Abstract
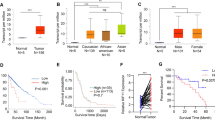
Kinesin family member 20A (KIF20A), an ideal cancer-testis antigen, was reported to be a promising immunotherapeutic target for pancreatic cancers. Clinical trials of KIF20A peptide vaccine immunotherapy have been conducted against pancreatic cancers. To demonstrate the efficacy of KIF20A as a candidate molecular target for gliomas, we analyzed the expression and function of KIF20A in gliomas. Western blot and quantitative PCR analyses showed that KIF20A expression in glioma cell lines and glioma tissues was high compared with that found in a normal brain. KIF20A immunostaining of glioma cells and glioma tissues demonstrated that KIF20A was involved in spindle formation and cytokinesis, and that KIF20A was highly expressed, especially in glioma cells undergoing mitosis. In silico analysis of a cancer microarray database revealed that KIF20A was highly expressed in gliomas depending on the pathological grade, and glioma patients with higher expression of KIF20A showed poorer prognosis. Down-regulating KIF20A reduced cell proliferation in glioma cells due to the failure of cytokinesis and generation of binucleated cells. Additionally, KIF20A inhibition induced significant apoptosis in SF126 glioma cells. Taken together, KIF20A is a tumor-associated antigen involved in the glioma cell growth and cell survival, suggesting that KIF20A is an oncoantigen of gliomas. Thus, KIF20A is a candidate novel immunotherapeutic target for gliomas.





Similar content being viewed by others
References
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B et al (2005) Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med 352:987–996
Imai K, Hirata S, Irie A, Senju S, Ikuta Y et al (2011) Identification of HLA-A2-restricted CTL epitopes of a novel tumour-associated antigen, KIF20A, overexpressed in pancreatic cancer. Br J Cancer 104:300–307
Nakamura T, Furukawa Y, Nakagawa H, Tsunoda T, Ohigashi H et al (2004) Genome-wide cDNA microarray analysis of gene expression profiles in pancreatic cancers using populations of tumor cells and normal ductal epithelial cells selected for purity by laser microdissection. Oncogene 23:2385–2400
Echard A, Jollivet F, Martinez O, Lacapere JJ, Rousselet A et al (1998) Interaction of a Golgi-associated kinesin-like protein with Rab6. Science 279:580–585
Hirokawa N, Noda Y, Okada Y (1998) Kinesin and dynein superfamily proteins in organelle transport and cell division. Curr Opin Cell Biol 10:60–73
Allan VJ, Schroer TA (1999) Membrane motors. Curr Opin Cell Biol 11:476–482
Gasnereau I, Boissan M, Margall-Ducos G, Couchy G, Wendum D et al (2012) KIF20A mRNA and its product MKlp2 are increased during hepatocyte proliferation and hepatocarcinogenesis. Am J Pathol 180:131–140
Yamashita J, Fukushima S, Jinnin M, Honda N, Makino K et al (2012) Kinesin family member 20 A is a novel melanoma-associated antigen. Acta Derm Venereol 92:593–597
Phuphanich S, Wheeler CJ, Rudnick JD, Mazer M, Wang H et al (2012) Phase I trial of a multi-epitope-pulsed dendritic cell vaccine for patients with newly diagnosed glioblastoma. Cancer Immunol Immunother 62:125–135
Yu JS, Liu G, Ying H, Yong WH, Black KL et al (2004) Vaccination with tumor lysate-pulsed dendritic cells elicits antigen-specific, cytotoxic T-cells in patients with malignant glioma. Cancer Res 64:4973–4979
Asahara S, Takeda K, Yamao K, Maguchi H, Yamaue H (2013) Phase I/II clinical trial using HLA-A24-restricted peptide vaccine derived from KIF20A for patients with advanced pancreatic cancer. J Transl Med 11:291
Taniuchi K, Nakagawa H, Nakamura T, Eguchi H, Ohigashi H et al (2005) Down-regulation of RAB6KIFL/KIF20A, a kinesin involved with membrane trafficking of discs large homologue 5, can attenuate growth of pancreatic cancer cell. Cancer Res 65:105–112
Hill E, Clarke M, Barr FA (2000) The Rab6-binding kinesin, Rab6-KIFL, is required for cytokinesis. EMBO J 19:5711–5719
McCollum D (2004) Cytokinesis: the central spindle takes center stage. Curr Biol 14:R953–R955
Gruneberg U, Neef R, Honda R, Nigg EA, Barr FA (2004) Relocation of Aurora B from centromeres to the central spindle at the metaphase to anaphase transition requires MKlp2. J Cell Biol 166:167–172
Echard A (1998) Interaction of a Golgi-Associated Kinesin-Like Protein with Rab6. Science 279:580–585
Hirokawa N, Noda Y, Tanaka Y, Niwa S (2009) Kinesin superfamily motor proteins and intracellular transport. Nat Rev Mol Cell Biol 10:682–696
Jordens I, Marsman M, Kuijl C, Neefjes J (2005) Rab proteins, connecting transport and vesicle fusion. Traffic 6:1070–1077
Okuyama R, Aruga A, Hatori T, Takeda K, Yamamoto M (2013) Immunological responses to a multi-peptide vaccine targeting cancer-testis antigens and VEGFRs in advanced pancreatic cancer patients. Oncoimmunology 2:e27010
Tomita Y, Yuno A, Tsukamoto H, Senju S, Kuroda Y et al (2013) Identification of promiscuous KIF20A long peptides bearing both CD4+ and CD8+ T-cell epitopes: KIF20A-specific CD4+ T-cell immunity in patients with malignant tumor. Clin Cancer Res 19:4508–4520
Eggert US, Mitchison TJ, Field CM (2006) Animal cytokinesis: from parts list to mechanisms. Annu Rev Biochem 75:543–566
Glotzer M (2005) The molecular requirements for cytokinesis. Science 307:1735–1739
Miki H, Setou M, Kaneshiro K, Hirokawa N (2001) All kinesin superfamily protein, KIF genes in mouse and human. Proc Natl Acad Sci USA 98:7004–7011
Straight AF, Field CM (2000) Microtubules, membranes and cytokinesis. Curr Biol 10:R760–R770
Wheatley SP, Wang Y (1996) Midzone microtubule bundles are continuously required for cytokinesis in cultured epithelial cells. J Cell Biol 135:981–989
Yan GR, Zou FY, Dang BL, Zhang Y, Yu G et al (2012) Genistein-induced mitotic arrest of gastric cancer cells by downregulating KIF20A, a proteomics study. Proteomics 12:2391–2399
Yu Y, Feng YM (2010) The role of kinesin family proteins in tumorigenesis and progression: potential biomarkers and molecular targets for cancer therapy. Cancer 116:5150–5160
Adams RR, Carmena M, Earnshaw WC (2001) Chromosomal passengers and the (aurora) ABCs of mitosis. Trends Cell Biol 11:49–54
D’Avino PP, Savoian MS, Glover DM (2005) Cleavage furrow formation and ingression during animal cytokinesis: a microtubule legacy. J Cell Sci 118:1549–1558
Ainsztein AM, Kandels-Lewis SE, Mackay AM, Earnshaw WC (1998) INCENP centromere and spindle targeting: identification of essential conserved motifs and involvement of heterochromatin protein HP1. J Cell Biol 143:1763–1774
Ruchaud S, Carmena M, Earnshaw WC (2007) Chromosomal passengers: conducting cell division. Nat Rev Mol Cell Biol 8:798–812
Buczkowicz P, Zarghooni M, Bartels U, Morrison A, Misuraca KL et al (2013) Aurora kinase B is a potential therapeutic target in pediatric diffuse intrinsic pontine glioma. Brain Pathol 23:244–253
Carvajal RD, Tse A, Schwartz GK (2006) Aurora kinases: new targets for cancer therapy. Clin Cancer Res 12:6869–6875
Portella G, Passaro C, Chieffi P (2011) Aurora B: a new prognostic marker and therapeutic target in cancer. Curr Med Chem 18:482–496
Yeung SC, Gully C, Lee MH (2008) Aurora-B kinase inhibitors for cancer chemotherapy. Mini Rev Med Chem 8:1514–1525
Zeng WF, Navaratne K, Prayson RA, Weil RJ (2007) Aurora B expression correlates with aggressive behaviour in glioblastoma multiforme. J Clin Pathol 60:218–221
Jung Y, Joo KM, Seong DH, Choi YL, Kong DS et al (2012) Identification of prognostic biomarkers for glioblastomas using protein expression profiling. Int J Oncol 40:1122–1132
Kajiwara Y, Yamasaki F, Hama S, Yahara K, Yoshioka H et al (2003) Expression of survivin in astrocytic tumors: correlation with malignant grade and prognosis. Cancer 97:1077–1083
Uematsu M, Ohsawa I, Aokage T, Nishimaki K, Matsumoto K et al (2005) Prognostic significance of the immunohistochemical index of survivin in glioma: a comparative study with the MIB-1 index. J Neurooncol 72:231–238
Zhen HN, Zhang X, Hu PZ, Yang TT, Fei Z et al (2005) Survivin expression and its relation with proliferation, apoptosis, and angiogenesis in brain gliomas. Cancer 104:2775–2783
Ngan CY, Yamamoto H, Takagi A, Fujie Y, Takemasa I et al (2008) Oxaliplatin induces mitotic catastrophe and apoptosis in esophageal cancer cells. Cancer Sci 99:129–139
Vitale I, Galluzzi L, Castedo M, Kroemer G (2011) Mitotic catastrophe: a mechanism for avoiding genomic instability. Nat Rev Mol Cell Biol 12:385–392
Wonsey DR, Follettie MT (2005) Loss of the forkhead transcription factor FoxM1 causes centrosome amplification and mitotic catastrophe. Cancer Res 65:5181–5189
Kenney-Herbert EM, Ball SL, Al-Mayhani TM, Watts C (2011) Glioblastoma cell lines derived under serum-free conditions can be used as an in vitro model system to evaluate therapeutic response. Cancer Lett 305:50–57
Lee J, Kotliarova S, Kotliarov Y, Li A, Su Q et al (2006) Tumor stem cells derived from glioblastomas cultured in bFGF and EGF more closely mirror the phenotype and genotype of primary tumors than do serum-cultured cell lines. Cancer Cell 9:391–403
Vik-Mo EO, Sandberg C, Olstorn H, Varghese M, Brandal P et al (2010) Brain tumor stemcells maintain overall phenotype and tumorigenicity after in vitro culturing in serum-free conditions. Neuro Oncol 12:1220–1230
Acknowledgements
We thank Ms. Y. Aikawa, Ms. N. Tsuzaki, and Ms. K. Koide for technical assistance. This research was supported by the Japan Society for the Promotion of Science Grants-in-Aid for Scientific Research (B), Grant Number 22390283.
Author contributions
Conceived and designed the experiments: KS, SO, YK, KS, and MT. Performed the experiments:KS, SO. Analyzed the data KS: Contributed reagents/materials/analysis tools: SO, YK, KS, and MT. Wrote the paper: KS, SO, and MT.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of Interest
The authors have no conflicts of interest to disclose.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11060_2016_2360_MOESM1_ESM.tif
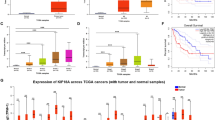
In silico analysis of REMBRANDT dataset, showing correlation between glioma patient overall survival and KIF20A expression. The relationship between patient prognosis and expression intensity of the target gene was analyzed in silico using the GlioVis portal (http://gliovis.bioinfo.cnio.es). Red line indicates high KIF20A expression (n=199), and blue line indicates low KIF20A expression (n=198). Patients with high expression (red line) have significantly shorter survival than those with a low expression (blue line) level of KIF20A (TIF 3315 KB)
11060_2016_2360_MOESM2_ESM.tif
siRNAs effect on expression of KIF20A in glioma cells. a KIF20A expression was investigated 2 and 5 days after KIF20A siRNA transfection in SF126 glioma cells, compared with mock (untreated) and control siRNA treatment as assessed by western blot analysis. b KIF20A expression was investigated 5 days after KIF20A siRNA transfection in U251MG glioma cells and U87MG glioma-like cells, compared with control siRNA treatment as assessed by western blot analysis (TIF 7600 KB)
Rights and permissions
About this article
Cite this article
Saito, K., Ohta, S., Kawakami, Y. et al. Functional analysis of KIF20A, a potential immunotherapeutic target for glioma. J Neurooncol 132, 63–74 (2017). https://doi.org/10.1007/s11060-016-2360-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-016-2360-1