Abstract
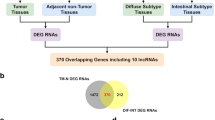
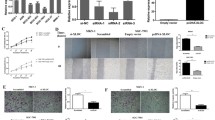
Long non-coding RNAs (LncRNAs) have been reported that play important roles in the progression and metastasis of some carcinomas. In the present study, we identified a new LncRNA, FRLnc1, from a microarray analysis in which those LncRNAs were regulated by FOXM1, an oncogene widely studied in most malignancies. Quantitative real-time PCR (qRT-PCR) results in gastric cancer cell lines indicated FRLnc1 expression is positively correlated with FOXM1 level, supporting the microarray data. Furthermore, the RNA level of FRLnc1 is upregulated in 49 % (20/41) of cancer samples compared with neighboring non-cancerous stomach tissues. The in vitro functional analyses demonstrated that FRLnc1 knockdown by RNA interference suppressed cell migration in MGC803 and AGS cells, whereas FRLnc1 overexpression promoted cell migration in BGC823 and SGC7901 cells. Moreover, FRLnc1 could enhance the distant metastasis of SGC7901 cells by tail vein injection approach in mice. We also identified TGFβ1 and Twist as the downstream effectors of FRLnc1 in the regulation of cell migration by qRT-PCR analysis. Taken together, our findings suggest that FRLnc1 is involved in gastric cancer cell migration and for the first time set up the link between FOXM1 and LncRNA in cancer.






Similar content being viewed by others
References
Tanpakushitsu HM, Koso K et al (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–945. doi:10.1038/nature03001
Tahira AC, Kubrusly MS, Faria MF, Dazzani B, Fonseca RS, Maracaja-Coutinho V, Verjovski-Almeida S, Machado MC, Reis EM (2011) Long noncoding intronic RNAs are differentially expressed in primary and metastatic pancreatic cancer. Mol Cancer 10:141. doi:10.1186/1476-4598-10-141
Ponting CP, Belgard TG (2010) Transcribed dark matter: meaning or myth? Hum Mol Genet 19:R162–R168. doi:10.1093/hmg/ddq362
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10:155–159. doi:10.1038/nrg2521
Brosnan CA, Voinnet O (2009) The long and the short of noncoding RNAs. Curr Opin Cell Biol 21:416–425. doi:10.1016/j.ceb.2009.04.001
Rinn JL, Chang HY (2012) Genome regulation by long noncoding RNAs. Annu Rev Biochem 81:145–166. doi:10.1146/annurev-biochem-051410-092902
Mattick JS (2009) The genetic signatures of noncoding RNAs. PLoS Genet 5:e1000459. doi:10.1371/journal.pgen.1000459
Ounzain S, Pezzuto I, Micheletti R, Burdet F, Sheta R, Nemir M, Gonzales C, Sarre A, Alexanian M, Blow MJ, May D, Johnson R, Dauvillier J, Pennacchio LA, Pedrazzini T (2014) Functional importance of cardiac enhancer-associated noncoding RNAs in heart development and disease. J Mol Cell Cardiol 76:55–70. doi:10.1016/j.yjmcc.2014.08.009
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, Guernec G, Martin D, Merkel A, Knowles DG, Lagarde J, Veeravalli L, Ruan X, Ruan Y, Lassmann T, Carninci P, Brown JB, Lipovich L, Gonzalez JM, Thomas M, Davis CA, Shiekhattar R, Gingeras TR, Hubbard TJ, Notredame C, Harrow J, Guigo R (2012) The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res 22:1775–1789. doi:10.1101/gr.132159.111
Marques AC, Ponting CP (2009) Catalogues of mammalian long noncoding RNAs: modest conservation and incompleteness. Genome Biol 10:R124. doi:10.1186/gb-2009-10-11-r124
Yang G, Lu X, Yuan L (2014) LncRNA: a link between RNA and cancer. Biochim Biophys Acta 1839:1097–1109. doi:10.1016/j.bbagrm.2014.08.012
Lu L, Zhu G, Zhang C, Deng Q, Katsaros D, Mayne ST, Risch HA, Mu L, Canuto EM, Gregori G, Benedetto C, Yu H (2012) Association of large noncoding RNA HOTAIR expression and its downstream intergenic CpG island methylation with survival in breast cancer. Breast Cancer Res Treat 136:875–883. doi:10.1007/s10549-012-2314-z
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, Wang Y, Brzoska P, Kong B, Li R, West RB, van de Vijver MJ, Sukumar S, Chang HY (2010) Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464:1071–1076. doi:10.1038/nature08975
Yang F, Xue X, Bi J, Zheng L, Zhi K, Gu Y, Fang G (2013) Long noncoding RNA CCAT1, which could be activated by c-Myc, promotes the progression of gastric carcinoma. J Cancer Res Clin Oncol 139:437–445. doi:10.1007/s00432-012-1324-x
He X, Tan X, Wang X, Jin H, Liu L, Ma L, Yu H, Fan Z (2014) C-Myc-activated long noncoding RNA CCAT1 promotes colon cancer cell proliferation and invasion. Tumour Biol 35:12181–12188. doi:10.1007/s13277-014-2526-4
Yuan JH, Yang F, Wang F, Ma JZ, Guo YJ, Tao QF, Liu F, Pan W, Wang TT, Zhou CC, Wang SB, Wang YZ, Yang Y, Yang N, Zhou WP, Yang GS, Sun SH (2014) A long noncoding RNA activated by TGF-beta promotes the invasion-metastasis cascade in hepatocellular carcinoma. Cancer Cell 25:666–681. doi:10.1016/j.ccr.2014.03.010
Li W, Kang Y (2014) A new Lnc in metastasis: long noncoding RNA mediates the prometastatic functions of TGF-beta. Cancer Cell 25:557–559. doi:10.1016/j.ccr.2014.04.014
Qi P, Du X (2013) The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod Pathol 26:155–165. doi:10.1038/modpathol.2012.160
Yamamoto H, Watanabe Y, Maehata T, Morita R, Yoshida Y, Oikawa R, Ishigooka S, Ozawa S, Matsuo Y, Hosoya K, Yamashita M, Taniguchi H, Nosho K, Suzuki H, Yasuda H, Shinomura Y, Itoh F (2014) An updated review of gastric cancer in the next-generation sequencing era: insights from bench to bedside and vice versa. World J Gastroenterol 20:3927–3937. doi:10.3748/wjg.v20.i14.3927
Smith MG, Hold GL, Tahara E, El-Omar EM (2006) Cellular and molecular aspects of gastric cancer. World J Gastroenterol 12:2979–2990
Okada K, Fujiwara Y, Takahashi T, Nakamura Y, Takiguchi S, Nakajima K, Miyata H, Yamasaki M, Kurokawa Y, Mori M, Doki Y (2013) Overexpression of forkhead box M1 transcription factor (FOXM1) is a potential prognostic marker and enhances chemoresistance for docetaxel in gastric cancer. Ann Surg Oncol 20:1035–1043. doi:10.1245/s10434-012-2680-0
Li Q, Zhang N, Jia Z, Le X, Dai B, Wei D, Huang S, Tan D, Xie K (2009) Critical role and regulation of transcription factor FoxM1 in human gastric cancer angiogenesis and progression. Cancer Res 69:3501–3509. doi:10.1158/0008-5472.can-08-3045
Zhang Y, Zhang N, Dai B, Liu M, Sawaya R, Xie K, Huang S (2008) FoxM1B transcriptionally regulates vascular endothelial growth factor expression and promotes the angiogenesis and growth of glioma cells. Cancer Res 68:8733–8742. doi:10.1158/0008-5472.can-08-1968
Zeng J, Wang L, Li Q, Li W, Bjorkholm M, Jia J, Xu D (2009) FoxM1 is up-regulated in gastric cancer and its inhibition leads to cellular senescence, partially dependent on p27 kip1. J Pathol 218:419–427. doi:10.1002/path.2530
Huang C, Du J, Xie K (2014) FOXM1 and its oncogenic signaling in pancreatic cancer pathogenesis. Biochim Biophys Acta 1845:104–116. doi:10.1016/j.bbcan.2014.01.002
Bella L, Zona S, de Moraes GN, Lam EW (2014) FOXM1: a key oncofetal transcription factor in health and disease. Semin Cancer Biol 29c:32–39. doi:10.1016/j.semcancer.2014.07.008
Lam EW, Brosens JJ, Gomes AR, Koo CY (2013) Forkhead box proteins: tuning forks for transcriptional harmony. Nat Rev Cancer 13:482–495. doi:10.1038/nrc3539
Behren A, Muhlen S, Acuna Sanhueza GA, Schwager C, Plinkert PK, Huber PE, Abdollahi A, Simon C (2010) Phenotype-assisted transcriptome analysis identifies FOXM1 downstream from Ras-MKK3-p38 to regulate in vitro cellular invasion. Oncogene 29:1519–1530. doi:10.1038/onc.2009.436
Fang XY, Pan HF, Leng RX, Ye DQ (2015) Long noncoding RNAs: novel insights into gastric cancer. Cancer Lett 356:357–366. doi:10.1016/j.canlet.2014.11.005
Bartscht T, Lehnert H, Gieseler F, Ungefroren H (2012) The Src family kinase inhibitors PP2 and PP1 effectively block TGF-beta1-induced cell migration and invasion in both established and primary carcinoma cells. Cancer Chemother Pharmacol 70:221–230. doi:10.1007/s00280-012-1904-0
Ge R, Rajeev V, Ray P, Lattime E, Rittling S, Medicherla S, Protter A, Murphy A, Chakravarty J, Dugar S, Schreiner G, Barnard N, Reiss M (2006) Inhibition of growth and metastasis of mouse mammary carcinoma by selective inhibitor of transforming growth factor-beta type I receptor kinase in vivo. Clin Cancer Res 12:4315–4330. doi:10.1158/1078-0432.ccr-06-0162
Mohammad KS, Javelaud D, Fournier PG, Niewolna M, McKenna CR, Peng XH, Duong V, Dunn LK, Mauviel A, Guise TA (2011) TGF-beta-RI kinase inhibitor SD-208 reduces the development and progression of melanoma bone metastases. Cancer Res 71:175–184. doi:10.1158/0008-5472.can-10-2651
Yang J, Mani SA, Donaher JL, Ramaswamy S, Itzykson RA, Come C, Savagner P, Gitelman I, Richardson A, Weinberg RA (2004) Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 117:927–939. doi:10.1016/j.cell.2004.06.006
Acknowledgments
This work was supported in part by grants from National Natural Science Foundation of China (81272917 and 81272241), Shanghai medical key specialty (ZK2012A26), Key Disciplines Group Construction Project of Pudong Health Bureau of Shanghai (PWZxq2014-04) and Outstanding Leaders Training Program of Pudong Health Bureau of Shanghai (PWRl2013-02).
Conflict of interest
The authors declare that no conflicts of interest exist.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Hui Cai and Jingde Chen have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Cai, H., Chen, J., He, B. et al. A FOXM1 related long non-coding RNA contributes to gastric cancer cell migration. Mol Cell Biochem 406, 31–41 (2015). https://doi.org/10.1007/s11010-015-2421-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-015-2421-3