Abstract
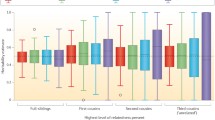
Apart from parent-offspring pairs and clones, relative pairs vary in the proportion of the genome that they share identical by descent. In the past, quantitative geneticists have used the expected value of sharing genes by descent to estimate genetic parameters and predict breeding values. With the possibility to genotype individuals for many markers across the genome it is now possible to empirically estimate the actual relationship between relatives. We review some of the theory underlying the variation in genetic identity, show applications to estimating genetic variance for height in humans and discuss other applications.



Similar content being viewed by others
References
Abecasis GR, Cherny SS, Cookson WO, Cardon LR (2002) Merlin-rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 30:97–101
Bulmer MG (1985) The mathematical theory of quantitative genetics. Clarendon Press, Oxford
Falconer DS, Mackay TFC (1996) Introduction to quantitative genetics, 4th edn. Longman, Harlow
Franklin IR (1977) The distribution of the proportion of the genome which is homozygous by descent in inbred individuals. Theor Popul Biol 11:60–80
Frazer KA, Ballinger DG, Cox DR, Hinds DA, Stuve LL, Gibbs RA, Belmont JW, Boudreau A, Hardenbol P, Leal SM et al (2007) A second generation human haplotype map of over 3.1 million SNPs. Nature 449:851–861
Gagnon A, Beise J, Vaupel JW (2005) Genome-wide identity-by-descent sharing among CEPH siblings. Genet Epidemiol 29:215–224
Guo SW (1994) Computation of identity-by-descent proportions shared by two siblings. Am J Hum Genet 54:1104–1109
Guo SW (1995) Proportion of genome shared identical by descent by relatives: concept, computation, and applications. Am J Hum Genet 56:1468–1476
Guo SW (1996) Variation in genetic identity among relatives. Hum Hered 46:61–70
Haseman JK, Elston RC (1972) The investigation of linkage between a quantitative trait and a marker locus. Behav Genet 2:3–19
Hill WG (1993a) Variation in genetic composition in backcrossing programs. J Hered 84:212–213
Hill WG (1993b) Variation in genetic identity within kinships. Heredity 71:652–653
Kent JW Jr, Dyer TD, Blangero J (2005) Estimating the additive genetic effect of the X chromosome. Genet Epidemiol 29:377–388
Kong X, Murphy K, Raj T, He C, White PS, Matise TC (2004) A combined linkage-physical map of the human genome. Am J Hum Genet 75:1143–1148
Lynch M, Walsh B (1998) Genetics and analysis of quantitative traits. Sinauer Associates, Sunderland
Meuwissen TH, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Risch N, Lange K (1979) Application of a recombination model in calculating the variance of sib pair genetic identity. Ann Hum Genet 43:177–186
Stam P (1980) The distribution of the fraction of the genome identical by descent in finite random mating populations. Genet Res 35:131–155
Stam P, Zeven AC (1981) The theoretical proportion of the donor genome in near-isogenic lines of self-fertilizers bred by backcrossing. Euphytica 30:227–238
Suarez BK, Reich T, Fishman PM (1979) Variability in sib pair genetic identity. Hum Hered 29:37–41
Thomas SC (2005) The estimation of genetic relationships using molecular markers and their efficiency in estimating heritability in natural populations. Philos Trans R Soc Lond B Biol Sci 360:1457–1467
Thomas SC, Pemberton JM, Hill WG (2000) Estimating variance components in natural populations using inferred relationships. Heredity 84(Pt 4):427–436
Thomas SC, Coltman DW, Pemberton JM (2002) The use of marker-based relationship information to estimate the heritability of body weight in a natural population: a cautionary tale. J Evol Biol 15:92–99
Visscher PM, Hopper JL (2001) Power of regression and maximum likelihood methods to map QTL from sib-pair and DZ twin data. Ann Hum Genet 65:583–601
Visscher PM, Medland SE, Ferreira MA, Morley KI, Zhu G, Cornes BK, Montgomery GW, Martin NG (2006) Assumption-free estimation of heritability from genome-wide identity-by-descent sharing between full siblings. PLoS Genet 2:e41
Visscher PM, Macgregor S, Benyamin B, Zhu G, Gordon S, Medland S, Hill WG, Hottenga JJ, Willemsen G, Boomsma DI, Liu Y-Z, Deng HW, Montgomery GW, Martin NG (2007) Genome partitioning of genetic variation for height from 11, 214 sibling pairs. Am J Hum Genet 81:1104–1110
Xu S (2006) Population genetics: separating nurture from nature in estimating heritability. Heredity 97:256–257
Acknowledgements
We acknowledge support from the Australian National Health and Medical Research Council (grants 389892 and 442915) and the Australian Research Council (grant DP0770096), and thank Bill Hill, Mike Goddard, Nick Martin and Naomi Wray for many stimulating discussions and two referees for helpful comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Visscher, P.M. Whole genome approaches to quantitative genetics. Genetica 136, 351–358 (2009). https://doi.org/10.1007/s10709-008-9301-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10709-008-9301-7