1H NMR metabolomics study of metastatic melanoma in C57BL/6J mouse spleen. Metabolomics 10, 1129–1144 (2014). https://doi.org/10.1007/s11306-014-0652-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-014-0652-z