Abstract
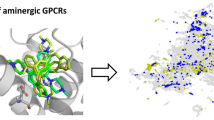
A physicochemical property-based desirability scoring scheme for fragment-based drug discovery was developed for class A aminergic GPCR targeted fragment libraries. Physicochemical property distributions of known aminergic GPCR-active fragments from the ChEMBL database were examined and used for a desirability function-based score. Property-distributions such as log D (at pH 7.4), PSA, pKa (strongest basic center), number of nitrogen atoms, number of oxygen atoms, and the number of rotatable bonds were combined into a desirability score (FrAGS). The validation of the scoring scheme was carried out using both public and proprietary experimental screening data. The scoring scheme is suitable for the design of aminergic GPCR targeted fragment libraries and might be useful for preprocessing fragments before structure based virtual or wet screening.





Similar content being viewed by others
Abbreviations
- FrAGS:
-
Fragment Aminergic GPCR Score
- GPCR:
-
G-protein coupled receptor
- PSA:
-
Polar surface area
- FBDD:
-
Fragment-based drug discovery
- HTS:
-
High-throughput screening
- FS:
-
Fragment-screening
- 7TM:
-
Seven-transmembrane
- SILE:
-
Size-independent ligand-efficiency
- SMILES:
-
Simplified molecular-input line-entry system
- EF:
-
Enrichment factor
- TPR:
-
True positive rate
- TNR:
-
True negative rate
- FPR:
-
False positive rate
- FNR:
-
False negative rate
- ROC:
-
Receiver operating characteristic
- TAAR1 :
-
Trace-amine receptor subtype 1
- 5HT1 :
-
5-Hydroxy-tryptamine receptor subtype 1
References
Fink T, Bruggesser H, Reymond JL (2005) Virtual exploration of the small-molecule chemical universe below 160 daltons. Angew Chem Int Ed 44:1504–1508
Bohacek RS, McMartin C, Guida WC (1996) The art and practice of structure-based drug design: a molecular modeling perspective. Med Res Rev 16:3–50
Leeson PD, Springthorpe B (2007) The influence of drug-like concepts on decision-making in medicinal chemistry. Nat Rev Drug Discov 6:881–890
Hann MM, Keserű GM (2012) Finding the sweet spot: the role of nature and nurture in medicinal chemistry. Nat Rev Drug Discov 11:355–365
Andrews SP, Brown GA, Christopher JA (2014) Structure-based and fragment-based GPCR drug discovery. ChemMedChem 9:256–275
ChEMBL GPCR SARfari home page. https://www.ebi.ac.uk/chembl/sarfari/gpcrsarfari
Knime Desktop (Konstanz Information Miner), version 2.9.1 (2014)
Willem J, Nissink M (2009) Simple size-independent measure of ligand efficiency. J Chem Inf Model 49(6):1617–1622
Hann MM, Leach AR, Green DVS, Oprea TI (eds) (2005) Methods and principles in medicinal chemistry, vol 23. Wiley-VCH, Weinheim, pp 43–57
Hopkins AL, Groom CR, Alex A (2004) Ligand efficiency: a useful metric for lead selection. Drug Discov Today 9:430–431
Congreve et al (2003) A ‘rule of three’ for fragment-based lead discovery. Drug Discov Today 8:876–877
GPCRDB information system for G protein-coupled receptors home page. http://www.gpcr.org/7tm/
ChEMBL home page. ftp://ebi.ac.uk/pub/databases/chembl/ChEMBLdb/
RDKit: cheminformatics and machine learning software home page. http://www.rdkit.org/
Balakin KV, Tkachenko SE, Lang SA, Okun I, Ivashchenko AA, Savchuk NP (2002) Property-based design of GPCR-targeted library. J Chem Inf Comput Sci 42:1332–1342
Marvin, version 5.2, JChem for Excel (2014) Chemaxon, Budapest
Statistica 12, Statsoft home page. http://www.statsoft.com/
Segall MD (2012) Multi-parameter optimization: identifying high quality compounds with a balance of properties. Curr Pharm Des 18:1292–1310
Harrington EC (1965) The desirability function. Ind Qual Control 21:494–498
Wager TT, Hou X, Verhoest PR, Villalobos A (2010) Moving beyond rules: the development of a central nervous system multiparameter optimization (CNS MPO) approach to enable alignment of drug-like properties. ACS Chem Neurosci 1:435–449
PubChem home page. https://pubchem.ncbi.nlm.nih.gov/
Tanimoto distance as defined in the “Distance Matrix Calculate” node of Knime
Acknowledgments
We thank for Gedeon Richter Plc for providing GPCR-related fragment-screening and high-throughput screening data for the validation studies and particularly Márton Vass for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kelemen, Á.A., Ferenczy, G.G. & Keserű, G.M. A desirability function-based scoring scheme for selecting fragment-like class A aminergic GPCR ligands. J Comput Aided Mol Des 29, 59–66 (2015). https://doi.org/10.1007/s10822-014-9804-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10822-014-9804-5