Abstract
Biosynthesis of sterols is a multistep process in higher plants where the precursor cycloartenol gets converted into functional phytosterols after removal of two methyl groups at C-4 by an enzyme complex involving a sterol C-4 methyl oxidase (SMO). We identified and cloned a cDNA from Artemisia annua designated as AaSMO1 showing similarity to SMO. The cDNA predicted to encode a polytopic protein with characteristic histidine-rich motifs and an ER retrieval signal. GFP-AaSMO1 fusion protein was localized in endoplasmic reticulum of transformed protoplast and onion epidermal cells. AaSMO1 expression was drastically induced upon osmotic/dehydration stress and its promoter showed the presence of abscisic acid responsive element. Transgenic tobacco plants ectopically overexpressing AaSMO1 were raised, and various biochemical and physiological analyses of transgenics revealed increased total sterol, better germination and growth in subsequent generations. They also exhibited reduced sensitivity towards osmotic/dehydration stress which may be attributed to enhanced SMO1 activity. Our studies demonstrated that apart from acting as phytohormones, plant sterols also participate in providing capability to plants for improved growth and adaptation during stress conditions. AaSMO1 can be used as an excellent candidate for generating dehydration/drought tolerant plants.






Similar content being viewed by others
Abbreviations
- 4-MU:
-
4-Methylumbelliferone
- ABA:
-
Abscisic acid
- ABRE:
-
Abscisic acid responsive element
- DRE/CRT:
-
Dehydration-responsive element/C-RepeaT
- ER:
-
Endoplasmic reticulum
- GA:
-
Gibberellic acid
- GFP:
-
Green fluorescent protein
- GUS:
-
β-Glucuronidase
- MeJa:
-
Methyl jasmonate
- MS:
-
Murashige and Skoog medium
- MUG:
-
4-Methylumbelliferyl glucuronide
- RT-qPCR:
-
Reverse transcription quantitative real-time PCR
- SMO:
-
Sterol-4α-methyl-oxidase/sterol C-4 methyl oxidase
- TCA:
-
Trans-cinnamic acid
- TLC:
-
Thin layer chromatography
References
Bard M, Bruner DA, Pierson CA, Lees ND, Biermann B, Frye L, Koegel C, Barbuch R (1996) Cloning and characterization of ERG25, the Saccharomyces cerevisiae gene encoding C-4 sterol methyl oxidase. Proc Natl Acad Sci USA 93:186–190
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Bishop GJ, Yokota T (2001) Plants steroid hormones, brassinosteroids: current highlights of molecular aspects on their synthesis/metabolism, transport, perception and response. Plant Cell Physiol 42:114–120
Borsani O, Cuartero J, Valpuesta V, Botella MA (2002) Tomato tos1 mutation identifies a gene essential for osmotic tolerance and abscisic acid sensitivity. Plant J 32:905–914
Cai R, Zhao Y, Wang Y, Lin Y, Peng X, Li Q, Chang Y, Jiang H, Xiang Y, Cheng B (2014) Overexpression of a maize WRKY58 gene enhances drought and salt tolerance in transgenic rice. Plant Cell Tiss Organ Cult 119:565–577
Chappell J, Wolf F, Proulx J, Cuellar R, Saunders C (1995) Is the reaction catalyzed by 3-hydroxy-3-methylglutaryl coenzyme A reductase a rate-limiting step for isoprenoid biosynthesis in plants? Plant Physiol 109:1337–1343
Citovsky V, Lee LY, Vyas S, Glick E, Chen MH, Vainstein A, Gafni Y, Gelvin SB, Tzfira T (2006) Subcellular localization of interacting proteins by bimolecular fluorescence complementation in planta. J Mol Biol 362:1120–1131
Darnet S, Rahier A (2004) Plant sterol biosynthesis: identification of two distinct families of sterol 4α-methyl oxidases. Biochem J 378:889–898
Darnet S, Bard M, Rahier A (2001) Functional identification of sterol-4α-methyl oxidase cDNAs from Arabidopsis thaliana by complementation of a yeast erg25 mutant lacking sterol-4α-methyl oxidation. FEBS Lett 508:39–43
Gachotte D, Barbuch R, Gaylor J, Nickel E, Bard M (1998) Characterization of the Saccharomyces cerevisiae ERG26 gene encoding the C-3 sterol dehydrogenase (C-4 decarboxylase) involved in sterol biosynthesis. Proc Natl Acad Sci USA 95:13794–13799
Gondet L, Weber T, Maillot-Vernier P, Benveniste P, Bach T (1992) Regulatory role of microsomal 3-hydroxy-3-methylglutaryl-coenzyme A reductase in a tobacco mutant that overproduces sterols. Biochem Biophys Res Commun 186:888–893
Guo P, Baum M, Grando S, Ceccarelli S, Bai G, Li R, von Korff M, Varshney RK, Graner A, Valkoun J (2009) Differentially expressed genes between drought-tolerant and drought-sensitive barley genotypes in response to drought stress during the reproductive stage. J Exp Bot 60:3531–3544
Holmberg N, Harker M, Gibbard CL, Wallace AD, Clayton JC, Rawlins S, Hellyer A, Safford R (2002) Sterol C-24 methyltransferase type 1 controls the flux of carbon into sterol biosynthesis in tobacco seed. Plant Physiol 130:303–311
Holmberg N, Harker M, Wallace AD, Clayton JC, Gibbard CL, Safford R (2003) Co-expression of N-terminal truncated 3-hydroxy-3-methylglutaryl CoA reductase and C24-sterol methyltransferse type 1 in transgenic tobacco enhances carbon flux towards end-product sterols. Plant J 36:12–20
Horsch RB, Fry JE, Hoffmann NL, Eichholtz D, Rogers SG, Farley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231
Jackson MR, Nilsson T, Peterson PA (1990) Identification of a consensus motif for retention of transmembrane proteins in the endoplasmic reticulum. EMBO J 9:3153–3162
Jang JC, Fujioka S, Tasaka M, Seto H, Takatsuto S, Ishii A, Aida M, Yoshida S, Sheen J (2000) A critical role of sterols in embryonic patterning and meristem programming revealed by the fackel mutants of Arabidopsis thaliana. Genes Dev 14:1485–1497
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 6:3901–3907
Jiang CJ, Shoji K, Matsuki R, Baba A, Inagaki N, Ban H, Iwasaki T, Imamoto N, Yoneda Y, Deng XW, Yamamoto N (2001) Molecular cloning of a novel importin alpha homologue from rice, by which constitutive photomorphogenic 1 (COP1) nuclear localization signal (NLS)-protein is preferentially nuclear imported. J Biol Chem 276:9322–9329
Jiang X, Zhang C, Lu P, Jiang G, Liu X, Dai F, Gao J (2014) RhNAC3, a stress-associated NAC transcription factor, has a role in dehydration tolerance through regulating osmotic stress-related genes in rose petals. Plant Biotechnol J 12:38–48
Josekutty PC (1998) Regulation by HMGR of sterol biosynthesis in a selected high sterol cell line of Solanum xanthocarpum Shrader & Wendl. Phyton-Int J Exp Bot 63:123–128
Kim T, Bohmer M, Hu H, Nishimura N, Schroeder J (2010) Guard cell signal transduction network: advances in understanding abscisic acid, CO2, and Ca2+ signaling. Annu Rev Plant Biol 61:561–591
Kishor KPB, Hong Z, Miao GH, Hu CAA, Verma DPS (1995) Overexpression of D1-pyrroline-5-carboxylate synthetase increases proline production and confers osmotolerance in transgenic plants. Plant Physiol 108:1387–1394
Lee M, Lenman M, Banaś A, Bafor M, Singh S, Schweizer M, Nilsson R, Liljenberg C, Dahlqvist A, Gummeson PO, Sjödahl S, Green A, Stymne S (1998) Identification of non-heme diiron proteins that catalyze triple bond and epoxy group formation. Science 280:915–918
Li L, Kaplan J (1996) Characterization of yeast methyl sterol oxidase (ERG25) and identification of a human homologue. J Biol Chem 271:16927–16933
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148:350–382
Lindsey K, Pullen ML, Topping JF (2003) Importance of plant sterols in pattern formation and hormone signalling. Trends Plant Sci 8:521–525
Lugan R, Niogret MF, Leport L, Gue´gan JP, Larher FR, Savoure´ A, Kopka J, Bouchereau A (2010) Metabolome and water homeostasis analysis of Thellungiella salsuginea suggests that dehydration tolerance is a key response to osmotic stress in this halophyte. Plant J 64:215–229
Mao X, Zhang H, Tian S, Chang X, Jing R (2009) TaSnRK2.4, an SNF1-type serine-threonine protein kinase of wheat (Triticum aestivum L.) confers enhanced multi-stress tolerance in Arabidopsis. J Exp Bot 61:683–696
Nair P, Misra A, Singh A, Shukla AK, Gupta MM, Gupta AK, Gupta V, Khanuja SP, Shasany AK (2013) Differentially expressed genes during contrasting growth stages of Artemisia annua for artemisinin content. PLoS ONE 8:e60375
Nelson BK, Cai X, Nebenführ A (2007) A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J 51:1126–1136
Pascal S, Taton M, Rahier A (1994) Plant sterol biosynthesis: identification of a NADPH dependent sterone reductase involved in sterol-4 demethylation. Arch Biochem Biophys 312:260–271
Pose´ D, Castanedo I, Borsani O, Nieto B, Rosado A, Taconnat L, Ferrer A, Dolan L, Valpuesta V, Botella MA (2009) Identification of the Arabidopsis dry2/sqe1-5 mutant reveals a central role for sterols in drought tolerance and regulation of reactive oxygen species. Plant J 59:63–76
Rahier A (2011) Dissecting the sterol C-4 demethylation process in higher plants. From structures and genes to catalytic mechanism. Steroids 76:340–352
Rahier A, Smith M, Taton M (1997) The role of cytochrome b5 in 4alpha-methyl-oxidation and C5(6) desaturation of plant sterol precursors. Biochem Biophys Res Commun 236:434–437
Rajakani R, Narnoliya L, Sangwan NS, Sangwan RS, Gupta V (2014) Subtractive transcriptomes of fruit and leaf reveal differential representation of transcripts in Azadirachta indica. Tree Genet Genomes 10:1331–1351
Rondet S, Taton M, Rahier A (1999) Identification, characterization, and partial purification of 4 alpha-carboxysterol-C3-dehydrogenase/C4-decarboxylase from Zea mays. Arch Biochem Biophys 366:249–260
Schaller H, Grausem B, Benveniste P, Chey ML, Tan CT, Song YH, Chua NH (1995) Expression of the Hevea brasiliensis (H.B.K.) Mull. Arg. 3-hydroxy-3-methylglutaryl coenzymes A reductase 1 in tobacco results in sterol overproduction. Plant Physiol 109:761–770
Schrick K, Mayer U, Horrichs A, Kuhnt C, Bellini C, Dangl J, Schmidt J, Jürgens G (2000) FACKEL is a sterol C-14 reductase required for organized cell division and expansion in Arabidopsis embryogenesis. Genes Dev 14:1471–1484
Seki M, Kamei A, Yamaguchi-Shinozaki K, Shinozaki K (2003) Molecular responses to drought, salinity and frost: common and different paths for plant protection. Curr Opin Biotechnol 14:194–199
Shanklin J, Whittle E, Fox BG (1994) Eight histidine residues are catalytically essential in a membrane-associated iron enzyme, stearoyl-CoA desaturase, and are conserved in alkane hydroxylase and xylene monooxygenase. Biochemistry 33:12787–12794
Sharma R, Sahoo A, Devendran R, Jain M (2014) Over-expression of a rice tau class glutathione S-transferase gene improves tolerance to salinity and oxidative stresses in Arabidopsis. PLoS ONE 9:e92900
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227
Silva EN, Ferreira-Silva SL, Fontenele Ade V, Ribeiro RV, Viégas RA, Silveira JA (2010) Photosynthetic changes and protective mechanisms against oxidative damage subjected to isolated and combined drought and heat stresses in Jatropha curcas plants. J Plant Physiol 167:1157–1164
Souter M, Topping J, Pullen M, Friml J, Palme K, Hackett R, Grierson D, Lindsey K (2002) Hydra mutants of Arabidopsis are defective in sterol profiles and auxin and ethylene signaling. Plant Cell 14:1017–1031
Taton M, Husselstein T, Benveniste P, Rahier A (2000) Role of highly conserved residues in the reaction catalyzed by recombinant ∆7-sterol-C5(6)-desaturase studied by site-directed mutagenesis. Biochemistry 39:701–711
Tusnády GE, Simon I (2001) The HMMTOP transmembrane topology prediction server. Bioinformatics 17:849–850
Tuteja N (2010) A method to confer salinity stress tolerance to plants by helicase overexpression. Methods Mol Biol 587:377–387
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97:11632–11637
Verpoorte R, Memelink J (2002) Engineering secondary metabolite production in plants. Curr Opin Biotechnol 13:181–187
Vivek PJ, Tuteja N, Soniya EV (2013) CDPK1 from ginger promotes salinity and drought stress tolerance without yield penalty by improving growth and photosynthesis in Nicotiana tabacum. PLoS ONE 8:e76392
Wang FQ, Zhao Y, Dai M, Liu J, Zheng GZ, Ren ZH, He JG (2008) Cloning and functional identification of C-4 methyl sterol oxidase genes from the penicillin-producing fungus Penicillium chrysogenum. FEMS Microbiol Lett 287:91–99
Wasternack C (2007) Jasmonates: an update on biosynthesis, signal transduction and action in plant stress response, growth and development. Ann Bot 100:681–697
Xiang Y, Huang YM, Xiong LZ (2007) Characterization of stress responsive CIPK genes in rice for stress tolerance improvement. Plant Physiol 144:1416–1428
Yang G, Wang Y, Xia D, Gao C, Wang C, Yang C (2014) Overexpression of a GST gene (ThGSTZ1) from Tamarix hispida improves drought and salinity tolerance by enhancing the ability to scavenge reactive oxygen species. Plant Cell Tiss Organ Cult 117:99–112
Yoo SD, Cho YH, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572
Yuan Y, Qi L, Yang J, Wu C, Liu Y, Huang L (2014) A Scutellaria baicalensis R2R3-MYB gene, SbMYB8, regulates flavonoid biosynthesis and improves drought stress tolerance in transgenic tobacco. Plant Cell Tiss Organ Cult. doi:10.1007/s11240-014-0650-x
Zhai S, Gao Q, Liu X, Sui Z, Zhang J (2013) Overexpression of a Zea mays phospholipase C1 gene enhances drought tolerance in tobacco in part by maintaining stability in the membrane lipid composition. Plant Cell Tiss Organ Cult 115:253–262
Acknowledgments
The initial work was supported by CSIR, Govt. of India, under NWP08 project, and further funding was provided by SERB, DST, Govt. of India under GAP265 project. Research fellowships from CSIR and UGC, Govt. of India, are acknowledged. The authors wish to express their sincere thanks to the Director, CSIR-CIMAP for providing necessary facilities. Help from Drs. R. K. Shukla, A. K. Shasany, A. S. Negi and Ms. Pooja Sharma is duly acknowledged. Dr. Samir Sawant from CSIR-NBRI, Lucknow and Dr. Jitendra Thakur from NIPGR, New Delhi, are acknowledged for extending their help in confocal microscopy.
Conflict of interest
The authors declare no competing interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11240_2014_692_MOESM1_ESM.ppt
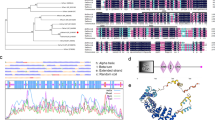
Supplemental Fig. S1 Comparative and phylogenetic analyses of AaSMO1. a Predicted amino acid sequence was aligned with selected sterol C-4 methyl oxidases reported in NCBI database by using ClustalW. Identical and similar amino acid residues are shaded in black and grey, respectively. Dashes denote gaps inserted to maximize the alignment. Signature characteristic motifs (domain I, II and III) of sterol C-4 methyl oxidases are boxed and ER retrieval signal is underlined. b Phylogenetic relationship (unrooted neighbor-joining tree) of AaSMO1 with related homologs from other plant species. The sequences used for alignment or phylogenetic analysis in ‘a’ and ‘b’ include: A. thaliana (CAB78254; NP_567670; NP_001077975; AAM65428; AAM64961; NP_567669; NP_850133; CAA62079; AAQ13424), A. tripolium (BAC57961), Brachypodium distachyon (XP_003574284), Glycine max (NP_001276171), Gossypium arboreum (AAO13795), Hordeum vulgare (BAK02768; BAJ99813), Komagataella pastoris/Pichia pastoris (XP_002492261), Medicago truncatula (XP_003608797; XP_003613314), N. benthamiana (AAQ83691), N. tabacum (AAD04034; AAD20458), Oryza sativa (AAN62786), P. trichocarpa (XP_002304440), Ricinus communis (EEF51010; XP_002509623), S. cerevisiae (EDN61653; P32353; AAA16608), Sorghum bicolor (XP_002455692), V. vinifera (XP_002282305; XP_002282653; XP_002270978), Z. mays (NP_001105744; NP_001148435; NP_001148698) and A. annua (AaSMO1; GQ847864). (PPT 565 kb)
11240_2014_692_MOESM2_ESM.ppt
Supplemental Fig. S2 Kyte-Doolittle hydrophobicity plot of AaSMO1. Amino acid regions with values above zero are hydrophobic in nature. The horizontal scale indicates the position of amino acids. Hydrophobic regions identified in the plot are encircled. (PPT 152 kb)
11240_2014_692_MOESM3_ESM.doc
Supplemental Fig. S3 In silico docking of AaSMO1 with 24-methylene cycloartanol and 24-ethylidene lophenol (i.e. substrates of SMO1 and SMO2, respectively) by homology modelling, and its comparison with that of the binding affinity of AtSMO1-1 isoform. (DOC 701 kb)
Rights and permissions
About this article
Cite this article
Singh, A., Jindal, S., Longchar, B. et al. Overexpression of Artemisia annua sterol C-4 methyl oxidase gene, AaSMO1, enhances total sterols and improves tolerance to dehydration stress in tobacco. Plant Cell Tiss Organ Cult 121, 167–181 (2015). https://doi.org/10.1007/s11240-014-0692-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-014-0692-0