Abstract
Height, an important agronomic trait, relates to lodging resistance, harvest index, and yield potential. Height is also a major issue in developmental biology. Systematic identification and characterization of dwarf plants that show defects in height have basic and applied significance. In previous research, we analyzed the agronomic and physiological features of the dominant maize dwarf plant Dwarf11 (D11). However, transcriptomic specialization of D11 remains largely unknown. Here, we combined RNA-seq and co-expression analysis to survey the transcriptome dynamics of D11. A total of 306 significantly differentially expressed genes (DEGs) were identified, including 171 upregulated and 135 downregulated DEGs. Functional annotation, gene ontology (GO), and pathway enrichment analysis indicated that roles of DEGs were diversified, from organ size control, cell wall character, vascular tissue development, leaf morphology, chlorophyll metabolism, chloroplast function to sugar homeostasis. Functional associations of DEGs were further revealed by the co-expression analysis. Results presented here are informative for the elucidation of D11-involved regulatory network, and lay a foundation for candidate gene prioritization and D11 gene cloning.




Similar content being viewed by others
References
Cassani E, Bertolini E, Badone FC, Landoni M, Gavina D, Sirizzotti A, Pilu R (2009) Characterization of the first dominant dwarf maize mutant carrying a single amino acid insertion in the VHYNP domain of the dwarf8 gene. Mol Breeding 24:375–385
Conesa A, Madrigal P, Tarazona S, Gomez-Cabrero D, Cervera A, McPherson A, Szcześniak MW, Gaffney DJ, Elo LL, Zhang X, Mortazavi A (2016) A survey of best practices for RNA-seq data analysis. Genome Biol 17:13
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70
Eisen MB, Spellman PT, Brown PO, Botstein D (1998) Cluster analysis and display of genome-wide expression patterns. Proc Natl Acad Sci U S A 95:14863–14868
Fujioka S, Yamane H, Spray CR, Gaskin P, Macmillan J, Phinney BO, Takahashi N (1988a) Qualitative and quantitative analyses of gibberellins in vegetative shoots of normal, dwarf-1, dwarf-2, dwarf-3, and dwarf-5 seedlings of Zea mays L. Plant Physiol 88:1367–1372
Fujioka S, Yamane H, Spray CR, Katsumi M, Phinney BO, Gaskin P, Macmillan J, Takahashi N (1988b) The dominant non-gibberellin-responding dwarf mutant (D8) of maize accumulates native gibberellins. Proc Natl Acad Sci U S A 85:9031–9035
Gene Ontology Consortium (2001) Creating the gene ontology resource: design and implementation. Genome Res 11:1425–1433
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M (2012) KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40:D109–D114
Kawahara Y, de la Bastide M, Hamilton JP, Kanamori H, McCombie WR, Ouyang S, Schwartz DC, Tanaka T, Wu J, Zhou S, Childs KL, Davidson RM, Lin H, Quesada-Ocampo L, Vaillancourt B, Sakai H, Lee SS, Kim J, Numa H, Itoh T, Buell CR, Matsumoto T (2013) Improvement of the Oryza sativa Nipponbare reference genome using next generation sequence and optical map data. Rice 6:4
Khush GS (2001) Green revolution: the way forward. Nat Rev Genet 2:815–822
Kir G, Ye H, Nelissen H, Neelakandan AK, Kusnandar AS, Luo A, Inzé D, Sylvester AW, Yin Y, Becraft PW (2015) RNA interference knockdown of BRASSINOSTEROID INSENSITIVE1 in maize reveals novel functions for brassinosteroid signaling in controlling plant architecture. Plant Physiol 169:826–839
Krishnakumar V, Hanlon MR, Contrino S, Ferlanti ES, Karamycheva S, Kim M, Rosen BD, Cheng CY, Moreira W, Mock SA, Stubbs J, Sullivan JM, Krampis K, Miller JR, Micklem G, Vaughn M, Town CD (2015) Araport: the Arabidopsis information portal. Nucleic Acids Res 43:D1003–D1009
Lawit SJ, Wych HM, Xu D, Kundu S, Tomes DT (2010) Maize DELLA proteins dwarf plant8 and dwarf plant9 as modulators of plant development. Plant Cell Physiol 51:1854–1868
Li R, Yu C, Li Y, Lam TW, Yiu SM, Kristiansen K, Wang J (2009) SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25:1966–1967
Li QC, Li YX, Yang ZZ, Liu C, Liu ZZ, Li CH, Peng B, Zhang Y, Wang D, Tan WW, Sun BC, Shi YS, Song YC, Zhang ZM, Pan GT, Wang TY, Li Y (2013) QTL mapping for plant height and ear height by using multiple related RIL populations in maize. Acta Agron Sin 39:1521–1529
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25:402–408
Makarevitch I, Thompson A, Muehlbauer GJ, Springer NM (2012) Brd1 gene in maize encodes a brassinosteroid C-6 oxidase. PLoS One 7:e30798
Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M, Sato M, Nasu S, Minobe Y (2002) Positional cloning of rice semidwarfing gene, sd-1: rice "green revolution gene" encodes a mutant enzyme involved in gibberellin synthesis. DNA Res 9:11–17
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628
Multani DS, Briggs SP, Chamberlin MA, Blakeslee JJ, Murphy AS, Johal GS (2003) Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 302:81–84
Peiffer JA, Romay MC, Gore MA, Flint-Garcia SA, Zhang Z, Millard MJ, Gardner CA, McMullen MD, Holland JB, Bradbury PJ, Buckler ES (2014) The genetic architecture of maize height. Genetics 196:1337–1356
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) Green revolution' genes encode mutant gibberellin response modulators. Nature 400:256–261
Roberts A, Pimentel H, Trapnell C, Pachter L (2011) Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics 27:2325–2329
de Saint GA, Ligerot Y, Dun EA, Pillot JP, Ross JJ, Beveridge CA, Rameau C (2013) Strigolactones stimulate internode elongation independently of gibberellins. Plant Physiol 163:1012–1025
Saldanha AJ (2004) Java Treeview—extensible visualization of microarray data. Bioinformatics 20:3246–3248
Spielmeyer W, Ellis MH, Chandler PM (2002) Semidwarf (sd-1), "green revolution" rice, contains a defective gibberellin 20-oxidase gene. Proc Natl Acad Sci U S A 99:9043–9048
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, Simonovic M, Roth A, Santos A, Tsafou KP, Kuhn M, Bork P, Jensen LJ, von Mering C (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43:D447–D452
Tello-Ruiz MK, Stein J, Wei S, Preece J, Olson A, Naithani S, Amarasinghe V, Dharmawardhana P, Jiao Y, Mulvaney J, Kumari S, Chougule K, Elser J, Wang B, Thomason J, Bolser DM, Kerhornou A, Walts B, Fonseca NA, Huerta L, Keays M, Tang YA, Parkinson H, Fabregat A, McKay S, Weiser J, D'Eustachio P, Stein L, Petryszak R, Kersey PJ, Jaiswal P, Ware D (2016) Gramene 2016: comparative plant genomics and pathway resources. Nucleic Acids Res 44:D1133–D1140
Teng F, Zhai L, Liu R, Bai W, Wang L, Huo D, Tao Y, Zheng Y, Zhang Z (2013) ZmGA3ox2, a candidate gene for a major QTL, qPH3.1, for plant height in maize. Plant J 73:405–416
Thyssen GN, Fang DD, Turley RB, Florane C, Li P, Naoumkina M (2015) Mapping-by-sequencing of Ligon-lintless-1 (Li 1 ) reveals a cluster of neighboring genes with correlated expression in developing fibers of upland cotton (Gossypium hirsutum L.) Theor Appl Genet 128:1703–1712
Wang Y, Deng D (2014) Molecular basis and evolutionary pattern of GA-GID1-DELLA regulatory module. Mol Gen Genomics 289:1–9
Wang Y, Li J (2005) The plant architecture of rice (Oryza sativa). Plant Mol Biol 59:75–84
Wang Y, Deng D, Ding H, Xu X, Zhang R, Wang S, Bian Y, Yin Z, Chen Y (2013) Gibberellin biosynthetic deficiency is responsible for maize dominant Dwarf11 (D11) mutant phenotype: physiological and transcriptomic evidence. PLoS One 8:e66466
Wang Y, Lu W, Chen Y, Deng D, Ding H, Bian Y, Yin Z, Zhu Y, Zhao J (2016a) Revealing physiological and genetic properties of a dominant maize dwarf Dwarf11 (D11) by integrative analysis. Mol Breeding 36:31
Wang Y, Lu W, Deng D (2016b) Bioinformatic landscapes for plant transcription factor system research. Planta 243:297–304
Weng J, Xie C, Hao Z, Wang J, Liu C, Li M, Zhang D, Bai L, Zhang S, Li X (2011) Genome-wide association study identifies candidate genes that affect plant height in Chinese elite maize (Zea mays L.) inbred lines. PLoS One 6:e29229
Xiao Y, Thatcher S, Wang M, Wang T, Beatty M, Zastrow-Hayes G, Li L, Li J, Li B, Yang X (2016) Transcriptome analysis of near-isogenic lines provides molecular insights into starch biosynthesis in maize kernel. J Integr Plant Biol 58:713–723
Xing A, Gao Y, Ye L, Zhang W, Cai L, Ching A, Llaca V, Johnson B, Liu L, Yang X, Kang D, Yan J, Li J (2015) A rare SNP mutation in Brachytic2 moderately reduces plant height and increases yield potential in maize. J Exp Bot 66:3791–3802
Zhang S, Liu F, Liu B, Wang L, Dong S (2007) Discovery of a new dominant dwarf gene in maize and its preliminary study. J Maize Sci 15:15–18
Zheng DB, Yang XH, Li JS, Yan JB, Zhang SL, He ZH, Huang YQ (2013) QTL identification for plant height and ear height based on SNP mapping in maize (Zea mays L.) Acta Agron Sin 39:549–556
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31571671 and 31201213), the National Key Research and Development Program of China (2016YFD0101002), the Priority Academic Program Development of Jiangsu Higher Education Institutions, and the Jiangsu Government Scholarship for Overseas Studies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Yijun Wang and Wenjie Lu contributed equally to this work.
Electronic supplementary material
Figure S1
Correlations between biological replicates. Biological replicates showed high correlations (r > 0.94) in gene expression (JPEG 638 kb)
Figure S2
Gene ontology analysis of up-regulated differentially expressed genes (JPEG 105 kb)
Figure S3
Gene ontology analysis of down-regulated differentially expressed genes (JPEG 429 kb)
Figure S4
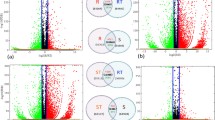
Shared differentially expressed genes between Zheng 58 and Mo17 populations (JPEG 102 kb)
Table S1
Primers used in this study (DOCX 17 kb)
Table S2
Sample statistics for RNA-seq and mapping (DOCX 15 kb)
Table S3
Information of all expressed genes in biological replicates of the wild-type and D11 (TXT 14557 kb)
Table S4
Information of differentially expressed genes in D11 relative to the wild-type (XLSX 66 kb)
Table S5
Differentially expressed genes revealed by RNA-seq and qRT-PCR (DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Lu, W., Zhao, J. et al. Transcriptome Dynamics of Dominant Maize Dwarf Dwarf11 (D11) Revealed by RNA-seq and Co-expression Analysis. Plant Mol Biol Rep 35, 355–365 (2017). https://doi.org/10.1007/s11105-017-1028-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-017-1028-0