Abstract
White spot syndrome virus (WSSV) is one of the major pathogens of cultured shrimp. Identification of envelope protein interactions has become a central issue for the understanding of WSSV assembly. In this paper, WSSV envelope protein VP52B was fused with GST-tag and expressed in Escherichia coli BL-21(DE3). Immunogold-electron microscopy revealed that VP52B was located on the outside surface of WSSV virions. Far-Western blotting analysis suggested that VP52B might directly interact with a major viral envelope protein VP26, and their interaction was confirmed by GST pull-down assay. Further investigation showed that the VP52B binding domain was located between residues 135–170 of VP26. These findings will enhance our understanding of the molecular mechanisms of WSSV morphogenesis.
Similar content being viewed by others
Introduction
White spot syndrome virus (WSSV), the sole member of the genus Whispovirus of the family Nimaviridae [1], is an extremely virulent pathogen of shrimp and crayfish [2, 3]. The virus contains a large double-stranded circular DNA genome (about 300 kbp) encoding more than 180 predicted proteins [4, 5]. WSSV structural proteins have been identified using mass spectrometry analysis and shotgun proteomics, and over 30 proteins are located to the viral envelope [6–8].
Viral envelope proteins are the first factors to interact with the host cell membrane during infection, resulting in host cell targeting and triggering host antiviral reaction [9]. In the case of WSSV, some evidences suggest that its infectivity is influenced by viral envelope proteins, indicating a potential relationship of viral envelope proteins and host antiviral reaction [10, 11]. Beside the interaction with host membrane, viral envelope proteins are essential for viral assembly and maturation. For example, an envelope protein complex of vaccinia virus was proved to be necessary for the formation of viral membranes and pre-mature virions [12]. So far, the assembly mechanism of WSSV virions is not well understood. Therefore, knowledge of the interaction among WSSV envelope proteins will assist our understanding of WSSV morphogenesis, as well as the early events of WSSV-host interaction.
VP28, VP26, VP24, and VP19 are four most abundance envelope proteins of WSSV. Pair-wise interactions among these proteins have been demonstrated, which include VP28-VP24, VP28-VP26, VP28-VP19, VP26-VP24, and VP24-VP19 [13–15]. Using two-dimensional blue native/SDS-PAGE and mass spectrometry, Li et al. predicted that VP26 and VP24 might function as hub proteins to recruit low-abundant WSSV structural proteins to complete virion morphogenesis and constructed a model of WSSV envelope assembly [16]. The model has been supported by a number of recent findings of WSSV protein–protein interactions, including interactions of VP26 with VP90 [17], VP37 [18], and VP52A (also termed VP51A in TW-isolate) [13], as well as interactions of VP24 with VP33 [19], VP38 [20], and WSV010 [21]. The two-dimensional blue native/SDS-PAGE analysis also suggested a potential interaction between a low-abundant structural protein VP52B (also termed VP51B) and VP26 or VP24 [16].
In this paper, we focus on viral envelope protein VP52B. The VP52B protein is encoded by wsv256. This protein consists of 383 amino acids. Topological predictions by TMHMM 2.0 suggested that VP52B contains a transmembrane region in its N-terminal (data not shown). No putative conserved domains have been detected using BLAST analyzing tool (data not shown). Since the low homology of VP52B to other proteins, its function remains hard to predict. Nevertheless, using the analysis methods of protein interaction, we demonstrate that VP26 can bind to WSSV envelop protein VP52B through a domain located between residues 135–170 of VP26. These results will extend our understanding of WSSV morphogenesis.
Materials and methods
Preparation of WSSV virions, envelope and nucleocapsid fractions
Crayfish (Procambarus clarkia), the host for viral replication, were purchased from a local market in Xiamen, China. WSSV virions were purified, and the envelope and nucleocapsid fractions were prepared as previously reported [8, 22].
Plasmid construction, protein purification, and antibody preparation
The vp52B ORF (wsv256) was amplified from the genomic DNA with the following primers vp52B+: 5′-GCGGGATCCATGATATTTTATACAATGCAAC-3′ and vp52B−: 5′-GCGGTCGACATCGTCTACCAAATTTTCTG-3′. The PCR product was cloned into pGEX-2TH vector in-frame with GST and poly-His tag (constructed by adding a poly-His tag to pGEX-2T) to generate pGEX-vp52B-2TH plasmid.
The expression plasmids of four major WSSV envelope proteins (VP28, VP26, VP24, and VP19), pET-vp28-His, pET-vp26-His, pET28a-vp24-V5, and pET-vp19-His, were constructed by our lab [8, 23]. The N-terminal hydrophobic regions of these proteins have been removed to ensure soluble expression, and each of the protein was fused with a poly-His tag.
The rVP26 mutant plasmids (Fig. 4c) were generated by inverse PCR using pET-vp26-His as the template, and were named as pET-vp26(68-204)-His, pET-vp26(102-204)-His, pET-vp26(135-204)-His, pET-vp26(36-102)-His, pET-vp26(36-135)-His, and pET-vp26(36-170)-His, according to the vp26 fragments they contained. The primers used to construct these recombinant plasmids were listed in Supplementary Table 1.
For protein expression and purification, the recombinant plasmids were transformed into E. coli BL21 (DE3), and each of the transformants was grown in 200 mL selective (Amp) LB medium at 37 °C until OD600 reaching 0.6–0.8. Recombinant protein expression was then induced by 0.05 mM IPTG for 24 h at 16 °C. Cells were harvested by centrifugation. The bacterial pellets were resuspended, sonicated, and centrifuged. GST-tagged VP52B in the supernatant was purified using the glutathione sepharose 4B (GE Healthcare). His-tagged or v5-tagged recombinant proteins (rVP28, rVP26, rVP24, rVP19, and six rVP26 mutations) in the supernatant of sonicated bacterial sample were directly used in the GST pull-down analysis without purification.
For antiserum preparation, the purified GST-tagged VP52B was used to immunize mice by subcutaneous injection as previously described [24].
Immunoelectron microscopy (IEM)
For IEM, purified WSSV virions and nucleocapsids were mounted on a carbon-coated copper grid for 30 min at room temperature (RT), and the excess liquid was carefully removed with filter paper. For immunogold labeling, the sample grids were blocked for 30 min in phosphate-buffered saline (PBS) containing 2 % BSA and 0.05 % Tween-20, then incubated with anti-rVP52B serum (1:200, diluted in 2 % BSA) for 1 h at RT in a wet chamber. After washing four times with PBS, the grids were incubated for 1 h at RT with 10 nm (diameter)-gold-conjugated goat anti-mouse IgG (Sigma) diluted 1:20 in 2 % PBS-BSA. The grids were then washed four times with PBS and stained with 2 % phosphotungstic acid (PTA, pH 7.0) for 1 min. Specimens were examined under transmission electron microscope (JEM-1230, JEOL). Pre-bled mouse serum was used as the negative control.
Western blotting
Proteins were separated by 12 % SDS-PAGE and transferred to PVDF membrane (Immobilon-P, Millipore). The membrane was blocked with blocking buffer (Thermo) for 1 h at RT and probed with the corresponding primary antibody for 1 h at RT. After washed in TBST for three times, the PVDF membrane was incubated with AP conjugated goat anti-mouse IgG secondary antibody (1:7500; Promega), and the signals were detected using NBT/BCIP substrate (Roche).
Far-western blotting
Far-Western assay was performed as previously described [24]. In Brief, the WSSV envelope fraction was separated by 12 % SDS-PAGE, following by transferred onto a PVDF membrane, and renatured by successive 30-min incubations at RT with 6 M, 3 M, 1.5 M, 0.75 M, 0.325 M, and 0.15 M Guanidine-HCl in renaturation buffer (20 mM Tris–HCl/PH 7.5, 150 mM NaCl, 1 mM EDTA, 1 mM dithiothreitol, 2 % nonfat milk, 0.1 % Tween-20, and 10 % glycerol). Subsequently, the membrane was immersed in renaturation buffer overnight at 4 °C. The membrane containing renatured WSSV envelope proteins was incubated with GST-VP52B or GST alone as negative control in blocking buffer for 2 h at RT, and then analyzed by Western blotting using anti-GST antibody (1:1000; GE Healthcare).
GST pull-down analysis
The glutathione sepharose 4B (Pierce Biotechnology) was pre-treated with blocking buffer for 30 min to avoid non-specific binding. Five micrograms of GST-VP52B protein or GST protein (negative control) was immobilized with 10 µL of glutathione beads for 2 h at 4 °C. The beads were then incubated with the supernatant of sonicated bacterial sample containing target proteins. The binding process was carried out in the binding buffer (20 mmol/L Tris–HCl, 300 mmol/L NaCl, 0.5 % Triton X-100, and 2 % BSA and protease inhibitors) for 2 h at 4 °C. The resins were extensively washed by washing buffer (20 mmol/L Tris–HCl, 500 mmol/L NaCl, and 0.5 % Triton X-100) to removed unbound proteins. After that, the immobilized proteins on the resins were separated by 12 % SDS-PAGE and transferred to PVDF membrane, and analyzed by Western blotting. Anti-VP28 (1:10000), anti-VP26 (1:8000), anti-VP24 (1:1000), and anti-VP19 (1:8000) antibodies were used as primary antibody to detect the four major envelope proteins, respectively. Anti-his antibody (1:3000; GE Healthcare) was used to detect the rVP26 mutants.
Results
VP52B is located on the surface of WSSV virions
Previous study indicated that VP52B might be presented in WSSV envelope [16]. To determine the localization of VP52B, IEM and Western blotting analysis were performed. The IEM results showed that when the sample was probed with anti-VP28 antibody, gold particles were observed on the surface of the virions (Fig. 1b), but not in the nucleocapsids (Fig. 1c). No gold particles were detected on the virions when the sample was probed with pre-bled mouse serum (Fig. 1d). Western blotting analysis showed that VP52B was presented in the envelope fraction, but not the nucleocapsid fraction of WSSV (Fig. 1e), which was coincident to the IEM results. Therefore, VP52B is an envelope protein located on the outside surface of WSSV virions.
Localization of VP52B in WSSV virions. a Electron micrograph of negatively stained purified virions (2 % PTA); b The virions incubated with anti-VP52B antiserum as primary antibody followed by gold-labeled secondary antibodies. Gold particles are indicated by arrows. c The viral nucleocapsids incubated with anti-VP52B antiserum as primary antibody followed by gold-labeled secondary antibodies. d The virions incubated with normal mouse serum as primary antibody followed by gold-labeled secondary antibodies. (Scale bar, 200 nm) e WSSV virions (lanes 1 and 4), envelope fraction (lanes 2 and 5), and nucleocapsid fraction (lanes 3 and 6) were separated by 12 % SDS-PAGE and detected by Coomassie blue staining (lanes 1, 2 and 3) or Western blotting using anti-VP52B antibody (lanes 4, 5, and 6). VP52B is indicated by arrow
Direct interaction between VP52B and VP26
Far-Western blotting was performed to analyze the interaction of VP52B with other viral envelope protein(s). The viral proteins in the envelope fractions were separated by SDS-PAGE, transferred onto PVDF membrane, and renatured. The membrane with renatured proteins was incubated with purified GST-VP52B or GST alone, and the target proteins were detected using anti-GST antibody. One single band (Fig. 2 lane 2) corresponded to VP26 (Fig. 2 lane 1, 4) was detected when the membrane was probed with GST-VP52B, while no band was detected when probed with GST alone (Fig. 2 lane 3). The result suggests that there is a direct interaction between VP52B and VP26.
Direct interaction of VP52B with VP26 by far-Western blotting. The viral envelope proteins were separated by 12 % SDS-PAGE and subjected to Coomassie blue staining (lane 1), as well as far-Western blotting using GST-VP52B (lane 2) or GST (lane 3, negative control) as overlay protein and detected with anti-GST monoclonal antibody. The position of VP26 was visualized by Western blotting using anti-VP26 antibody (lane 4). The four major envelope proteins, VP28, VP26, VP24, and VP19 are marked with asterisks
To further confirm the interaction between VP52B and VP26, four major viral envelope proteins VP28, VP26, VP24, and VP19 were expressed and subjected to GST pull-down assays. The rVP28, rVP26, rVP24, and rVP19 were incubated with GST-VP52B-bound or GST-bound glutathione resins. The pull-down fractions were then examined by SDS/PAGE and Western blotting. As expected, GST-VP52B exhibited interaction with rVP26 (Fig. 3b) but not rVP28, rVP24, and rVP19 (Fig. 3a, c, d). There is no non-specific interaction between GST protein and the four WSSV envelope proteins.
Interaction of VP52B with VP26 by GST pull-down. rVP28 (a, lane 1), rVP26 (b, lane 1), rVP24 (c, lane 1), and rVP19 (d, lane 1) were incubated with GST-VP52B (a–d, lane 2) or GST alone (a–d, lane 3) in the binding buffer for 2 h, and were detected using anti-VP28, anti-VP26, anti-VP24, or anti-VP19 antibodies, respectively
Identification of the VP52B binding domain in VP26
To identify the VP52B binding domain in VP26, six VP26 deletion mutants, VP26 (68-204), VP26 (102-204), VP26 (135-204), VP26 (36-102), VP26 (36-135), and VP26 (36-170), were incubated with GST-VP52B-bounded resins. The pull-down fractions were analyzed by Western blotting using anti-His antibody. As shown in Fig. 4b, four VP26 mutants including VP26 (68-204), VP26 (102-204), VP26 (135-204), and VP26 (36-170) could interact with VP52B, while VP26 (36-135) and VP26 (36-102) failed to bind to VP52B. Therefore, the VP52B binding domain of VP26 is located between residues 135 and 170.
Interactions between VP26 deletion mutants and VP52B. a Expression of six VP26 mutants, VP26(68-204), VP26(102-204), VP26(135-204), VP26(36-102), VP26(36-135), and VP26(36-170), were examined by Western blotting using anti-His antibody. b GST-VP52B was immobilized onto glutathione sepharose 4B and incubated with VP26(68-204), VP26(102-204), VP26(135-204), VP26(36-102), VP26(36-135), or VP26(36-170). The binding proteins on the resins were detected by Western blotting using anti-His antibody. c Schematic illustration of VP26 deletion mutants and the results of their interaction with VP52B. Plus or minus sign indicates interaction or no interaction with VP52B, respectively
Discussion
WSSV is an enveloped virus composed of about 30 structural proteins [8, 25]. VP28, VP26, VP24, and VP19 are four major envelope proteins of WSSV, accounting for about 90 % of all viral envelope proteins and can form multi-protein complexes (MPCs) [14]. Recent work suggests that the WSSV envelope consists of a series of MPCs. Low-abundant envelope proteins may be incorporated into the MPCs via interacting with viral major envelope proteins, especially hub proteins VP26 and VP24 [16]. Based on the co-migrated model in blue native PAGE [16], we speculate that there may be an interaction between VP52B and either or both of these two hub proteins. In the current study, we first confirmed the distribution of VP52B on WSSV virions by Western blotting and IEM analysis, and demonstrated that VP52B is a envelop protein located on the outside surface of WSSV. Far-Western blotting was performed to explore the interaction between VP52B and other WSSV envelop proteins. The results suggested a direct interaction between VP52B and VP26 (Fig. 2). Their interaction was confirmed by GST pull-down analysis in which GST VP52B was able to pull-down rVP26 but not rVP28, rVP24, or rVP19 (Fig. 3). The results further support the hypothesis that VP26 serves as a hub protein to recruit other low-abundant envelope proteins during viral envelope formation.
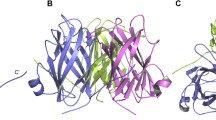
Although new evidence suggests that VP26 is a hub protein which not only interacts with other envelope proteins, but also acts as an envelope-nucleocapsid linker through its binding with capsid protein [26], the specific interaction domain of VP26 with other viral structural proteins remains largely unknown. To explore the regions in VP26 that involved in its interaction with VP52B, a series of VP26 deletion mutants were generated and their interaction with VP52B was investigated by GST pull-down assay. As shown in Fig. 4, the VP52B-interacting domain of VP26 was located in residues 135–170 (Fig. 4). Base on the crystal structure of VP26, residues 135–170 are located on the outside of the β-barrel structure of VP26 [27]. Therefore this region is available for interaction with other proteins.
Taken together, our findings suggested that WSSV envelop protein VP52B might be incorporated into WSSV MPCs via its interaction with VP26 and participate in WSSV envelope assembly. This will improve our understanding of WSSV morphogenesis.
References
M.A. Mayo, A summary of taxonomic changes recently approved by ICTV. Arch. Virol. 147(8), 1655–1663 (2002)
V. Corbel, Z. Zuprizal, C. Shi, J. Arcier, J. Bonami, Experimental infection of European crustaceans with white spot syndrome virus (WSSV). J. Fish Dis. 24, 377–382 (2001)
C.F. Lo, C.H. Ho, S.E. Peng, C.H. Chen, H.C. Hsu, Y.L. Chiu et al., White spot syndrome baculovirus (WSBV) detected in cultured and captured shrimp, crabs and other arthropods. Dis. Aquat. Org. 27, 215–225 (1996)
M.C. van Hulten, J. Witteveldt, S. Peters, N. Kloosterboer, R. Tarchini, M. Fiers et al., The white spot syndrome virus DNA genome sequence. Virology 286(1), 7–22 (2001)
F. Yang, J. He, X. Lin, Q. Li, D. Pan, X. Zhang, X. Xu, Complete genome sequence of the shrimp white spot bacilliform virus. J. Virol. 75(23), 11811–11820 (2001)
Z. Li, Q. Lin, J. Chen, J.L. Wu, T.K. Lim, S.S. Loh et al., Shotgun identification of the structural proteome of shrimp white spot syndrome virus and iTRAQ differentiation of envelope and nucleocapsid subproteomes. Mol. Cell. Proteomics 6(9), 1609–1620 (2007)
J.M. Tsai, H.C. Wang, J.H. Leu, H.H. Hsiao, A.H. Wang, G.H. Kou, C.F. Lo, Genomic and proteomic analysis of thirty-nine structural proteins of shrimp white spot syndrome virus. J. Virol. 78(20), 11360–11370 (2004)
X. Xie, L. Xu, F. Yang, Proteomic analysis of the major envelope and nucleocapsid proteins of white spot syndrome virus. J. Virol. 80(21), 10615–10623 (2006)
N. Chazal, D. Gerlier, Virus entry, assembly, budding, and membrane rafts. Microbiol. Mol. Biol. Rev. 67(2), 226–237 (2003)
L.J. Li, J.F. Yuan, C.A. Cai, W.G. Gu, Z.L. Shi, Multiple envelope proteins are involved in white spot syndrome virus (WSSV) infection in crayfish. Arch. Virol. 151(7), 1309–1317 (2006)
W. Wu, L. Wang, X. Zhang, Identification of white spot syndrome virus (WSSV) envelope proteins involved in shrimp infection. Virology 332(2), 578–583 (2005)
P. Szajner, H. Jaffe, A.S. Weisberg, B. Moss, A complex of seven vaccinia virus proteins conserved in all chordopoxviruses is required for the association of membranes and viroplasm to form immature virions. Virology 330(2), 447–459 (2004)
Y.S. Chang, W.J. Liu, T.L. Chou, Y.T. Lee, T.L. Lee, W.T. Huang et al., Characterization of white spot syndrome virus envelope protein VP51A and its interaction with viral tegument protein VP26. J. Virol. 82(24), 12555–12564 (2008)
Q. Zhou, L. Xu, H. Li, Y.P. Qi, F. Yang, Four major envelope proteins of white spot syndrome virus bind to form a complex. J. Virol. 83(9), 4709–4712 (2009)
X. Xie, F. Yang, White spot syndrome virus VP24 interacts with VP28 and is involved in virus infection. J. Gen. Virol. 87, 1903–1908 (2006)
Z. Li, L. Xu, F. Li, Q. Zhou, F. Yang, Analysis of white spot syndrome virus envelope protein complexome by two-dimensional blue native/SDS-PAGE combined with mass spectrometry. Arch. Virol. 156(7), 1125–1135 (2011)
Q. Li, Q.H. Liu, J. Huang, VP90 of white spot syndrome virus interacts with VP26 and VP28. Acta Virol. 56(1), 57–62 (2012)
Q.H. Liu, C.Y. Ma, W.B. Chen, X.L. Zhang, Y. Liang, S.L. Dong et al., White spot syndrome virus VP37 interacts with VP28 and VP26. Dis. Aquat. Org. 85(1), 23–30 (2009)
Y. Lin, L. Xu, F. Yang, Tetramerization of white spot syndrome virus envelope protein VP33 and its interaction with VP24. Arch. Virol. 155(6), 833–838 (2010)
Z. Jie, L. Xu, F. Yang, The C-terminal region of envelope protein VP38 from white spot syndrome virus is indispensable for interaction with VP24. Arch. Virol. 153(11), 2103–2106 (2008)
J. Chen, Z. Li, C.L. Hew, Characterization of a novel envelope protein WSV010 of shrimp white spot syndrome virus and its interaction with a major viral structural protein VP24. Virology 364(1), 208–213 (2007)
X. Xie, H. Li, L. Xu, F. Yang, A simple and efficient method for purification of intact white spot syndrome virus (WSSV) viral particles. Virus Res. 108(1–2), 63–67 (2005)
X. Xie, F. Yang, Interaction of white spot syndrome virus VP26 protein with actin. Virology 336(1), 93–99 (2005)
J. Li, L. Xu, F. Li, F. Yang, Low-abundance envelope protein VP12 of white spot syndrome virus interacts with envelope protein VP150 and capsid protein VP51. Virus Res. 178(2), 206–210 (2013)
J.M. Tsai, H.C. Wang, J.H. Leu, A.H. Wang, Y. Zhuang, P.J. Walker et al., Identification of the nucleocapsid, tegument, and envelope proteins of the shrimp white spot syndrome virus virion. J. Virol. 80(6), 3021–3029 (2006)
Q. Wan, L. Xu, F. Yang, VP26 of white spot syndrome virus functions as a linker protein between the envelope and nucleocapsid of virions by binding with VP51. J. Virol. 82(24), 12598–12601 (2008)
X. Tang, J. Wu, J. Sivaraman, C.L. Hew, Crystal structures of major envelope proteins VP26 and VP28 from white spot syndrome virus shed light on their evolutionary relationship. J. Virol. 81(12), 6709–6717 (2007)
Acknowledgments
This work was supported by Natural Science Foundation of China (No. 31272698), the Special Fund for Agro-scientific Research in the Public Interest (No. 201103034) and China Agriculture Research System (CARS-47). We would like to thank Analytical and Testing Center (TIO, SOA) for the instrumental assistance.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lin, F., Jie, Z., Hou, L. et al. Characterization of white spot syndrome virus VP52B and its interaction with VP26. Virus Genes 50, 46–51 (2015). https://doi.org/10.1007/s11262-014-1126-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-014-1126-0