Abstract
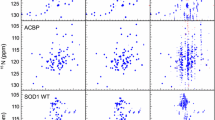
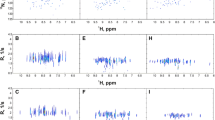
The application of non-uniform sampling (NUS) to relaxation experiments traditionally used to characterize the fast internal motion of proteins is quantitatively examined. Experimentally acquired Poisson-gap sampled data reconstructed with iterative soft thresholding are compared to regular sequentially sampled (RSS) data. Using ubiquitin as a model system, it is shown that 25 % sampling is sufficient for the determination of quantitatively accurate relaxation rates. When the sampling density is fixed at 25 %, the accuracy of rates is shown to increase sharply with the total number of sampled points until eventually converging near the inherent reproducibility of the experiment. Perhaps contrary to some expectations, it is found that accurate peak height reconstruction is not required for the determination of accurate rates. Instead, inaccuracies in rates arise from inconsistencies in reconstruction across the relaxation series that primarily manifest as a non-linearity in the recovered peak height. This indicates that the performance of an NUS relaxation experiment cannot be predicted from comparison of peak heights using a single RSS reference spectrum. The generality of these findings was assessed using three alternative reconstruction algorithms, eight different relaxation measurements, and three additional proteins that exhibit varying degrees of spectral complexity. From these data, it is revealed that non-linearity in peak height reconstruction across the relaxation series is strongly correlated with errors in NUS-derived relaxation rates. Importantly, it is shown that this correlation can be exploited to reliably predict the performance of an NUS-relaxation experiment by using three or more RSS reference planes from the relaxation series. The RSS reference time points can also serve to provide estimates of the uncertainty of the sampled intensity, which for a typical relaxation times series incurs no penalty in total acquisition time.






Similar content being viewed by others
References
Aoto PC, Fenwick RB, Kroon GJA, Wright PE (2014) Accurate scoring of non-uniform sampling schemes for quantitative NMR. J Magn Reson 246:31–35. doi:10.1016/j.jmr.2014.06.020
Barna JCJ, Laue ED, Mayger MR, Skilling J, Worrall SJP (1987) Exponential sampling, an alternative method for sampling in two-dimensional NMR experiments. J Magn Reson 73:69–77. doi:10.1016/0022-2364(87)90225-3
Becker S, Bobin J, Candes EJ (2011) NESTA: a fast and accurate first-order method for sparse recovery SIAM. J Imaging Sci 4:1–39. doi:10.1137/090756855
Bodenhausen G, Ernst RR (1982) Direct determination of rate constants of slow dynamic processes by two-dimensional accordion spectroscopy in nuclear magnetic-resonance. J Am Chem Soc 104:1304–1309. doi:10.1021/ja00369a027
Boggs PT, Rogers JE (1990) Orthogonal distance regression. Contemp Math 112:186
Candes EJ, Wakin MB (2008) An introduction to compressive sampling. ISPM 25:21–30. doi:10.1109/Msp.2007.914731
Candes EJ, Romberg JK, Tao T (2006) Stable signal recovery from incomplete and inaccurate measurements. Commun Pure Appl Math 59:1207–1223. doi:10.1002/cpa.20124
Candes EJ, Wakin MB, Boyd SP (2008) Enhancing sparsity by reweighted l(1) minimization. J Fourier Anal Appl 14:877–905. doi:10.1007/s00041-008-9045-x
Clore GM, Szabo A, Bax A, Kay LE, Driscoll PC, Gronenborn AM (1990) Deviations from the simple 2-parameter model-free approach to the interpretation of N-15 nuclear magnetic-relaxation of proteins. J Am Chem Soc 112:4989–4991. doi:10.1021/ja00168a070
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPIPE—a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293. doi:10.1007/bf00197809
Dodevski I, Nucci NV, Valentine KG, Sidhu GK, O’Brien ES, Pardi A, Wand AJ (2014) Optimized reverse micelle surfactant system for high-resolution NMR spectroscopy of encapsulated proteins and nucleic acids dissolved in low viscosity fluids. J Am Chem Soc 136:3465–3474. doi:10.1021/ja410716w
Donoho DL (1995) De-noising by soft-thresholding. IEEE Trans Inf Theory 41:613–627. doi:10.1109/18.382009
Donoho DL, Stark PB (1989) Uncertainty principles and signal recovery SIAM. J Appl Math 49:906–931. doi:10.1137/0149053
Drori I (2007) Fast l(1) minimization by iterative thresholding for multidimensional NMR Spectroscopy. EURASIP J Adv Signal Process. doi:10.1155/2007/20248
Farrow NA et al (1994) Backbone dynamics of a free and a phosphopeptide-complexed Src homology-2 domain studied by N-15 NMR relaxation. Biochemistry 33:5984–6003. doi:10.1021/bi00185a040
Ferrage F, Piserchio A, Cowburn D, Ghose R (2008) On the measurement of N-15–{H-1} nuclear Overhauser effects. J Magn Reson 192:302–313. doi:10.1016/j.jmr.2008.03.011
Findeisen M, Brand T, Berger S (2007) A H-1-NMR thermometer suitable for cryoprobes. Magn Reson Chem 45:175–178. doi:10.1002/mrc.1941
Frederick KK, Marlow MS, Valentine KG, Wand AJ (2007) Conformational entropy in molecular recognition by proteins. Nature 448:325-U323. doi:10.1038/nature05959
Fu Y, Kasinath V, Moorman VR, Nucci NV, Hilser VJ, Wand AJ (2012) Coupled motion in proteins revealed by pressure perturbation. J Am Chem Soc 134:8543–8550. doi:10.1021/ja3004655
Gledhill JM Jr, Walters BT, Wand AJ (2009) AMORE-HX: a multidimensional optimization of radial enhanced NMR-sampled hydrogen exchange. J Biomol NMR 45:233–239. doi:10.1007/s10858-009-9357-4
Harden BJ, Frueh DP (2014) SARA: a software environment for the analysis of relaxation data acquired with accordion spectroscopy. J Biomol NMR 58:83–99. doi:10.1007/s10858-013-9807-x
Henzler-Wildman KA, Lei M, Thai V, Kerns SJ, Karplus M, Kern D (2007) A hierarchy of timescales in protein dynamics is linked to enzyme catalysis. Nature 450:913-U927. doi:10.1038/nature06407
Hiller S, Ibraghimov I, Wagner G, Orekhov VY (2009) Coupled decomposition of four-dimensional NOESY spectra. J Am Chem Soc 131:12970–12978. doi:10.1021/ja902012x
Hoch JC (1985) Maximum-entropy signal-processing of two-dimensional NMR data. J Magn Reson 64:436–440. doi:10.1016/0022-2364(85)90106-4
Hyberts SG, Takeuchi K, Wagner G (2010) Poisson-gap sampling and forward maximum entropy reconstruction for enhancing the resolution and sensitivity of protein NMR data. J Am Chem Soc 132:2145–2147. doi:10.1021/ja908004w
Hyberts SG, Arthanari H, Wagner G (2012a) Applications of non-uniform sampling and processing. In: Novel sampling approaches in higher dimensional NMR, vol 316. Top Curr Chem, pp 125–148. doi:10.1007/128_2011_187
Hyberts SG, Milbradt AG, Wagner AB, Arthanari H, Wagner G (2012b) Application of iterative soft thresholding for fast reconstruction of NMR data non-uniformly sampled with multidimensional Poisson gap scheduling. J Biomol NMR 52:315–327. doi:10.1007/s10858-012-9611-z
Hyberts SG, Robson SA, Wagner G (2013) Exploring signal-to-noise ratio and sensitivity in non-uniformly sampled multi-dimensional NMR spectra. J Biomol NMR 55:167–178. doi:10.1007/s10858-012-9698-2
Hyberts SG, Arthanari H, Robson SA, Wagner G (2014) Perspectives in magnetic resonance: NMR in the post-FFT era. J Magn Reson 241:60–73. doi:10.1016/j.jmr.2013.11.014
Igumenova TI, Frederick KK, Wand AJ (2006) Characterization of the fast dynamics of protein amino acid side chains using NMR relaxation in solution. Chem Rev 106:1672–1699. doi:10.1021/cr040422h
Jaravine VA, Zhuravleva AV, Permi P, Ibraghimov I, Orekhov VY (2008) Hyperdimensional NMR spectroscopy with nonlinear sampling. J Am Chem Soc 130:3927–3936. doi:10.1021/ja077282o
Jarymowycz VA, Stone MJ (2006) Fast time scale dynamics of protein backbones: NMR relaxation methods, applications, and functional consequences. Chem Rev 106:1624–1671. doi:10.1021/cr040421p
Jones JA (1997) Optimal sampling strategies for the measurement of relaxation times in proteins. J Magn Reson 126:283–286. doi:10.1006/jmre.1997.1167
Kamath U, Shriver JW (1989) Characterization of themotropic state changes in myosin subfragment-1 and heavy-meromyosin by UV difference spectroscopy. J Biol Chem 264:5586–5592
Kay LE, Muhandiram DR, Farrow NA, Aubin Y, Forman-Kay JD (1996) Correlation between dynamics and high affinity binding in an SH2 domain interaction. Biochemistry 35:361–368. doi:10.1021/bi9522312
Kazimierczuk K, Orekhov VY (2011a) Accelerated NMR spectroscopy by using compressed sensing. Angew Chem Int Ed Engl 50:5556–5559. doi:10.1002/anie.201100370
Kazimierczuk K, Orekhov VY (2011b) Accelerated NMR spectroscopy by using compressed sensing. Angew Chem 50:5556–5559. doi:10.1002/anie.201100370
Kazimierczuk K, Orekhov VY (2012) A comparison of convex and non-convex compressed sensing applied to multidimensional NMR. J Magn Reson 223:1–10. doi:10.1016/j.jmr.2012.08.001
Kranz JK, Lee EK, Nairn AC, Wand AJ (2002) A direct test of the reductionist approach to structural studies of calmodulin activity—relevance of peptide models of target proteins. J Biol Chem 277:16351–16354. doi:10.1074/jbc.C200139200
Lakomek N-A, Ying J, Bax A (2012) Measurement of N-15 relaxation rates in perdeuterated proteins by TROSY-based methods. J Biomol NMR 53:209–221. doi:10.1007/s10858-012-9626-5
Laue ED, Skilling J, Staunton J, Sibisi S, Brereton RG (1985) Maximum-entropy method in nuclear magnetic-resonance spectroscopy. J Magn Reson 62:437–452
Lee AL, Wand AJ (1999) Assessing potential bias in the determination of rotational correlation times of proteins by NMR relaxation. J Biomol NMR 13:101–112. doi:10.1023/a:1008304220445
Li Z, Raychaudhuri S, Wand AJ (1996) Insights into the local residual entropy of proteins provided by NMR relaxation. Protein Sci 5:2647–2650. doi:10.1002/pro.5560051228
Linnet TE, Teilum K (2016) Non-uniform sampling of NMR relaxation data. J Biomol NMR. doi:10.1007/s10858-016-0020-6
Lipari G, Szabo A (1982a) Model-free approach to the interpretation of nuclear magnetic-resonance relaxation in macromolecules 1. Theory and range of validity. J Am Chem Soc 104:4546–4559. doi:10.1021/ja00381a009
Lipari G, Szabo A (1982b) Model-free approach to the interpretation of nuclear magnetic-resonance relaxation in macromolecules 2. Analysis of experimental results. J Am Chem Soc 104:4559–4570. doi:10.1021/ja00381a010
Logan BF (1965) Properties of high-pass signals. Ph.D. thesis. Columbia University
Long D, Delaglio F, Sekhar A, Kay LE (2015) Probing invisible, excited protein states by non-uniformly sampled pseudo-4D CEST spectroscopy. Angew Chem 54:10507–10511. doi:10.1002/anie.201504070
Marlow MS, Dogan J, Frederick KK, Valentine KG, Wand AJ (2010) The role of conformational entropy in molecular recognition by calmodulin. Nat Chem Biol 6:352–358. doi:10.1038/nchembio.347
Matsuki Y, Eddy MT, Herzfeld J (2009) Spectroscopy by integration of frequency and time domain information for fast acquisition of high-resolution dark spectra. J Am Chem Soc 131:4648–4656. doi:10.1021/ja807893k
Matsuki Y, Eddy MT, Griffin RG, Herzfeld J (2010) Rapid three-dimensional MAS NMR spectroscopy at critical sensitivity. Angew Chem 49:9215–9218. doi:10.1002/anie.201003329
Matsuki Y, Konuma T, Fujiwara T, Sugase K (2011) Boosting protein dynamics studies using quantitative nonuniform sampling NMR spectroscopy. J Phys Chem B 115:13740–13745. doi:10.1021/jp2081116
Mayzel M, Rosenlow J, Isaksson L, Orekhov VY (2014) Time-resolved multidimensional NMR with non-uniform sampling. J Biomol NMR 58:129–139. doi:10.1007/s10858-013-9811-1
Millet O, Muhandiram DR, Skrynnikov NR, Kay LE (2002) Deuterium spin probes of side-chain dynamics in proteins. 1. Measurement of five relaxation rates per deuteron in C-13-labeled and fractionally H-2-enriched proteins in solution. J Am Chem Soc 124:6439–6448. doi:10.1021/ja012497y
Mobli M, Hoch JC (2015) Nonuniform sampling and non-Fourier signal processing methods in multidimensional NMR. Prog Nucl Magn Reson Spectrosc 86–87:80. doi:10.1016/j.pnmrs.2015.02.001
Nesterov Y (2005) Smooth minimization of non-smooth functions. Math Program 103:127–152. doi:10.1007/s10107-004-0552-5
Nucci NV et al (2011) Optimization of NMR spectroscopy of encapsulated proteins dissolved in low viscosity fluids. J Biomol NMR 50:421–430. doi:10.1007/s10858-011-9528-y
Oliphant TE (2007) Python for scientific computing. Comput Sci Eng 9:10–20. doi:10.1109/MCSE.2007.58
Orekhov VY, Jaravine VA (2011) Analysis of non-uniformly sampled spectra with multi-dimensional decomposition. Prog Nucl Magn Reson Spectrosc 59:271–292. doi:10.1016/j.pnmrs.2011.02.002
Oyen D, Fenwick RB, Stanfield RL, Dyson HJ, Wright PE (2015) Cofactor-mediated conformational dynamics promote product release from Escherichia coli dihydrofolate reductase via an allosteric pathway. J Am Chem Soc 137:9459–9468. doi:10.1021/jacs.5b05707
Palmer MR, Suiter CL, Henry GE, Rovnyak J, Hoch JC, Polenova T, Rovnyak D (2015) Sensitivity of nonuniform sampling NMR. J Phys Chem B 119:6502–6515. doi:10.1021/jp5126415
Rovnyak D, Hoch JC, Stern AS, Wagner G (2004) Resolution and sensitivity of high field nuclear magnetic resonance spectroscopy. J Biomol NMR 30:1–10. doi:10.1023/b:jnmr.0000042946.04002.19
Ryabov Y, Clore GM, Schwieters CD (2012) Coupling between internal dynamics and rotational diffusion in the presence of exchange between discrete molecular conformations. J Chem Phys. doi:10.1063/1.3675602
Schmieder P, Stern AS, Wagner G, Hoch JC (1997) Quantification of maximum-entropy spectrum reconstructions. J Magn Reson 125:332–339. doi:10.1006/jmre.1997.1117
Sibisi S, Skilling J, Brereton RG, Laue ED, Staunton J (1984) Maximum-entropy signal-processing in practical NMR-spectroscopy. Nature 311:446–447. doi:10.1038/311446a0
Skelton NJ, Palmer AG, Akke M, Kordel J, Rance M, Chazin WJ (1993) Practical aspects of 2-dimensional proton-detected N-15 spin relaxation measurements. J Magn Reson, Ser B 102:253–264. doi:10.1006/jmrb.1993.1095
Stern AS, Donoho DL, Hoch JC (2007) NMR data processing using iterative thresholding and minimum l(1)-norm reconstruction. J Magn Reson 188:295–300. doi:10.1016/j.jmr.2007.07.008
Stevens SY, Sanker S, Kent C, Zuiderweg ERP (2001) Delineation of the allosteric mechanism of a cytidylyltransferase exhibiting negative cooperativity. Nat Struct Biol 8:947–952. doi:10.1038/nsb1101-947
Sun S, Gill M, Li Y, Huang M, Byrd RA (2015) Efficient and generalized processing of multidimensional NUS NMR data: the NESTA algorithm and comparison of regularization terms. J Biomol NMR 62:105–117. doi:10.1007/s10858-015-9923-x
Tugarinov V, Kay LE (2005) Quantitative C-13 and H-2 NMR relaxation studies of the 723-residue enzyme malate synthase g reveal a dynamic binding interface. Biochemistry 44:15970–15977. doi:10.1021/bi0519809
Tugarinov V, Kanelis V, Kay LE (2006) Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat Protoc 1:749–754. doi:10.1038/nprot.2006.101
Tzeng S-R, Kalodimos CG (2009) Dynamic activation of an allosteric regulatory protein. Nature 462:368-U139. doi:10.1038/nature08560
Tzeng S-R, Kalodimos CG (2012) Protein activity regulation by conformational entropy. Nature 488:236–240. doi:10.1038/nature11271
Wand AJ (2001) Dynamic activation of protein function: a view emerging from NMR spectroscopy. Nat Struct Biol 8:926–931. doi:10.1038/nsb1101-926
Wand AJ, Urbauer JL, McEvoy RP, Bieber RJ (1996) Internal dynamics of human ubiquitin revealed by C-13-relaxation studies of randomly fractionally labeled protein. Biochemistry 35:6116–6125. doi:10.1021/bi9530144
Wand AJ, Moorman VR, Harpole KW (2013) A surprising role for conformational entropy in protein function. In: Dynamics in enzyme catalysis, vol 337. Top Curr Chem, pp 69–94. doi:10.1007/128_2012_418
Zidek L, Novotny MV, Stone MJ (1999) Increased protein backbone conformational entropy upon hydrophobic ligand binding. Nat Struct Biol 6:1118–1121
Acknowledgments
This work was supported by NIH GM102447 awarded to A.J.W. M.A.S. is an NIH predoctoral trainee (GM008275). We thank Prof. Gerhard Wagner and Dr. Scott Robson for providing the hmsIST software and early assistance with data processing. We also thank Dr. Sven Hyberts and Dr. Haribabu Arthanari for helpful discussion. We are grateful to Dr. Kathy Valentine for technical assistance and extensive discussion. We gratefully acknowledge Li Liang for providing the AK sample, Dr. Nathaniel Nucci for providing the ubiquitin sample in 30 % glycerol, and Bryan Marques for providing the MBP sample.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Stetz, M.A., Wand, A.J. Accurate determination of rates from non-uniformly sampled relaxation data. J Biomol NMR 65, 157–170 (2016). https://doi.org/10.1007/s10858-016-0046-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-016-0046-9