Abstract
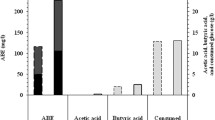
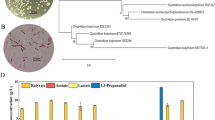
Three hyper 2-propanol producing strains were isolated from Singapore environment using an enrichment step and a high through-put screening step. The analysis of the amplified 16S rDNA revealed that the isolates belonged to Clostridium species and they were named as Clostridium sp. BT10-1, Clostridium sp. M10-1 and Clostridium sp. PU31-4. At 1 L scale, the 2-propanol titer of these positive strains was 1.6–2.1 times of that of Clostridium beijerinckii NRRL B593, which is so far the most efficient natural 2-propanol producer. The highest 2-propanol titer was achieved by isolate BT10-6 and it was 5.26 g/L (87.5 mM). These three positive strains BT10-6, M10-1 and PU31-4 consumed glucose almost completely in 40–48 h and gave 2-propanol productivity at 0.132, 0.118 and 0.087 g/L/h, respectively, which is 3.0–4.6 times of 0.029 g/L/h given by C. beijerinckii NRRL B593. Butanol was also produced by these positive strains with a slightly lower butanol titer and higher butanol productivity, compared to butanol control strain C. beijerinckii NCIMB 8052.





Similar content being viewed by others
References
Andreesen JR, Bahl H, Gottschalk G (1989) Introduction to the physiology and biochemistry of the genus Clostridium. In: Minton NP, Clarke DJ (eds) Biotechnology handbooks, vol 3., ClostridiaPlenum press, New York, pp 27–62
Annous BA, Blaschek HP (1991) Isolation and characterization of Clostridium acetobutylicum mutants with enhanced amylolytic activity. Appl Environ Microb 57:2544–2548
Chen JS, Hiu SF (1986) Acetone-butanol-isopropanol production by Clostridium beijerinckii (synonym, Clostridium butylicum). Biotechnol Lett 8:371–376. doi:10.1007/BF01040869
Evans PJ, Wang HW (1988) Enhancement of butanol formation by Clostridium acetobutylicum in the presence of decanol-oleyl alcohol mixed extractants. Appl Environ Microbiol 54:1662–1667
Ezeji TC, Qureshi N, Blaschek HP (2004) Acetone-butanol-ethanol (ABE) production from concentrated substrate: reduction in substrate inhibition by fed-batch technique and product inhibition by gas stripping. Appl Microbiol Biotechnol 63:653–658. doi:10.1007/s00253-003-1400-x
Ezeji TC, Qureshi N, Karcher PM, Blaschek HP (2005) Improving the performance of a gas stripping-based recovery system to remove butanol from Clostridium beijerinckii fermentation. Bioprocess Biosyst Eng 27:207–214. doi:10.1007/s00449-005-0403-7
Ezeji TC, Milne C, Price ND, Blaschek HP (2010) Achievements and perspectives to overcome the poor solvent resistance in acetone and butanol-producing microorganisms. Appl Microbiol Biotechnol 85:1697–1712. doi:10.1007/s00253-009-2390-0
Garcia A, Iannotti EL, Fischer JL (1986) Butanol fermentation liquor production and separation by reverse osmosis. Biotechnol Bioeng 28:785–791. doi:10.1002/bit.260280603
Groot WJ, Luyben KChAM (1986) In situ product recovery by adsorption in the butanol/isopropanol batch fermentation. Appl Microbiol Biotechnol 25:29–31. doi:10.1007/BF00252508
Groot WJ, van den Oever CE, Kossen NWF (1984) Pervaporation for simultaneous product recovery in the butanol/isopropanol batch fermentation. Biotechnol Lett 6:709–714. doi:10.1007/BF00133061
Hanai T, Atsumi S, Liao JC (2007) Engineered synthetic pathway for isopropanol production in Escherichia coli. Appl Environ Microb 73:7814–7818. doi:10.1128/AEM.01140-07
Heipieper HJ, Weber FJ, Sikkema J, Keweloh H, Debont JAM (1994) Mechanisms of resistance of whole cells to toxic organic-solvents. Trends Biotechnol 12:409–415. doi:10.1016/0167-7799(94)90029-9
Jojima T, Inui M, Yukawa H (2008) Production of isopropanol by metabolically engineered Escherichia coli. Appl Microbiol Biotechnol 77:1219–1224. doi:10.1007/s00253-007-1246-8
Kasap M (2002) Nitrogen metabolism and solvent production in Clostridium beijerinckii NRRL B593. Virginia Polytechnic Institute and State University, Blacksburg Dissertation
Kibby CL, Hall WK (1972) Studies of acid catalyzed reactions. XII. Alcohol decomposition over hydroxyapatite catalysts. J Catal 29:144–159. doi:10.1016/0021-9517(73)90213-3
Krouwel PG, Groot WJ, Kossen NWF, van der Laan WFM (1983) Continuous isopropanol-butanol-ethanol fermentation by immobilized Clostridium beijerinckii cells in a packed bed fermenter. Enzym Microb Technol 5:46–54. doi:10.1016/0141-0229(83)90064-9
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Lee I, Johnson LA, Hammond EG (1995) Use of branched chain esters to reduce the crystallization temperature of biodiesel. J Am Oil Chem Soc 72:1155–1160. doi:10.1007/BF02540982
Matsumura M, Takehara S, Kataoka H (1992) Continuous butanol/isopropanol fermentation in down-flow column reactor coupled with pervaporation using supported liquid membrane. Biotechnol Bioeng 39:148–156. doi:10.1002/bit.260390205
Montoya D, Spitia S, Silva E, Schwarz WH (2000) Isolation of mesophilic solvent-producing Clostridia from Colombian sources: physiological characterization, solvent production and polysaccharide hydrolysis. J Biotechnol 79:117–126. doi:10.1016/S0168-1656(00)00218-2
Osburn OL, Brown RW, Werkman CH (1937) The butyl alcohol-isoppropyl alcohol fermentation. J Biol Chem 121:685–695
Papa AJ (2005) Propanols in: Ullmann’s encyclopedia of industrial chemistry. Wiley-VCH, Weinheim. doi:10.1002/14356007.a22_173
Qureshi N, Blaschek HP (1999) Butanol recovery from model solution/fermentation broth by pervaporation: evaluation of membrane performance. Biomass Bioenergy 17:175–184. doi:10.1016/S0961-9534(99)00030-6
Qureshi N, Maddox IS, Friedl A (1992) Application of continuous substrate feeding to the ABE fermentation: relief of product inhibition using extraction, perstraction, stripping and pervaporation. Biotechnol Prog 8:382–390. doi:10.1021/bp00017a002
Sangster J (1989) Octanol-water partition coefficient of simple organic compounds. J Phys Chem Ref Data 18:1111–1227. doi:10.1063/1.555833
Shi Z, Blaschek HP (2008) Transcriptional analysis of Clostridium beijerinckii NCIMB 8052 and the hyper-butanol-producing mutant BA101 during the shift from acidogenesis to solventogenesis. Appl Environ Microb 74:7709–7714. doi:10.1128/AEM.01948-08
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Bio Evol 24:1596–1599. doi:10.1093/molbev/msm092
Thauer RK, Jungermann K, Decker K (1977) Energy conservation in thermotropic anaerobic bacteria. Bacteriol Rev 41:100–180
Vermue M, Sikkema J, Verheul A, Bakker R, Tramper J (1993) Toxicity of homologous series of organic-solvents for the gram positive bacteria Arthrobacter and Nocardia sp. and the gram negative bacteria Acinetobacter and Pseudomonas sp. Biotechnol Bioeng 42:747–758. doi:10.1002/bit.260420610
Zoutberg GR, Willemsberg R, Smit G, Teixeira de Mattos MJ, Neijssel OM (1989) Solvent production by an aggregate-forming variant of Clostridium butyricum. Appl Microbiol Biotechnol 32:22–26. doi:10.1007/BF00164817
Acknowledgments
This work was supported by the Agency for Science, Technology and Research (A*star), Singapore (ICES/08-374B01) and was collaborated with Mitsui Chemical Singapore R&D Centre, Pte Ltd.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ng, Z.R., Takahashi, K. & Liu, Z. Isolation, characterization and evaluation of hyper 2-propanol producing bacteria from Singapore environment. World J Microbiol Biotechnol 29, 1059–1065 (2013). https://doi.org/10.1007/s11274-013-1269-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-013-1269-5