Abstract
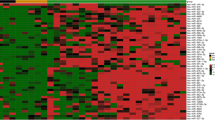
The genetic factors of cancer predisposition remain elusive in the majority of familial and/or early-onset cases of breast cancer (BC). This type of BC is promoted by germ-line mutations that inactivate BRCA1 or BRCA2. On the other hand, recent studies have indicated that alterations in the levels of miRNA expression are linked to this disease. Although BRCA1 and BRCA2 gene mutations have been reported to commonly lead to alterations in genes that encode cancer-related proteins, little is known regarding the putative impact of these mutations on noncoding miRNAs. In the present study, we aimed to determine whether miRNA dysregulation is involved in the pathogenesis of BRCA-mutated BC. An expression analysis of 14 human miRNAs previously shown to be related to BC diagnosis, prognosis, and drug resistance was conducted using tissues from 60 familial and/or early-onset patients whose peripheral blood samples had been screened for BRCA1 and BRCA2 mutations through sequence analysis. Let-7a and miR-335 expression levels were significantly downregulated in the tumors of patients with a BRCA mutation compared with those of patients without a BRCA mutation (P = 0.04 and P = 0.02, respectively). Our results defined the associations between the expression status of let-7a and miR-335 and BRCA mutations. The expression analysis of these miRNAs might be used as biomarkers of the BRCA mutation status of early-onset and/or familial BC.



Similar content being viewed by others
Abbreviations
- BC:
-
Breast cancer
- ER:
-
Estrogen receptor
- PR:
-
Progesterone receptor
- HER-2:
-
Human epidermal growth factor receptor 2
References
Egeli U, Cecener G, Tunca B, Tasdelen I (2006) Novel germline BRCA1 and BRCA2 mutations in Turkish women with breast and/or ovarian cancer and their relatives. Cancer Invest 24:484–491
Schnitt SJ (2010) Classification and prognosis of invasive breast cancer: from morphology to molecular taxonomy. Mod Pathol 2:60–64
Ozmen V, Ozcinar B, Karanlik H, Cabioglu N, Tukenmez M, Disci R, Ozmen T, Igci A, Muslumanoglu M, Kecer M, Soran A (2009) Breast cancer risk factors in Turkish women—a university hospital based nested case control study. World J Surg Oncol 37:1–8
Parkin DM, Bray F, Ferlay J, Pisani P (2005) Global cancer statistics, 2002. CA Cancer J Clin 55:74–108
Kim Y, Choi JY, Lee KM, Park SK, Ahn SH, Noh DY, Hong YC, Kang D, Yoo KY (2007) Dose dependent protective effect of breast feeding against breast cancer among ever lactated women in Korea. Eur J Cancer Prev 16:124–129
Tuncer M (2008) Significance of cancer in Turkey, the burden of disease and cancer control policies. In: Tuncer M (ed) Cancer control in Turkey, vol 74. Ankara, Onur Press, Health Ministry Publication, pp 5–9
Szabo CI, King MC (1997) Population genetics of BRCA1 and BRCA2. Am J Hum Genet 60:1013–1020
Ferla R, Calò V (2007) Founder mutations in BRCA1 and BRCA2 genes. Ann Oncol 18:93–98
Kenemans P, Verstraeten RA, Verheijen RHM (2004) Oncogenic pathways in hereditary and sporadic breast cancer. Maturitas 49:34–43
Imyanitov EN, Moiseyenko VM (2011) Drug therapy for hereditary cancers. Hered Cancer Clin Pract 6:5–9
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, Aldler H, Rattan S, Keating M, Rai K, Rassenti L, Kipps T, Negrini M, Bullrich F, Croce CM (2002) Frequent deletions and down regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci 99:15524–15529
Calin GA, Sevignani C, Dumitru CD, Hyslop T, Noch E, Yendamuri S, Shimizu M, Rattan S, Bullrich F, Negrini M, Croce CM (2004) Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc Natl Acad Sci 101:2999–3004
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ, Hammond SM (2005) A microRNA polycistron as a potential human oncogene. Nature 435:828–833
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D, Sweet-Cordero A, Ebert BL, Mak RH, Ferrando AA, Downing JR, Jacks T, Horvitz HR, Golub TR (2005) MicroRNA expression profiles classify human cancers. Nature 435:834–838
Calin GA, Croce C (2006) MicroRNA signatures in human cancers. Nat Rev Cancer 6:857–866
Esquela-Kerscher A, Slack FJ (2006) OncomiRs—microRNAs with a role in cancer. Nat Rev Cancer 6:259–269
Garzon R, Fabbri M, Cimmino A, Calin GA, Croce CM (2006) MicroRNA expression and function in cancer. Trends Mol Med 12:580–587
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of posttranscriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114
Iorio M, Casalini P, Tagliabue E, Ménard S, Croce C (2008) MicroRNA profiling as a tool to understand prognosis, therapy response and resistance in breast cancer. Eur J Cancer 44:2753–2759
Ng EK, Wong CL, Ma ES, Kwong A (2009) MicroRNAs as new players for diagnosis, prognosis, and therapeutic targets in breast cancer. J Oncol 305420:1–6
Visone R, Croce C (2009) MicroRNAs and cancer. Am J Pathol 174:1131–1138
Pinto R, Pilato B, Ottini L, Lambo R, Simone G, Paradiso A, Tommasi S (2013) Different methylation and microRNA expression pattern in male and female familial breast cancer. J Cell Physiol 228:1264–1269
Findlay VJ (2010) MicroRNAs and breast cancer. Open Cancer J 3:55–61
Brewster BL, Rossiello F, French JD, Edwards SL, Wong M, Wronski A, Whiley P, Waddell N, Chen X, Bove B, kConFab, Hopper JL, John EM, Andrulis I, Daly M, Volorio S, Bernard L, Peissel B, Manoukian S, Barile M, Pizzamiglio S, Verderio P, Spurdle AB, Radice P, Godwin AK, Southey MC, Brown MA, Peterlongo P (2012) Identification of fifteen novel germline variants in the BRCA1 3′UTR reveals a variant in a breast cancer case that introduces a functional miR-103 target site. Hum Mutat 33:1665–1675
Lheureux S, Lambert B, Krieger S, Legros A, Vaur D, Denoyelle C, Berthet P, Poulain L, Hardouin A (2011) Two novel variants in the 3′UTR of the BRCA1 gene in familial breast and/or ovarian cancer. Breast Cancer Res Treat 125:885–891
Kontorovich T, Levy A, Korostishevsky M, Nir U, Friedman E (2010) Single nucleotide polymorphisms in miRNA binding sites and miRNA genes as breast/ovarian cancer risk modifiers in Jewish high-risk women. Int J Cancer 127:589–597
Shen J, Ambrosone CB, Zhao H (2009) Novel genetic variants in microRNA genes and familial breast cancer. Int J Cancer 124:1178–1182
Barh D, Malhotra R, Ravi B, Sindhurani P (2010) MicroRNA let-7: an emerging nextgeneration cancer therapeutic. Curr Oncol 17:70–80
O’Day E, Lal A (2010) MicroRNAs and their target gene networks in breast cancer. Breast Cancer Res 12:201–211
Cecener G, Egeli U, Tunca B, Erturk E, Ak S, Gokgoz S, Tasdelen I, Tezcan G, Demirdogen E, Bayrom N, Avcı N, Evrensel T (2014) BRCA1/2 Germline mutations and their clinical importance in Turkish breast cancer patients. Cancer Invest 1–13. doi:10.3109/07357907.2014.919302
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(–∆ ∆ C(T)) method. Methods 25:402–408
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE, Iorio MV, Visone R, Sever NI, Fabbri M, Iuliano R, Palumbo T, Pichiorri F, Roldo C, Garzon R, Sevignani C, Rassenti L, Alder H, Volinia S, Liu CG, Kipps TJ, Negrini M, Croce CM (2005) A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 353:1793–1801
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F, Visone R, Iorio M, Roldo C, Ferracin M, Prueitt RL, Yanaihara N, Lanza G, Scarpa A, Vecchione A, Negrini M, Harris CC, Croce CM (2006) A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci 103:2257–2261
Yanaihara N, Caplen N, Bowman E, Seike M, Kumamoto K, Yi M, Stephens RM, Okamoto A, Yokota J, Tanaka T, Calin GA, Liu CG, Croce CM, Harris CC (2006) Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9:189–198
Barbarotto E, Schmittgen DT, Calin G (2008) MicroRNAs and cancer: profile, profile, profile. Int J Cancer 122:969–977
Lee CH, Subramanian S, Beck AH, Espinosa I, Senz J, Zhu SX, Huntsman D, van de Rijn M, Gilks CB (2009) MicroRNA profiling of BRCA1/2 mutation-carrying and non-mutation-carrying high-grade serous carcinomas of ovary. PLoS One 4:7314–7325
Zhang H, Li Y, Lai M (2010) The microRNA network and tumor metastasis. Oncogene 29:937–948
Zhao Y, Deng C, Lu W, Xiao J, Ma D, Guo M, Recker RR, Gatalica Z, Wang Z, Xiao GG (2011) Let-7 microRNAs induce tamoxifen sensitivity by down regulation of estrogen receptor alpha signaling in breast cancer. Mol Med 17:1233–1241
Kim SJ, Shin JY, Lee KD, Bae YK, Sung KW, Nam SJ, Chun KH (2012) MicroRNA let-7a suppresses breast cancer cell migration and invasion through down regulation of C–C chemokine receptor type 7. Breast Cancer Res 18:14–26
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H, Harano T, Yatabe Y, Nagino M, Nimura Y, Mitsudomi T, Takahashi T (2004) Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res 64:3753–3756
Johnson CD, Esquela-Kerscher A, Stefani G, Byrom M, Kelnar K, Ovcharenko D, Wilson M, Wang X, Shelton J, Shingara J, Chin L, Brown D, Slack FJ (2007) The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 67:7713–7722
Marson A, Levine SS, Cole MF, Frampton GM, Brambrink T, Johnstone S, Guenther MG, Johnston WK, Wernig M, Newman J, Calabrese JM, Dennis LM, Volkert TL, Gupta S, Love J, Hannett N, Sharp PA, Bartel DP, Jaenisch R, Young RA (2008) Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134:521–533
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C, Huang Y, Hu X, Su F, Lieberman J, Song E (2007) Let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131:1109–1123
Kubista M, Rosner M, Kubista E, Bernaschek G, Hengstschlager M (2002) Brca1 regulates in vitro differentiation of mammary epithelial cells. Oncogene 21:4747–4756
Furuta S, Jiang X, Gu B, Cheng E, Chen PL, Lee WH (2005) Depletion of BRCA1 impairs differentiation but enhances proliferation of mammary epithelial cells. Proc Natl Acad Sci USA 102:9176–9181
Bane A, Viloria-Petit A, Pinnaduwage D, Mulligan AM, O’Malley FP, Andrulis IL (2013) Clinical–pathologic significance of cancer stem cell marker expression in familial breast cancers. Breast Cancer Res Treat 140:195–205
Heyn H, Engelmann M, Schreek S, Ahrens P, Lehmann U, Kreipe H, Schlegelberger B, Beger C (2011) MicroRNA miR-335 is crucial for the BRCA1 regulatory cascade in breast cancer development. Int J Cancer 129:2797–2806
Hafez MM, Hassan ZK, Zekri AR, Gaber AA, Al Rejaie SS, Sayed-Ahmed MM, Al Shabanah O (2012) MicroRNAs and metastasis-related gene expression in Egyptian breast cancer patients. Asian Pac J Cancer Prev 13:591–598
Tavazoie SF, Alarcón C, Oskarsson T, Padua D, Wang Q, Bos PD, Gerald WL, Massagué J (2008) Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451:147–152
Sorrentino A, Liu C, Addario A, Peschle C, Scambia G, Ferlini C (2008) Role of microRNAs in drug-resistant ovarian cancer cells. Gynecol Oncol 111:478–486
Verghese ET, Hanby AM, Speirs V, Hughes TA (2008) Small is beautiful: microRNAs and breast cancer—where are we now? J Pathol 215:214–221
Shı M, Lıu D, Duan H, Shen B, Guo N (2010) Metastasis-related miRNAs, active players in breast cancer invasion and metastasis. Cancer Metastasis Rev 29:785–799
Inwald EC, Klinkhammer-Schalke M, Hofstädter F, Zeman F, Koller M, Gerstenhauer M, Ortmann O (2013) Ki-67 is a prognostic parameter in breast cancer patients: results of a large population-based cohort of a cancer registry. Breast Cancer Res Treat 139:539–552
Polley MY, Leung SC, McShane LM, Gao D, Hugh JC, Mastropasqua MG, Viale G, Zabaglo LA, Penault-Llorca F, Bartlett JM, Gown AM, Symmans WF, Piper T, Mehl E, Enos RA, Hayes DF, Dowsett M, Nielsen TO (2013) On behalf of the International Ki67 in Breast Cancer Working Group of the Breast International Group and North American Breast Cancer Group: an International Ki67 Reproducibility Study. J Natl Cancer Inst 105:1897–1906
Zhao JJ, Lin J, Yang H, Kong W, He L, Ma X, Coppola D, Cheng JQ (2008) MicroRNA-221/222 negatively regulates estrogen receptor alpha and is associated with tamoxifen resistance in breast cancer. J Biol Chem 283:31079–31086
Acknowledgments
We would like to thank Professor Kay Huebner for her scientific advice and careful reading of the draft manuscript. Additionally, we thank Professor Huebner and Professor Hansjuerg Alder in the Molecular Virology, Immunology and Medical Genetics Human Cancer Genetics Program, Department of the Ohio State University, who kindly performed the RT-PCR experimental stage in the Comprehensive Cancer Center for in this study. This study was supported by a grant from the Scientific Research Projects Foundation (BAP) of the Uludag University of Turkey [Project No: UAP(T)-2011/1].
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
E. Erturk and G. Cecener have contributed equally to this work.
Rights and permissions
About this article
Cite this article
Erturk, E., Cecener, G., Egeli, U. et al. Expression status of let-7a and miR-335 among breast tumors in patients with and without germ-line BRCA mutations. Mol Cell Biochem 395, 77–88 (2014). https://doi.org/10.1007/s11010-014-2113-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-014-2113-4