Abstract
The differential expression profiling of the fruit-specific genes between a blood orange (Citrus sinensis cv. Tarocco) and a blonde orange (Citrus sinensis cv. twenty-first century navel orange) was investigated by fruit transcriptome sequencing. The relative expression folds for Cs6g17570 and Cs5g31400 were significantly greater in the “Tarocco” fruits, but Cs9g04810 transcripts had no significant differences between two accessions by digital gene expression profiling. In particular, R2R3-MYB, bHLH-type, and WD40-repeat proteins (encoded by Cs6g17570, Cs5g31400, and Cs9g04810) were transcription factors with each conserved domain and interacted in vitro and in vivo. Moreover, the dihydroflavonol 4-reductase (DFR), a key enzyme involved into anthocyanin biosynthesis pathway, had greater transcription levels in Tarocco fruit and also in the transgenic Arabidopsis thaliana by Cs6g17570 ectopic expression. Furthermore, Cs6g17570 and Cs5g31400 in Tarroco were upregulated by cold induction and then had similar expression profiling during cold storage. In conclusion, Cs6g17570, Cs5g31400, and Cs9g04810 encoding proteins might be involved into anthocyanin regulation in Tarocco by MYB-bHLH-WD40 complex as well as in other fruit trees. Studies on the anthocyanin regulatory mechanism will aid in development of new biotechnological tools for Citrus breeding.









Similar content being viewed by others
References
An XH, Tian Y, Chen KQ, Wang XF, Hao YJ (2012) The apple WD40 protein MdTTG1 interacts with bHLH but not MYB protein to regulate anthocyanin accumulation. J Plant Physiol 169:710–717
Ben-Simhon Z, Judeinstein S, Nadler-Hassar T, Trainin T, Bar-Ya'akov I, Borochov-Neori H, Holland D (2011) A pomegranate (Punica granatum L.) WD40-repeat gene is a functional homologue of Arabidopsis TTG1 and is involved in the regulation of anthocyanin biosynthesis during pomegranate fruit development. Planta 234(5):865–881
Butelli E, Licciardello C, Zhang Y, Liu J, Mackay S, Bailey P (2012) Retrotransposons control fruit-specific, cold-dependent accumulation of anthocyanins in blood oranges. Plant Cell 24:1242–1255
Chen H, Zou Y, Shang Y (2008) Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol 146:368–376
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2gO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676
Crifo T, Petrone G, Lo Cicero L, Lo Piero AR (2012) Short cold storage enhances the anthocyanin contents and level of transcripts related to their biosynthesis in blood oranges. J Agric Food Chem 60(1):476–81
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Cultrone AA, Cotroneo PS, Recupero GR (2010) Cloning and molecular characterization of R2R3-MYB and bHLH-MYC transcription factors from Citrus sinensis. Tree Genet Genomes 6:101–112
Cotroneo PS, Russo MP, Ciuni M, Recupero GR, Lo Piero AR (2006) Quantitative real-time reverse transcriptase-PCR profiling of anthocyanin biosynthetic genes during orange fruit ripening. J Amer Soc Hort Sci 131(4):537–547
Dong S, Liu Y, Niu J, Ning Y, Lin S, Zhang Z (2014) De novo transcriptome analysis of the Siberian apricot (Prunus sibirica L.) and search for potential SSR markers by 454 pyrosequencing. Gene 544:220–227
Gonzalez A, Zhao M, Leavitt JM, Alan ML (2008) Regulation of the anthocyanin biosynthetic pathway by the TTG1/bHLH/Myb transcriptional complex in Arabidopsis seedlings. Plant J 53:814–827
He J, Chen KL, Liu JJ, Guan B, Li HW, Wang JH (2013) Effect of liquid fertilizer by injecting into on quality of Tarocco blood orange. Southwest China J Agric Sci 26:299–303
Heppel SC, Jaffe FW, Takos AM, Schellmann S, Rausch T, Walker AR, Bogs J (2013) Identification of key amino acids for the evolution of promoter target specificity of anthocyanin and proanthocyanidin regulating MYB factors. Plant Mol Biol 82(4-5):457–71
Hillebrand S, Schwarz M, Winterhalter P (2004) Characterization of anthocyanins and pyranoanthocyanins from blood orange [Citrus sinensis (L.) osbeck] juice. J Agric Food Chem 52:7331–7338
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1071–1083
Li S (2014) Transcriptional control of flavonoid biosynthesis: fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Signal Behav 9(1):e27522
Licciardello C, Russo MP, Valè G, Reforgiato Recupero G (2008) Identification of differentially expressed genes in the flesh of blood and common oranges. Tree Genet Genomes 4:315–31
Liang M, Yang X, Li H, Su S, Yi H, Chai L, Deng X (2015) De novo transcriptome assembly of pummelo and molecular marker development. PLoS One 10(3):e0120615. doi:10.1371/journal.pone.0120615
Lo Piero AR, Puglisi I, Rapisarda P, Petrone G (2005) Anthocyanins accumulation and related gene expression in red orange fruit induced by low temperature storage. J Agric Food Chem 53:9083–9088
Luo C, Wu HX, Yao QS, Wang SB, Xu WT (2015) Development of EST-SSR and TRAP markers from transcriptome sequencing data of the mango. Genet Mol Res 14(3):7914–7919
Lo Piero AR, Puglisi I, Petrone G (2006) Gene isolation, analysis of expression, and in vitro synthesis of glutathione s-transferase from orange fruit [Citrus sinensis L. (Osbeck)]. J Agric Food Chem 54(24):9227–9233
Moriguchi T, Kita M, Tomono Y, Endo-Inagaki T, Omura M (1999) One type of chalcone synthase gene expressed during embryogenesis regulates the flavonoid accumulation in Citrus cell cultures. Plant Cell Physiol 40:651–65
Muccilli V, Licciardello C, Fontanini D, Patrizia Russo M, Cunsolo V, Saletti R, Reforgiato Recupero G, Foti S (2009) Proteome analysis of Citrus sinensis L. (Osbeck) flesh at ripening time. J Proteomics 73(1):134–152
Pollier J, Rombauts S, Goossens A (2013) Analysis of RNA-Seq data with TopHat and Cufflinks for genome-wide expression analysis of jasmonate-treated plants and plant cultures. Methods Mol Biol 1011:305–315
Rahim MA, Busatto N, Trainotti L (2014) Regulation of anthocyanin biosynthesis in peach fruits. Planta 240(5):913–29
Rapisarda P, Fanella F, Maccarone E (2000) Reliability of analytical methods for determining anthocyanins in blood orange juices. J Agric Food Chem 48:2249–2252
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, Trush V, Quackenbush J (2003) TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34(2):374–378
Schaart JG, Dubos C, Fuente IRDL, Houwelingen AMML, Vos RCH, Jonker HH, Xu WJ, Routaboul JM, Lepiniec L, Bovy AG (2013) Identification and characterization of MYB-bHLH-WD40 regulatory complexes controlling proanthocyanidin biosynthesis in strawberry fruits. New Phytologist 197:454–467
Szklarczyk D, Franceschini A, Wyder S, Forslund K, Heller D, Huerta-Cepas J, Simonovic M, Roth A, Santos A, Tsafou KP, Kuhn M, Bork P, Jensen LJ, von Mering C (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43:447–452. doi:10.1093/nar/gku1003
Tang Q, Ma X, Mo C, Wilson IW, Song C, Zhao H, Yang Y, FuW QD (2011) An efficient approach to finding Siraitia grosvenorii triterpene biosynthetic genes by RNA-seq and digital gene expression analysis. BMC Genomics 12:343
Wei BY, Zhang JZ, Pang CX, Yu H, Guo DS, Jiang H, Ding MX, Chen ZY, Tao Q, Gu HY, Qu LJ, Qin GJ (2015) The molecular mechanism of SPOROCYTELESS/NOZZLE in controlling Arabidopsis ovule development. Cell Res 25:121–134
Xu Q, Chen LL, Ruan XA, Chen DJ, Zhu AD, Chen CL, Bertrand D, Jiao WB, Hao BH, Matthew PL (2013) The draft genome of sweet orange (Citrus sinensis). Nat Genet 45(1):59–66
Zhang QJ, Tao ST, Li M, Qi XX, Wu J, Yin H, Deng JL, Zhang SL (2015) Identification of differentially expressed genes using digital gene expression profiles in Pyrus pyrifolia Nakai cv. Hosui bud release following early defoliation. Tree Genet Genomes 11:34. doi:10.1007/s11295-015-0858-x
Acknowledgement
This work was supported by the Applied and Basic Research Project of Sichuan Province.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
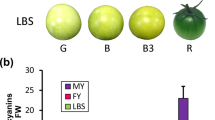
Fruit quality analysis between two accessions. 1, 2 indicated Tarocco and twenty-first century navel orange, respectively. The total acidity (a), anthocyanin contents (b), and total soluble solids (c) were determined. The key genes involved into the anthocyanin biosynthesis and chalcone synthase, were expressed differentially between two accessions (d) (JPG 380 kb)
Rights and permissions
About this article
Cite this article
Wang, Jh., Liu, Jj., Chen, Kl. et al. Anthocyanin Biosynthesis Regulation in the Fruit of Citrus sinensis cv. Tarocco. Plant Mol Biol Rep 34, 1043–1055 (2016). https://doi.org/10.1007/s11105-016-0984-0
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-016-0984-0