Abstract
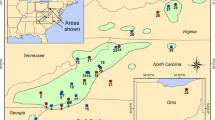
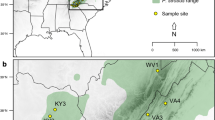
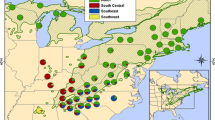
Eastern hemlock (Tsuga canadensis [L.] Carr.) is a widespread and ecologically important conifer species of eastern North America that is threatened by the hemlock woolly adelgid (Adelges tsugae Annand), a pest introduced into the United States from Asia in the 1920s. Information about the genetic composition of eastern hemlock is necessary to guide ex situ conservation efforts in the southeastern United States, where the species is expected to harbor relatively high amounts of genetic variation in areas of Pleistocene glacial refuge. Nineteen allozyme markers were used to quantify the genetic variation present in 20 eastern hemlock populations in the southeastern United States. Results indicate that the species has low levels of genetic diversity in the region compared to most other conifers, but greater population differentiation (F ST = 0.126). Populations along the eastern periphery and in the Appalachian interior exhibited higher levels of diversity than those along the western periphery of its geographic range. The results suggest that the glacial refuge area for eastern hemlock was likely located east of the southern Appalachian Mountains, and indicate that ex situ conservation seed collections should be concentrated in these areas of higher diversity.


Similar content being viewed by others
References
Ally D, El-Kassaby Y, Ritland K (2000) Genetic diversity, differentiation and mating system in mountain hemlock (Tsuga mertensiana) across British Columbia. For Genet 7(2):97–108
Alvarez-Buylla ER, Chaos A, Pinero D et al (1996) Demographic genetics of a pioneer tropical tree species: patch dynamics, seed dispersal, and seed banks. Evolution 50(3):1155–1166
Bhiry N, Filion L (1996) Mid-Holocene hemlock decline in eastern North America linked with phytophagous insect activity. Quat Res 45(3):312–320
Bohonak AJ (2002) IBD (isolation by distance): a program for analyses of isolation by distance. J Hered 93(2):153–154
Brooks RT (2001) Effects of the removal of overstory hemlock from hemlock-dominated forests on eastern redback salamanders. For Ecol Manage 149(1–3):197–204
Brown AHD, Hardner CM (2000) Sampling the gene pools of forest trees for ex situ conservation. In: Young AG, Boshier D, Boyle TJ (eds) Forest conservation genetics: principles and practice. CABI Publishing, Collingwood, Australia
Camcore (2005) 2005 Camcore Annual Report. Camcore International Tree Conservation and Domestication, Department of Forestry and Environmental Resources, North Carolina State University, Raleigh, NC
Camcore (2006) 2006 Camcore Annual Report. Camcore International Tree Conservation and Domestication, Department of Forestry and Environmental Resources, North Carolina State University, Raleigh, NC
Cavers S, Degen B, Caron H et al (2005) Optimal sampling strategy for estimation of spatial genetic structure in tree populations. Heredity 95(4):281–289
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144(4):2001–2014
Cronin TM, Szabo BJ, Ager TA et al (1981) Quaternary climates and sea levels of the United States Atlantic Coastal Plain. Science 211(4479):233–240
Cwynar LC, Macdonald GM (1987) Geographical variation of lodgepole pine in relation to population history. Am Nat 129(3):463–469
Davis MB (1981) Quaternary history and the stability of forest communities. In: Shugart HH, Botkin DB, West DC (eds) Forest succession, concepts and applications. Springer, New York
Delcourt PA, Delcourt HR (1987) Long-term forest dynamics of the temperate zone. Springer, New York
Delcourt PA, Delcourt HR, Brister RC et al (1980) Quaternary vegetation history of the Mississippi embayment. Quat Res 13(1):111–132
Ellison AM, Bank MS, Clinton BD et al (2005) Loss of foundation species: consequences for the structure and dynamics of forested ecosystems. Front Ecol Environ 3(9):479–486
Eriksson G, Namkoong G, Roberds JH (1993) Dynamic gene conservation for uncertain futures. For Ecol Manage 62(1–4):15–37
ESRI (2001) ArcGIS 8.1. Environmental Systems Research Institute, Redlands, CA
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Felsenstein J (2005) PHYLIP (phylogeny inference package), version 3.6. Department of Genome Sciences, University of Washington, Seattle
Ford CR, Vose JM (2007) Tsuga canadensis (L.) Carr. mortality will impact hydrologic processes in southern appalachian forest ecosystems. Ecol Appl 17(4):1156–1167
Fuller JL (1998) Ecological impact of the mid-holocene hemlock decline in southern Ontario, Canada. Ecology 79(7):2337–2351
Godman RM, Lancaster K (1990) Eastern hemlock. In: Burns RM, Barbara H. Honkala (eds) Silvics of North America: 1 Conifers. United States Department of Agriculture Forest Service, Washington, DC
Hamrick JL, Godt MJW (1996) Effects of life history traits on genetic diversity in plant species. Philos Trans R Soc B 351(1345):1291–1298
Hardin JW, Cooper AW (1967) Mountain disjuncts in the eastern Piedmont of North Carolina. J Elisha Mitch Sci S 83(3):139–150
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. Biol J Linn Soc 58(3):247–276
Hewitt GM (2000) The genetic legacy of the quaternary ice ages. Nature 405(6789):907–913
Jenkins JC, Aber JD, Canham CD (1999) Hemlock woolly adelgid impacts on community structure and N cycling rates in eastern hemlock forests. Can J For Res 29(5):630–645
Jorgensen SM, Hamrick JL (1997) Biogeography and population genetics of whitebark pine, Pinus albicaulis. Can J For Res 27(10):1574–1585
Kimura M, Crow JF (1964) Number of alleles that can be maintained in a finite population. Genetics 49(4):725–738
Kimura M, Ohta T (1978) Stepwise mutation model and distribution of allelic frequencies in a finite population. Proc Natl Acad Sci USA 75(6):2868–2872
Kizlinski ML, Orwig DA, Cobb RC et al (2002) Direct and indirect ecosystem consequences of an invasive pest on forests dominated by eastern hemlock. J Biogeogr 29(10–11):1489–1503
Ledig FT (2000) Founder effects and the genetic structure of Coulter pine. J Hered 91(4):307–315
Li P, MackKay J, Bousquet J (1992) Genetic diversity in Canadian hardwoods: implications for conservation. Forest Chron 68(6):709–719
McClure MS, Salom SM, Shields KS (2003) Hemlock woolly Adelgid. Forest Health Technology Enterprise Team. United States Department of Agriculture, Forest Service, Morgantown, WV
McWilliams WH, Schmidt TL (2000) Composition, structure and sustainability of hemock ecosystems in eastern North America. In: Proceedings of the symposium on sustainable management of hemlock ecosystems in Eastern North America, Durham, New Hampshire, United States Department of Agriculture, Forest Service
Nei M (1972) Genetic distance between populations. Am Nat 106(949):283–292
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89(3):583–590
Nienstaedt H, Olson JS (1961) Effects of photoperiod and source on seedling growth of eastern hemlock. Forest Sci 7(1):81–96
Olson JS, Nienstaedt H (1957) Photoperiod and chilling control growth of hemlock. Science 125(3246):492–494
Oosting HJ, Hess DW (1956) Microclimate and a relic stand of Tsuga canadensis in the lower piedmont of North Carolina. Ecology 37(1):28–39
Orwig DA, Foster DR (1998) Forest response to the introduced hemlock woolly adelgid in southern New England, USA. J Torrey Bot Soc 125(1):60–73
Orwig DA, Foster DR, Mausel DL (2002) Landscape patterns of hemlock decline in New England due to the introduced hemlock woolly adelgid. J Biogeogr 29(10–11):1475–1487
Parker KC, Hamrick JL, Parker AJ et al (1997) Allozyme diversity in Pinus virginiana (Pinaceae): intraspecific and interspecific comparisons. Am J Bot 84(10):1372–1382
Parshall T (2002) Late holocene stand-scale invasion by hemlock (Tsuga canadensis) at its western range limit. Ecology 83(5):1386–1398
Petit RJ, Aguinagalde I, de Beaulieu JL et al (2003) Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300(5625):1563–1565
Piry S, Luikart G, Cornuet JM (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90(4):502–503
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959
Pritchard JK, Wen W (2004) Documentation for STRUCTURE software: version 2.0. Department of Human Genetics, University of Chicago, Chicago, IL
Rajora OP, Mosseler A (2001) Challenges and opportunities for conservation of forest genetic resources. Euphytica 118(2):197–212
Ross RM, Bennett RM, Snyder CD et al (2003) Influence of eastern hemlock (Tsuga canadensis L.) on fish community structure and function in headwater streams of the Delaware River basin. Ecol Freshw Fish 12(1):60–65
SAS Institute Inc. (2003) The SAS System for Windows, Version 9.1. Cary, NC
Small MJ, Small CJ, Dreyer GD (2005) Changes in a hemlock-dominated forest following woolly adelgid infestation in southern New England. J Torrey Bot Soc 132(3):458–470
Snyder CD, Young JA, Lemarie DP et al (2002) Influence of eastern hemlock (Tsuga canadensis) forests on aquatic invertebrate assemblages in headwater streams. Can J Fish Aquat Sci 59(2):262–275
Tingley MW, Orwig DA, Field R (2002) Avian response to removal of a forest dominant: consequences of hemlock woolly adelgid infestations. J Biogeogr 29(10–11):1505–1516
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13(4):921–935
Wang C, Perlin MH, VanStockum RR et al (1997) Chloroplast DNA polymorphisms in Tsuga canadensis and Tsuga caroliniana. Can J For Res 27(9):1329–1335
Watts WA (1970) Full-glacial vegetation of northwestern Georgia. Ecology 51(1):17–33
Wellman H, Ritland C, Ritland K (2003) Genetic effects of domestication in western hemlock, Tsuga heterophylla. For Genet 10(5):229–239
Whitehead DR (1973) Late-Wisconsin vegetational changes in unglaciated eastern North America. Quat Res 3:621–631
Whitlock MC, McCauley DE (1999) Indirect measures of gene flow and migration: F ST ≠ 1/(4N m + 1). Heredity 82:117–125
Yeh FC, Yang R-C, Boyle TBJ et al (2000) PopGene32, microsoft windows-based freeware for population genetic analysis, version 1.32. Molecular Biology and Biotechnology Centre, University of Alberta, Edmonton, AB, Canada
Young JA, Young CG (1992) Seeds of woody plants in North America. Dioscorides Press, Portland, OR
Zabinski C (1992) Isozyme variation in eastern hemlock. Can J For Res 22(12):1838–1842
Acknowledgments
We appreciate the assistance from the following individuals in identifying eastern hemlock populations in the southeastern United States, and in procuring permits for collecting foliage samples: Russ MacFarlane, Joe McGuiness, Tina Tilley, Paul Bradley, Randall Burgess, and Mark Robison (USDA Forest Service); Tom Remaley (National Park Service); Allen Rogers (South Mountains State Park, North Carolina); Tim Lee (Caesar’s Head State Park, South Carolina); Zeb Weese (Natural Bridge State Park, Kentucky); Dean Henson (Pine Mountain State Park, Kentucky); Ed Smith (Lone Mountain State Forest, Tennessee); Justin Walden (Scott State Forest, Tennessee); Jerry Moody (N.C. Extension Service); and Winston Church (private landowner). We thank Jennifer DeWoody and Ricardo Hernandez at the National Forest Genetic Electrophoresis Laboratory for their technical support, and three anonymous reviewers for their comments on the manuscript. We also thank the USDA Forest Service and the members of Camcore for their financial support of this project. The foliage collection and general project support were funded by USDA Forest Service grant 05-DG-11083150–210.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Potter, K.M., Dvorak, W.S., Crane, B.S. et al. Allozyme variation and recent evolutionary history of eastern hemlock (Tsuga canadensis) in the southeastern United States. New Forests 35, 131–145 (2008). https://doi.org/10.1007/s11056-007-9067-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11056-007-9067-2