Abstract
Ion homeostasis is considered to be one of the most important mechanisms underlying salt stress tolerance. We used the Steptoe × Morex barley doubled haploid population to screen for genetic variation in response to salinity stress at an early development stage in a hydroponics system, focusing on ion homeostasis. Salinity induced a strong adverse effect on growth of the parents and their derived population, with Steptoe as the more tolerant parent. Steptoe maintained higher concentrations of K+, Na+ and Cl− in the roots and a similar shoot/root ion ratio (<1) under stress conditions compared to control conditions. In contrast, Morex had higher concentrations of these ions in the shoots under stress and a doubled shoot/root ion ratio relative to control conditions, indicating that salt exclusion might contribute to the higher tolerance of Steptoe. Correlation and path analysis demonstrated that shoot Cl− contents most strongly affected salt tolerance and suggest that both Na+ and Cl− contents are important for salinity stress tolerance in barley. We identified 11 chromosomal regions involved in the control of the variation observed for salt tolerance and various salt stress response traits, including Na+, Cl− and K+ contents in shoots. Two specific regions on chromosomes 2H and 3H were found controlling ion contents and salt tolerance, pointing to genes involved in ion homeostasis that contribute to salt tolerance.


Similar content being viewed by others
References
Abel GH (1969) Inheritance of capacity for chloride inclusion and chloride exclusion by soybeans. Crop Sci 9(6):697
Byrt CS, Platten JD, Spielmeyer W, James RA, Lagudah ES, Dennis ES, Tester M, Munns R (2007) HKT1;5-like cation transporters linked to Na + exclusion loci in wheat, Nax2 and Kna1. Plant Physiol 143(4):1918–1928. doi:10.1104/pp.093476
Chen F, Hayes PM (1989) A comparison of Hordeum bulbosum-mediated haploid production efficiency in barley using in vitro floret and tiller culture. Theor Appl Genet 77:701–704
Chen Z, Newman I, Zhou M, Mendham N, Zhang G, Shabala S (2005) Screening plants for salt tolerance by measuring K + flux: a case study for barley. Plant, Cell Environ 28(10):1230–1246
Chen Z, Pottosin II, Cuin TA, Fuglsang AT, Tester M, Jha D, Zepeda-Jazo I, Zhou M, Palmgren MG, Newman IA, Shabala S (2007) Root plasma membrane transporters controlling K +/Na + homeostasis in salt-stressed barley. Plant Physiol 145(4):1714–1725. doi:10.1104/pp.107.110262
Choi DW, Close TJ (2000) A newly identified barley gene, Dhn12 encoding a YSK2 DHN, is located on chromosome 6H and has embryo-specific expression. Theor Appl Genet 100(8):1274–1278
Close TJ, Bhat PR, Lonardi S, Wu Y, Rostoks N, Ramsay L, Druka A, Stein N, Svensson JT, Wanamaker S, Bozdag S, Roose ML, Moscou MJ, Chao S, Varshney RK, Szucs P, Sato K, Hayes PM, Matthews DE, Kleinhofs A, Muehlbauer GJ, DeYoung J, Marshall DF, Madishetty K, Fenton RD, Condamine P, Graner A, Waugh R (2009) Development and implementation of high-throughput SNP genotyping in barley. BMC Genomics 10:582
Dewey DRLK (1959) A correlation and path coefficient analysis of components of crested wheatgrass seed production. Agron J 51:515–518
Dubcovsky J, Maria GS, Epstein E, Luo MC, Dvorak J (1996) Mapping of the K +/Na + discrimination locus Kna1 in wheat. Theor Appl Genet 92(3–4):448–454
Ellis RP, Forster BP, Waugh R, Bonar N, Handley LL, Robinson D, Gordon DC, Powell W (1997) Mapping physiological traits in barley. New Phytol 137(1):149–157
Ellis RP, Forster BP, Gordon DC, Handley LL, Keith RP, Lawrence P, Meyer R, Powell W, Robinson D, Scrimgeour CM, Young G, Thomas WT (2002) Phenotype/genotype associations for yield and salt tolerance in a barley mapping population segregating for two dwarfing genes. J Exp Bot 53(371):1163–1176
FAO (2008) FAO land and plant nutrition management service. http://wwwfaoorg/ag/agl/agll/spush
Flowers TJ (2004) Improving crop salt tolerance. J Exp Bot 55(396):307–319
Flowers TJ, Hajibagheri MA (2001) Salinity tolerance in Hordeum vulgare: ion concentrations in root cells of cultivars differing in salt tolerance. Plant Soil 231(1):1–9
Garcia de Moral LF, Rharrabti Y, Elhani S, Martos V, Royo C (2005) Yield formation in Mediterranean durum wheat under two contrasting water regimes based on path-coefficient analysis. Euphytica 146:203–212
Genc Y, Oldach K, Verbyla AP, Lott G, Hassan M, Tester M, Wallwork H, McDonald GK (2010a) Sodium exclusion QTL associated with improved seedling growth in bread wheat under salinity stress. Theor Appl Genet 121(5):877–894. doi:10.1007/s00122-010-1357-y
Genc Y, Tester M, McDonald GK (2010b) Calcium requirement of wheat in saline and non-saline conditions. Plant Soil 327(1–2):331–345. doi:10.1007/s11104-009-0057-3
Gorham J, Bridges J, Dubcovsky J, Dvorak J, Hollington PA, Luo MC, Khan JA (1997) Genetic analysis and physiology of a trait for enhanced K +/Na + discrimination in wheat. New Phytol 137(1):109–116
Greenway H, Munns R (1980) Mechanisms of salt tolerance in non-halophytes. Annu Rev Plant Physiol 31:149–190
Han F, Ullrich SE, Romagosa I, Clancy JA, Froseth JA, Wesenberg DM (2003) Quantitative genetic analysis of acid detergent fibre content in barley grain. J Cereal Sci 38(2):167–172. doi:10.1016/S0733-5210(03)00020-1
Hauser F, Horie T (2010) A conserved primary salt tolerance mechanism mediated by HKT transporters: a mechanism for sodium exclusion and maintenance of high K(+)/Na(+) ratio in leaves during salinity stress. Plant, Cell Environ 33(4):552–565
Huang SB, Spielmeyer W, Lagudah ES, James RA, Platten JD, Dennis ES, Munns R (2006) A sodium transporter (HKT7) is a candidate for Nax1, a gene for salt tolerance in durum wheat. Plant Physiol 142(4):1718–1727. doi:10.1104/pp.106.088864
Huang S, Spielmeyer W, Lagudah ES, Munns R (2008) Comparative mapping of HKT genes in wheat, barley, and rice, key determinants of Na + transport, and salt tolerance. J Exp Bot 59(4):927–937. doi:10.1093/jxb/ern033
Hui Z, Zhang ZB, Shao HB, Ping X, Foulkes MJ (2008) Genetic correlation and path analysis of transpiration efficiency for wheat flag leaves. Environ Exp Bot 64(2):128–134. doi:10.1016/j.envexpbot.2007.11.001
Isla R, Aragues R, Royo A (1998) Validity of various physiological traits as screening criteria for salt tolerance in barley. Field Crop Res 58(2):97–107
James RA, Davenport RJ, Munns R (2006a) Physiological characterization of two genes for Na + exclusion in durum wheat, Nax1 and Nax2. Plant Physiol 142(4):1537–1547
James RA, Munns R, Von Caemmerer S, Trejo C, Miller C, Condon AG (2006b) Photosynthetic capacity is related to the cellular and subcellular partitioning of Na + , K + and Cl-salt-affected barley and durum wheat. Plant, Cell Environ 29:2185–2197
James RA, Blake C, Byrt CS, Munns R (2011) Major genes for Na + exclusion Nax1 and Nax2 (wheat HKT1;4 and HKT1;5) decrease Na + accumulation in bread wheat under saline and waterlogged conditions. J Exp Bot 62:2939–2947
Kaneko T, Zhang WS, Takahashi H, Ito K, Takeda K (2001) QTL mapping for enzyme activity and thermostability of beta-amylase in barley (Hordeum vulgare L.). Breed Sci 51(2):99–105
Kleinhofs A, Kilian A, Maroof MAS, Biyashev RM, Hayes P, Chen FQ, Lapitan N, Fenwick A, Blake TK, Kanazin V, Ananiev E, Dahleen L, Kudrna D, Bollinger J, Knapp SJ, Liu B, Sorrells M, Heun M, Franckowiak JD, Hoffman D, Skadsen R, Steffenson BJ (1993) A molecular, isozyme and morphological map of the barley (Hordeum vulgare) genome. Theor Appl Genet 86(6):705–712
Koyama ML, Levesley A, Koebner RM, Flowers TJ, Yeo AR (2001) Quantitative trait loci for component physiological traits determining salt tolerance in rice. Plant Physiol 125(1):406–422
Lee SY, Ahn JH, Cha YS, Yun DW, Lee MC, Ko JC, Lee KS, Eun MY (2006) Mapping of quantitative trait loci for salt tolerance at the seedling stage in rice. Mol Cell 21(2):192–196
Ligaba A, Katsuhara M (2010) Insights into the salt tolerance mechanism in barley (Hordeum vulgare) from comparisons of cultivars that differ in salt sensitivity. J Plant Res 123(1):105–118. doi:10.1007/s10265-009-0272-2
Lin HX, Zhu MZ, Yano M, Gao JP, Liang ZW, Su WA, Hu XH, Ren ZH, Chao DY (2004) QTLs for Na + and K + uptake of the shoots and roots controlling rice salt tolerance. Theor Appl Genet 108(2):253–260. doi:10.1007/s00122-003-1421-y
Lindsay MP, Lagudah ES, Hare RA, Munns R (2004) A locus for sodium exclusion (Nax1), a trait for salt tolerance, mapped in durum wheat. Funct Plant Biol 31(11):1105–1114. doi:10.1071/Fp04111
Liu JP, Zhu JK (1998) A calcium sensor homolog required for plant salt tolerance. Science 280(5371):1943–1945
Maas EV, Hoffman GJ (1977) Crop salt tolerance-current assessment. J Irrig Drain 103(IR2):115–134
Mano Y, Takeda K (1997) Mapping quantitative trait loci for salt tolerance at germination and the seedling stage in barley (Hordeum vulgare L). Euphytica 94(3):263–272
Mano Y, Nakazumi H, Takeda K (1996) Varietal variation in and effects of some major genes on salt tolerance at the germination stage in barley. Breed Sci 46:227–233
Mather DE (1995) Base maps of NABGMP crosses. http://gnome.agrenv.mcgill.ca/info/basemaps.htm
Munns R, Rawson HM (1999) Effect of salinity on salt accumulation and reproductive development in the apical meristem of wheat and barley. Aust J Plant Physiol 26(5):459–464
Munns R, Tester M (2008) Mechanisms of salinity tolerance. Annu Rev Plant Biol 59:651–681. doi:10.1146/annurev.arplant.59.032607.092911
Munns R, Husain S, Rivelli AR, James RA, Condon AG, Lindsay MP, Lagudah ES, Schachtman DP, Hare RA (2002) Avenues for increasing salt tolerance of crops, and the role of physiologically based selection traits. Plant Soil 247(1):93–105
Munns R, James RA, Lauchli A (2006) Approaches to increasing the salt tolerance of wheat and other cereals. J Exp Bot 57(5):1025–1043. doi:10.1093/jxb/erj100
Munns R, James RA, Xu B, Athman A, Conn SJ, Jordans C, Byrt CS, Hare RA, Tyerman SD, Tester M, Plett D, Gilliham M (2012) Wheat grain yield on saline soils is improved by an ancestral Na(+) transporter gene. Nat Biotechnol. doi:10.1038/nbt.2120
Rajendran K, Tester M, Roy SJ (2009) Quantifying the three main components of salinity tolerance in cereals. Plant, Cell Environ 32(3):237–249. doi:10.1111/j.1365-3040.2008.01916.x
Ren ZH, Gao JP, Li LG, Cai XL, Huang W, Chao DY, Zhu MZ, Wang ZY, Luan S, Lin HX (2005) A rice quantitative trait locus for salt tolerance encodes a sodium transporter. Nat Genet 37(10):1141–1146. doi:10.1038/Ng1643
Rivandi J, Miyazaki J, Hrmova M, Pallotta M, Tester M, Collins NC (2011) A SOS3 homologue maps to HvNax4, a barley locus controlling an environmentally sensitive Na(+) exclusion trait. J Exp Bot 62:1201–1216
Salvi S, Tuberosa R (2005) To clone or not to clone plant QTLs: present and future challenges. Trends Plant Sci 10(6):297–304
Shabala SN, Shabala SI, Martynenko AI, Babourina O, Newman IA (1998) Salinity effect on bioelectric activity, growth, Na + accumulation and chlorophyll fluorescence of maize leaves: a comparative survey and prospects for screening. Aust J Plant Physiol 25(5):609–616
Shabala S, Cuin TA, Pang J, Percey W, Chen Z, Conn S, Eing C, Wegner LH (2010) Xylem ionic relations and salinity tolerance in barley. Plant J 61(5):839–853. doi:10.1111/j.1365-313X.2009.04110.x
Shavrukov Y, Gupta NK, Miyazaki J, Baho MN, Chalmers KJ, Tester M, Langridge P, Collins NC (2010) HvNax3—a locus controlling shoot sodium exclusion derived from wild barley (Hordeum vulgare ssp. spontaneum). Funct Integr Genomics 10(2):277–291. doi:10.1007/s10142-009-0153-8
Siahsar BA, Narouei M (2010) Mapping QTLs of physiological traits associated with salt tolerance in ‘Steptoe’x‘Morex’ doubled haploid lines of barley at seedling stage. J Food Agric Environ 8(2):751–759
Silva P, Geros H (2009) Regulation by salt of vacuolar H + -ATPase and H + -pyrophosphatase activities and Na +/H + exchange. Plant Signal Behav 4(8):718–726
Sirault XRR, James RA, Furbank RT (2009) A new screening method for osmotic component of salinity tolerance in cereals using infrared thermography. Funct Plant Biol 36(10–11):970–977. doi:10.1071/Fp09182
Storey R, Jones RGW (1978) Salt stress and comparative physiology in gramineae. I-Ion relations of two salt and water stressed barley cultivars, California Mariout and Arimar. Aust J Plant Physiol 5:801–816
Tavakkoli E, Rengasamy P, McDonald GK (2010) The response of barley to salinity stress differs between hydroponic and soil systems. Funct Plant Biol 37(7):621–633. doi:10.1071/Fp09202
Tavakkoli E, Fatehi F, Coventry S, Rengasamy P, McDonald GK (2011) Additive effects of Na + and Cl- ions on barley growth under salinity stress. J Exp Bot 62(6):2189–2203. doi:10.1093/Jxb/Erq422
Teakle NL, Tyerman SD (2010) Mechanisms of Cl- transport contributing to salt tolerance. Plant, Cell Environ 33(4):566–589. doi:10.1111/j.1365-3040.2009.02060.x
Van Ooijen JW (2009) MapQTL6, software for the mapping of quantitative trait loci in experimental populations of diploid species. Kyazma BV, Wageningen
Verhoeven KJ, Biere A, Nevo E, van Damme JM (2004) Differential selection of growth rate-related traits in wild barley, Hordeum spontaneum, in contrasting greenhouse nutrient environments. J Evol Biol 17(1):184–196
Volis S, Verhoeven KJ, Mendlinger S, Ward D (2004) Phenotypic selection and regulation of reproduction in different environments in wild barley. J Evol Biol 17(5):1121–1131. doi:10.1111/j.1420-9101.2004.00738.xJEB738
Volkov V, Amtmann A (2006) Thellungiella halophila, a salt-tolerant relative of Arabidopsis thaliana, has specific root ion-channel features supporting K +/Na + homeostasis under salinity stress. Plant J 48:342–353
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93(1):77–78
Walia H, Wilson C, Wahid A, Condamine P, Cui X, Close TJ (2006) Expression analysis of barley (Hordeum vulgare L.) during salinity stress. Funct Integr Genomics 6(2):143–156. doi:10.1007/s10142-005-0013
Walia H, Wilson C, Condamine P, Liu X, Ismail AM, Close TJ (2007) Large-scale expression profiling and physiological characterization of jasmonic acid-mediated adaptation of barley to salinity stress. Plant, Cell Environ 30(4):410–421. doi:10.1111/j.1365-3040.2006.01628.x
White PJ, Broadley MR (2001) Chloride in soils and its uptake and movement within the plant: a review. Ann Bot 88(6):967–988. doi:10.1006/anbo.2001.1540
Witzel K, Weidner A, Surabhi GK, Borner A, Mock HP (2009) Salt stress-induced alterations in the root proteome of barley genotypes with contrasting response towards salinity. J Exp Bot 60(12):3545–3557. doi:10.1093/Jxb/Erp198
Witzel K, Weidner A, Surabhi GK, Varshney RK, Kunze G, Buck-Sorlin GH, Borner A, Mock HP (2010) Comparative analysis of the grain proteome fraction in barley genotypes with contrasting salinity tolerance during germination. Plant, Cell Environ 33(2):211–222. doi:10.1111/j.1365-3040.2009.02071
Xue DW, Huang YZ, Zhang XQ, Wei K, Westcott S, Li CD, Chen MC, Zhang GP, Lance R (2009) Identification of QTLs associated with salinity tolerance at late growth stage in barley. Euphytica 169(2):187–196. doi:10.1007/s10681-009-9919-2
Zhou GF, Johnson P, Ryan PR, Delhaize E, Zhou MX (2012) Quantitative trait loci for salinity tolerance in barley (Hordeum vulgare L.). Mol Breed 29:427–436
Acknowledgments
This work was financially supported by the Ministry of EL&I, The Netherlands and the Vietnamese Ministry of Education. The expert assistance of Geurt Versteeg in the greenhouse is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11032_2012_9777_MOESM1_ESM.doc
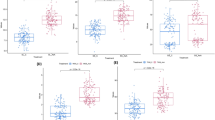
Ion contents in shoot (a) and in root (b) of Steptoe, Morex and the mean of the derived DH population. Ion contents were measured from pooled samples of four replications grown in two environments, viz. control (C) and salt stress (S). Shoot and root cation (Na+, K+, Mg2+ and Ca2+), anion (Cl−; PO4 3− and SO4 2−) and total ion content (sum of all studied cation and anion) ratio of Steptoe (St), Morex (Mx) and DH population mean (c). (DOC 157 kb)
11032_2012_9777_MOESM2_ESM.xls
Coefficient of correlations for the traits under both control and stress condition grown for 3 weeks on hydroponics (XLS 58 kb)
Rights and permissions
About this article
Cite this article
Nguyen, V.L., Ribot, S.A., Dolstra, O. et al. Identification of quantitative trait loci for ion homeostasis and salt tolerance in barley (Hordeum vulgare L.). Mol Breeding 31, 137–152 (2013). https://doi.org/10.1007/s11032-012-9777-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-012-9777-9