Abstract
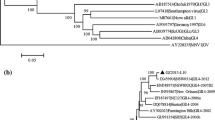
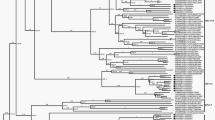
The complete nucleotide and deduced amino acid sequences of the RNA genome of a recently isolated norovirus (NoV) from Korea, designated Hu/GII-4/CBNU2/2007/KR (CBNU2), were determined and characterized by phylogenetic comparison with several genetically diverse NoV sequences. The RNA genome of CBNU2 is 7,560 nucleotides in length, excluding the 3′ poly (A) tract. It includes three open reading frames (ORFs): ORF1, which encodes the nonstructural polyprotein (5–5,104); ORF2, which encodes VP1 (5,085–6,707); and ORF3, which encodes VP2 (6,707–7,513). ORF2-based phylogenetic analysis revealed that CBNU2 belonged to the GII.4 genotype, the most prevalent genotype, and formed a cluster with NoVs isolated from Asian regions, between 2006 and 2008. Comparative analysis with the consensus sequence of 207 completely sequenced NoV genomes showed 47 mismatched nucleotides: 26 in ORF1, 14 in ORF2, and 7 in ORF3, resulting in 8 amino acid changes: 3 in ORF1, 2 in ORF2, and 3 in ORF3. Phylogenetic analysis with full genome ORF1, ORF2, and ORF3 nucleotide sequences obtained from CBNU2 and each of the other representative NoV genomes suggested that CBNU2 had not undergone recombination with any of the other NoVs. A SimPlot analysis further supported this finding.




Similar content being viewed by others
References
K.Y. Green, T. Ando, M.S. Balayan, T. Berke, I.N. Clarke, M.K. Estes, D.O. Matson, S. Nakata, J.D. Neill, M.J. Studdert, H.J. Thiel, J. Infect. Dis. 181(Suppl 2), S322–S330 (2000)
P.R. Lambden, E.O. Caul, C.R. Ashley, I.N. Clarke, Science 259, 516–519 (1993)
S. Inouye, K. Yamashita, S. Yamadera, M. Yoshikawa, N. Kato, N. Okabe, J. Infect. Dis. 181(Suppl 2), S270–S274 (2000)
M.C. Medici, M. Martinelli, L.A. Abelli, F.M. Ruggeri, I. Di Bartolo, M.C. Arcangeletti, F. Pinardi, F. De Conto, G. Izzi, S. Bernasconi, C. Chezzi, G. Dettori, J. Med. Virol. 78, 1486–1492 (2006)
R.L. Fankhauser, S.S. Monroe, J.S. Noel, C.D. Humphrey, J.S. Bresee, U.D. Parashar, T. Ando, R.I. Glass, J. Infect. Dis. 186, 1–7 (2002)
K.Y. Green, G. Belliot, J.L. Taylor, J. Valdesuso, J.F. Lew, A.Z. Kapikian, F.Y. Lin, J. Infect. Dis. 185, 133–146 (2002)
M.A. Widdowson, E.H. Cramer, L. Hadley, J.S. Bresee, R.S. Beard, S.N. Bulens, M. Charles, W. Chege, E. Isakbaeva, J.G. Wright, E. Mintz, D. Forney, J. Massey, R.I. Glass, S.S. Monroe, J. Infect. Dis. 190, 27–36 (2004)
J.Y. Chung, T.H. Han, S.H. Park, S.W. Kim, E.S. Hwang, J. Med. Virol. 82, 146–152 (2010)
T.H. Han, C.H. Kim, J.Y. Chung, S.H. Park, E.S. Hwang, Arch. Virol. 156, 323–329 (2011)
J.W. Huh, W.H. Kim, S.G. Moon, J.B. Lee, Y.H. Lim, J. Clin. Virol. 44, 152–156 (2009)
S.H. Kim, D.S. Cheon, J.H. Kim, D.H. Lee, W.H. Jheong, Y.J. Heo, H.M. Chung, Y. Jee, J.S. Lee, J. Clin. Microbiol. 43, 4836–4839 (2005)
V.P. Le, Y.C. Jung, K.S. Kang, I. Lim, S.C. Myung, W. Kim, J. Med. Virol. 82, 1065–1070 (2010)
S.G. Lee, S.H. Lee, S.W. Park, C.I. Suh, W.H. Jheong, S. Oh, S.Y. Paik, Virol. J. 8, 260 (2011)
S.G. Lee, W.H. Jheong, C.I. Suh, S.H. Kim, J.B. Lee, Y.S. Jeong, G. Ko, K.L. Jang, G.C. Lee, S.Y. Paik, Appl. Environ. Microbiol. 77, 1466–1474 (2011)
K.S. Park, H.S. Jeong, K.A. Baek, C.G. Lee, S.M. Park, J.S. Park, Y.J. Choi, H.J. Choi, D.S. Cheon, Arch. Virol. 155, 635–641 (2010)
J.W. Park, S.G. Lee, Y.M. Lee, W.H. Jheong, S. Ryu, S.Y. Paik, Virol. J. 8, 167 (2011)
J.S. Yoon, S.G. Lee, S.K. Hong, S.A. Lee, W.H. Jheong, S.S. Oh, M.H. Oh, G.P. Ko, C.H. Lee, S.Y. Paik, J. Clin. Microbiol. 46, 1474–1477 (2008)
S.I. Yun, J.K. Kim, B.H. Song, A.Y. Jeong, Y.M. Jee, C.H. Lee, S.Y. Paik, Y. Koo, I. Jeon, S.J. Byun, Y.M. Lee, Virus Res. 152, 137–152 (2010)
D.J. Allen, R. Noad, D. Samuel, J.J. Gray, P. Roy, M. Iturriza-Gómara, Virol. J. 6, 150 (2009)
T. Kageyama, S. Kojima, M. Shinohara, K. Uchida, S. Fukushi, F.B. Hoshino, N. Takeda, K. Katayama, J. Clin. Microbiol. 41, 1548–1557 (2003)
D.P. Zheng, T. Ando, R.L. Fankhauser, R.S. Beard, R.I. Glass, S.S. Monroe, Virology 346, 312–323 (2006)
K. Ambert-Balay, F. Bon, F. Le Guyader, P. Pothier, E. Kohli, J. Clin. Microbiol. 43, 5179–5186 (2005)
P. Chhabra, A.M. Walimbe, S.D. Chitambar, Virus Res. 147, 242–246 (2010)
S.K. Dey, T.G. Phan, M. Mizuguchia, S. Okitsua, H. Ushijima, Virus Genes 40, 362–364 (2010)
M. Jin, H.P. Xie, Z.J. Duan, N. Liu, Q. Zhang, B.S. Wu, H.Y. Li, W.X. Cheng, S.H. Yang, J.M. Yu, Z.Q. Xu, S.X. Cui, L. Zhu, M. Tan, X. Jiang, Z.Y. Fang, J. Med. Virol. 80, 1997–2004 (2008)
A. Kroneman, H. Vennema, K. Deforche, H.V. Avoort, S. Peñaranda, M.S. Oberste, J. Vinjé, M. Koopmans, J. Clin. Virol. 51, 121–125 (2011)
B. Lopman, H. Vennema, E. Kohli, P. Pothier, A. Sanchez, A. Negredo, J. Buesa, E. Schreier, M. Reacher, D. Brown, J. Gray, M. Iturriza, C. Gallimore, B. Bottiger, K.O. Hedlund, M. Torvén, C.H. von Bonsdorff, L. Maunula, M. Poljsak-Prijatelj, J. Zimsek, G. Reuter, G. Szücs, B. Melegh, L. Svennson, Y. van Duijnhoven, M. Koopmans, Lancet 363, 682–688 (2004)
R.A. Bull, E.T. Tu, C.J. McIver, W.D. Rawlinson, P.A. White, J. Clin. Microbiol. 44, 327–333 (2006)
T.G. Phan, T. Kuroiwa, K. Kaneshi, Y. Ueda, S. Nakaya, S. Nishimura, A. Yamamoto, K. Sugita, T. Nishimura, F. Yagyu, S. Okitsu, W.E. Müller, N. Maneekarn, H. Ushijima, J. Med. Virol. 78, 971–978 (2006)
C.I. Gallimore, M. Iturriza-Gomara, J. Xerry, J. Adigwe, J.J. Gray, Arch. Virol. 152, 1295–1303 (2007)
D.J. Allen, J.J. Gray, C.I. Gallimore, J. Xerry, M. Iturriza-Gómara, PLoS One 3, e1485 (2008)
K. Sdiri-Loulizi, K. Ambert-Balay, H. Gharbi-Khelifi, N. Sakly, M. Hassine, S. Chouchane, M.N. Guediche, P. Pothier, M. Aouni, J. Clin. Microbiol. 47, 421–429 (2009)
D.P. Zheng, M.A. Widdowson, R.I. Glass, J. Vinjé, J. Clin. Microbiol. 48, 168–177 (2010)
D.S. Cheon, J. Lee, K. Lee, S. Lee, K. Park, J. Ahn, Y. Jee, J. Yoon, H. Cho, J. Med. Virol. 73, 439–442 (2004)
G. Lee, C. Lee, J. Microbiol. 46, 319–324 (2008)
L. Li, Y. He, H. Yang, J. Zhu, X. Xu, J. Dong, Y. Zhu, Q. Jin, J. Clin. Microbiol. 43, 3835–3839 (2005)
K. Katayama, H. Shirato-Horikoshi, S. Kojima, T. Kageyama, T. Oka, F. Hoshino, S. Fukushi, M. Shinohara, K. Uchida, Y. Suzuki, T. Gojobori, N. Takeda, Virology 299, 225–239 (2002)
T. Kageyama, M. Shinohara, K. Uchida, S. Fukushi, F.B. Hoshino, S. Kojima, R. Takai, T. Oka, N. Takeda, K. Katayama, J. Clin. Microbiol. 42, 2988–2995 (2004)
N. Jothikumar, J.A. Lowther, K. Henshilwood, D.N. Lees, V.R. Hill, J. Vinjé, Appl. Environ. Microbiol. 71, 1870–1875 (2005)
S. Kojima, T. Kageyama, S. Fukushi, F.B. Hoshino, M. Shinohara, K. Uchida, K. Natori, N. Takeda, K. Katayama, J. Virol. Methods 100, 107–114 (2002)
Acknowledgments
We thank Dr. J.I. Lee (Seoul Metropolitan Research Institute of Public Health & Environment, Korea) for kindly providing the stool samples and co-workers (Mogam Biotechnology Research Institute, Korea) for discussions that helped us to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Gyu-Cheol Lee and Gyoo Seung Jung contributed equally to this study.
Rights and permissions
About this article
Cite this article
Lee, GC., Jung, G.S. & Lee, C.H. Complete genomic sequence analysis of norovirus isolated from South Korea. Virus Genes 45, 225–236 (2012). https://doi.org/10.1007/s11262-012-0779-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-012-0779-9