Abstract
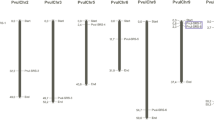
The present study is aimed to identify and characterize HSP70 (PvHSP70) genes in two different common bean cultivars under salt stress. For this purpose various in silico methods such as RNAseq data and qRT-PCR analysis were used. A total of 24 candidate PvHSP70 gene were identified. Except for chromosome 4 and 7, these candidate PvHSP70 genes were distributed on the remaining chromosomes. While the lowest number of PvHSP70 genes was determined on chromosomes 1, 3, 5, 7, 9, 10 and 11 (one HSP70 gene), the highest number of PvHSP70s was on chromosomes 6 and 8 (seven HSP70 genes each). Three genes; PvHSP70-5, -9, and -10 were found to have no-introns. In addition, four tandemly and six segmentally duplicated gene couples were detected. A total of 13 PvHSP70 genes were targeted by miRNAs of 44 plant species and the most targeted genes were PvHSP70-5 and -23. The expression profile of PvHSP70 genes based on publicly available RNA-seq data was identified and salt treated leaf tissue was found to have more gene expression levels compared to the root. qRT-PCR analysis showed that the transcript concentrations of upregulated PvHSP70 genes in leaves of Zulbiye (sensitive) were mostly higher than those of Yakutiye (resistant). The present study revealed that PvHSP70 genes might play an important role in salt stress response for common bean cultivars and variability between cultivars also suggests that these genes could be used as functional markers for salt tolerance in common bean.







Similar content being viewed by others
References
Timperio AM, Egidi MG, Zolla L (2008) Proteomics applied on plant abiotic stresses: role of heat shock proteins (HSP). J proteom 71(4):391–411. doi:10.1016/j.jprot.2008.07.005
Ahuja I, de Vos RC, Bones AM, Hall RD (2010) Plant molecular stress responses face climate change. Trends Plant Sci 15(12):664–674. doi:10.1016/j.tplants.2010.08.002
Li Z, Srivastava P (2004) Heat-shock proteins. Current protocols in immunology/edited by John E Coligan [et al] Appendix 1:Appendix 1T.10.1002/0471142735.ima01ts58
Cashikar AG, Duennwald M, Lindquist SL (2005) A chaperone pathway in protein disaggregation. Hsp26 alters the nature of protein aggregates to facilitate reactivation by Hsp104. J Biol Chem 280(25):23869–23875. doi:10.1074/jbc.M502854200
Sarkar NK, Kim YK, Grover A (2009) Rice sHsp genes: genomic organization and expression profiling under stress and development. BMC Genom 10:393. doi:10.1186/1471-2164-10-393
Wang W, Vinocur B, Shoseyov O, Altman A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9(5):244–252. doi:10.1016/j.tplants.2004.03.006
Zhu X, Zhao X, Burkholder WF, Gragerov A, Ogata CM, Gottesman ME, Hendrickson WA (1996) Structural analysis of substrate binding by the molecular chaperone DnaK. Science 272(5268):1606–1614
Masand S, Yadav SK (2016) Overexpression of MuHSP70 gene from Macrotyloma uniflorum confers multiple abiotic stress tolerance in transgenic Arabidopsis thaliana. Mol Biol Rep 43(2):53–64. doi:10.1007/s11033-015-3938-y
Richter K, Haslbeck M, Buchner J (2010) The heat shock response: life on the verge of death. Mol Cell 40(2):253–266. doi:10.1016/j.molcel.2010.10.006
Masand S, Yadav SK (2016) Overexpression of MuHSP70 gene from Macrotyloma uniflorum confers multiple abiotic stress tolerance in transgenic Arabidopsis thaliana. Mol Biol Rep 43(2):53–64
Sung DY, Kaplan F, Guy CL (2001) Plant Hsp70 molecular chaperones: protein structure, gene family, expression and function. Physiol Plant 113(4):443–451. doi:10.1034/j.1399-3054.2001.1130402.x
Guy CL, Li QB (1998) The organization and evolution of the spinach stress 70 molecular chaperone gene family. Plant Cell 10(4):539–556
Zhou SJ, Jing Z, Shi JL (2013) Genome-wide identification, characterization, and expression analysis of the MLO gene family in Cucumis sativus. Genet mol res 12(4):6565–6578. doi:10.4238/2013.December.11.8
Frydman J (2001) Folding of newly translated proteins in vivo: the role of molecular chaperones. Annu Rev Biochem 70:603–647. doi:10.1146/annurev.biochem.70.1.603
Daugaard M, Rohde M, Jaattela M (2007) The heat shock protein 70 family: highly homologous proteins with overlapping and distinct functions. FEBS Lett 581(19):3702–3710. doi:10.1016/j.febslet.2007.05.039
Jego G, Hazoume A, Seigneuric R, Garrido C (2013) Targeting heat shock proteins in cancer. Cancer Lett 332(2):275–285. doi:10.1016/j.canlet.2010.10.014
Alvim FC, Carolino SM, Cascardo JC, Nunes CC, Martinez CA, Otoni WC, Fontes EP (2001) Enhanced accumulation of BiP in transgenic plants confers tolerance to water stress. Plant Physiol 126(3):1042–1054
Cho EK, Hong CB (2006) Over-expression of tobacco NtHSP70-1 contributes to drought-stress tolerance in plants. Plant Cell Rep 25(4):349–358. doi:10.1007/s00299-005-0093-2
Su PH, Li HM (2008) Arabidopsis stromal 70-kD heat shock proteins are essential for plant development and important for thermotolerance of germinating seeds. Plant Physiol 146(3):1231–1241. doi:10.1104/pp.107.114496
Jungkunz I, Link K, Vogel F, Voll LM, Sonnewald S, Sonnewald U (2011) AtHsp70-15-deficient Arabidopsis plants are characterized by reduced growth, a constitutive cytosolic protein response and enhanced resistance to TuMV. Plant J 66(6):983–995. doi:10.1111/j.1365-313X.2011.04558.x
Augustine SM, Cherian AV, Syamaladevi DP, Subramonian N (2015) Erianthus arundinaceus HSP70 (EaHSP70) acts as a key regulator in the formation of anisotropic interdigitation in sugarcane (saccharum spp. hybrid) in response to drought stress. Plant Cell Physiol 56(12):2368–2380
Yer EN, Baloglu MC, Ziplar UT, Ayan S, Unver T (2015) Drought-responsive Hsp70 gene analysis in populus at genome-wide level. Plant Molecular Biology Reporter.1–18
Goodstein DM, Shu SQ, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res 40(D1):D1178–D1186. doi:10.1093/nar/gkr944
Guo AY, Zhu QH, Chen X, Luo JC (2007) [GSDS: a gene structure display server]. Yi chuan = Hereditas/Zhongguo yi chuan xue hui bian ji 29 (8):1023–1026
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J hered 93(1):77–78
Yang ZF, Gu SL, Wang XF, Li WJ, Tang ZX, Xu CW (2008) Molecular evolution of the CPP-like gene family in plants: insights from comparative genomics of Arabidopsis and rice. J Mol Evol 67(3):266–277. doi:10.1007/s00239-008-9143-z
Bailey TL, Williams N, Misleh C, Li WW (2006) MEME: discovering and analyzing DNA and protein sequence motifs. Nucleic acids research 34 (Web Server issue):W369–373. 10.1093/nar/gkl198
Quevillon E, Silventoinen V, Pillai S, Harte N, Mulder N, Apweiler R, Lopez R (2005) InterProScan: protein domains identifier. Nucleic acids research 33 (Web Server issue):W116–120. 10.1093/nar/gki442
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25(24):4876–4882. doi:10.1093/nar/25.24.4876
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. doi:10.1093/molbev/msr121
Letunic I, Bork P (2011) Interactive tree of life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478. doi:10.1093/nar/gkr201
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35 (Web Server issue):W585–587. 10.1093/nar/gkm259
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP SignalP and related tools. Nat Protoc 2(4):953–971
Conesa A, Gotz S, Garcia-Gomez JM, Terol J, Talon M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18):3674–3676. doi:10.1093/bioinformatics/bti610
Zhang YJ (2005) miRU: an automated plant miRNA target prediction server. Nucleic Acids Res 33:W701–W704. doi:10.1093/nar/gki383
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28(1):235–242. doi:10.1093/Nar/28.1.235
Kelley LA, Sternberg MJE (2009) Protein structure prediction on the Web: a case study using the Phyre server. Nat Protoc 4(3):363–371. doi:10.1038/nprot.2009.2
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34 (Web Server issue):W609-612.10.1093/nar/gkl315
Lynch M, Conery JS (2003) The evolutionary demography of duplicate genes. J Struct Funct Genom 3(1–4):35–44
Yang Z, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17(1):32–43
Hiz MC, Canher B, Niron H, Turet M (2014) Transcriptome analysis of salt tolerant common bean (Phaseolus vulgaris L.) under saline conditions. PLoS One 9(3):e92598. doi:10.1371/journal.pone.0092598
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628. doi:10.1038/nmeth.1226
Caraux G, Pinloche S (2005) PermutMatrix: a graphical environment to arrange gene expression profiles in optimal linear order. Bioinformatics 21(7):1280–1281. doi:10.1093/bioinformatics/bti141
Guler NS, Saglam A, Demiralay M, Kadioglu A (2012) Apoplastic and symplastic solute concentrations contribute to osmotic adjustment in bean genotypes during drought stress. Turk J Biol 36(2):151–160. doi:10.3906/biy-1101-177
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method. Methods 25(4):402–408
Lin BL, Wang JS, Liu HC, Chen RW, Meyer Y, Barakat A, Delseny M (2001) Genomic analysis of the Hsp70 superfamily in Arabidopsis thaliana. Cell Stress Chaperones 6(3):201–208. doi:10.1379/1466-1268(2001)006<0201:Gaoths>2.0.Co;2
Zhou SJ, Jing Z, Shi JL (2013) Genome-wide identification, characterization, and expression analysis of the MLO gene family in Cucumis sativus. Genet Mol Res 12(4):6565–6578. doi:10.4238/2013.December.11.8
Wang Y, Lin S, Song Q, Li K, Tao H, Huang J, Chen X, Que S, He H (2014) Genome-wide identification of heat shock proteins (Hsps) and Hsp interactors in rice: hsp70s as a case study. BMC Genom 15:344. doi:10.1186/1471-2164-15-344
Zhang Y, Wang M, Chen J, Rong J, Ding M (2014) [Genome-wide analysis of HSP70 superfamily in Gossypium raimondii and the expression of orthologs in Gossypium hirsutum]. Yi chuan = Hereditas/Zhongguo yi chuan xue hui bian ji 36 (9):921–933
Nishizawa A, Yabuta Y, Yoshida E, Maruta T, Yoshimura K, Shigeoka S (2006) Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J 48(4):535–547. doi:10.1111/j.1365-313X.2006.02889.x
Jiang C, Xu J, Zhang H, Zhang X, Shi J, Li M, Ming F (2009) A cytosolic class I small heat shock protein, RcHSP17.8, of Rosa chinensis confers resistance to a variety of stresses to Escherichia coli, yeast and Arabidopsis thaliana. Plant Cell Environ 32(8):1046–1059. doi:10.1111/j.1365-3040.2009.01987.x
Zhang J, Li J, Liu B, Zhang L, Chen J, Lu M (2013) Genome-wide analysis of the Populus Hsp90 gene family reveals differential expression patterns, localization, and heat stress responses. BMC Genom 14:532. doi:10.1186/1471-2164-14-532
Yang JY, Sun Y, Sun AQ, Yi SY, Qin J, Li MH, Liu J (2006) The involvement of chloroplast HSP100/ClpB in the acquired thermotolerance in tomato. Plant Mol Biol 62(3):385–395. doi:10.1007/s11103-006-9027-9
Sarkar NK, Kundnani P, Grover A (2013) Functional analysis of Hsp70 superfamily proteins of rice (Oryza sativa). Cell Stress Chaperones 18(4):427–437. doi:10.1007/s12192-012-0395-6
Zhang J, Li JB, Liu BB, Zhang L, Chen J, Lu MZ (2013) Genome-wide analysis of the Populus Hsp90 gene family reveals differential expression patterns, localization, and heat stress responses. BMC Genom 14:532. doi:10.1186/1471-2164-14-532
Flagel LE, Wendel JF (2009) Gene duplication and evolutionary novelty in plants. New phytol 183(3):557–564. doi:10.1111/j.1469-8137.2009.02923.x
Magadum S, Banerjee U, Murugan P, Gangapur D, Ravikesavan R (2013) Gene duplication as a major force in evolution. J Genet 92(1):155–161
Wang YP, Wang XY, Tang HB, Tan X, Ficklin SP, Feltus FA, Paterson AH (2011) Modes of gene duplication contribute differently to genetic novelty and redundancy, but show parallels across divergent angiosperms. PLoS One 6(12):e28150
Zhang L, Zhao HK, Dong QL, Zhang YY, Wang YM, Li HY, Xing GJ, Li QY, Dong YS (2015) Genome-wide analysis and expression profiling under heat and drought treatments of HSP70 gene family in soybean (Glycine max L.). Front. Plant Sci 6:773. doi:10.3389/fpls.2015.00773
Hou JJ, Jiang PP, Qi SM, Zhang K, He QX, Xu CZ, Ding ZH, Zhang KW, Li KP (2016) Isolation and functional validation of salinity and osmotic stress inducible promoter from the maize type-ii h + -pyrophosphatase gene by deletion analysis in transgenic tobacco plants. PLoS One 11(4):e0154041
Czarnecka E, Key JL, Gurley WB (1989) Regulatory domains of the Gmhsp17.5-E heat shock promoter of soybean. Mol Cell Biol 9(8):3457–3463
Cho EK, Choi YJ (2009) A nuclear-localized HSP70 confers thermoprotective activity and drought-stress tolerance on plants. Biotechnol Lett 31(4):597–606. doi:10.1007/s10529-008-9880-5
Kose S, Furuta M, Imamoto N (2012) Hikeshi, a nuclear import carrier for Hsp70 s, protects cells from heat shock-induced nuclear damage. Cell 149(3):578–589. doi:10.1016/j.cell.2012.02.058
Li WX, Oono Y, Zhu J, He XJ, Wu JM, Iida K, Lu XY, Cui X, Jin H, Zhu JK (2008) The Arabidopsis NFYA5 transcription factor is regulated transcriptionally and posttranscriptionally to promote drought resistance. Plant Cell 20(8):2238–2251. doi:10.1105/tpc.108.059444
Chi X, Yang Q, Chen X, Wang J, Pan L, Chen M, Yang Z, He Y, Liang X, Yu S (2011) Identification and characterization of microRNAs from peanut (Arachis hypogaea L.) by high-throughput sequencing. PLoS One 6(11):e27530. doi:10.1371/journal.pone.0027530
Debernardi JM, Rodriguez RE, Mecchia MA, Palatnik JF (2012) Functional specialization of the plant mir396 regulatory network through distinct microrna-target interactions. PLoS Genet. doi:10.1371/journal.pgen.1002419
Rodriguez RE, Ercoli MF, Debernardi JM, Breakfield NW, Mecchia MA, Sabatini M, Cools T, De Veylder L, Benfey PN, Palatnik JF (2015) MicroRNA miR396 regulates the switch between stem cells and transit-amplifying cells in arabidopsis roots. Plant Cell 27(12):3354–3366
Liu DM, Yu DQ (2009) MicroRNA (miR396) negatively regulates expression of ceramidase-like genes in Arabidopsis. Prog Nat Sci 19(6):781–785. doi:10.1016/j.pnsc.2008.09.006
Dong CH, Pei HX (2014) Over-expression of miR397 improves plant tolerance to cold stress in Arabidopsis thaliana. J Plant Biol 57(4):209–217. doi:10.1007/s12374-013-0490-y
Guleria P, Yadav SK (2011) Identification of miR414 and expression analysis of conserved miRNAs from Stevia rebaudiana. Genom proteom bioinform 9(6):211–217. doi:10.1016/S1672-0229(11)60024-7
Nijhawan A, Jain M, Tyagi AK, Khurana JP (2008) Genomic survey and gene expression analysis of the basic leucine zipper transcription factor family in rice. Plant Physiol 146(2):333–350. doi:10.1104/pp.107.112821
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17(6):369–381. doi:10.1016/j.tplants.2012.02.004
Soding J (2005) Protein homology detection by HMM-HMM comparison. Bioinformatics 21(7):951–960. doi:10.1093/bioinformatics/bti125
Mulaudzi-Masuku T, Mutepe RD, Mukhoro OC, Faro A, Ndimba B (2015) Identification and characterization of a heat-inducible Hsp70 gene from Sorghum bicolor which confers tolerance to thermal stress. Cell Stress Chaperones 20(5):793–804. doi:10.1007/s12192-015-0591-2
Guo HM, Li ZC, Zhou ML, Cheng HM (2014) cDNA-AFLP analysis reveals heat shock proteins play important roles in mediating cold, heat, and drought tolerance in Ammopiptanthus mongolicus. Funct Integr Genomics 14(1):127–133. doi:10.1007/s10142-013-0347-y
Montero-Barrientos M, Hermosa R, Cardoza RE, Gutierrez S, Nicolas C, Monte E (2010) Transgenic expression of the Trichoderma harzianum hsp70 gene increases Arabidopsis resistance to heat and other abiotic stresses. J Plant Physiol 167(8):659–665. doi:10.1016/j.jplph.2009.11.012
Hiz MC, Canher B, Niron H, Turet M (2014) Transcriptome Analysis of Salt Tolerant Common Bean (Phaseolus vulgaris L.) under Saline Conditions. PLoS One. doi:10.1371/journal.pone.0092598
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Golldack D, Li C, Mohan H, Probst N (2014) Tolerance to drought and salt stress in plants: unraveling the signaling networks. Front Plant Sci. doi:10.3389/Fpls.2014.00151
Bartels D, Sunkar R (2005) Drought and salt tolerance in plants. Crit Rev Plant Sci 24(1):23–58
Sun WN, Bernard C, van de Cotte B, Van Montagu M, Verbruggen N (2001) At-HSP17.6A, encoding a small heat-shock protein in Arabidopsis, can enhance osmotolerance upon overexpression. Plant J 27(5):407–415
Golldack D, Li C, Mohan H, Probst N (2014) Tolerance to drought and salt stress in plants: unraveling the signaling networks. Front plant sci 5:151. doi:10.3389/fpls.2014.00151
Garg DSS, Dalal S, Tiwari R, Singh R (2012) Heat shock protein based SNP marker for terminal heat stress in wheat (Triticum aestivum L.). Aust J Crop Sci 6(11):1516–1521
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Büyük, İ., Inal, B., Ilhan, E. et al. Genome-wide identification of salinity responsive HSP70s in common bean. Mol Biol Rep 43, 1251–1266 (2016). https://doi.org/10.1007/s11033-016-4057-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-016-4057-0