Abstract
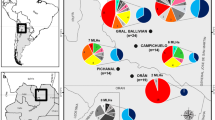
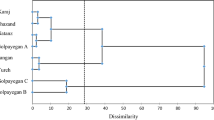
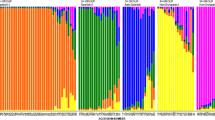
Grape downy mildew is a destructive oomycete disease worldwide for viticulture. A total of 440 infected leaf lesions were collected from nineteen vineyards distributing in the major viticulture regions in China. Six specific SSR markers were used to genetically characterize the Plasmopara viticola populations. A total of 60 alleles were generated and 324 distinct genotypes were identified based on these multi-locus markers. Most populations had high levels of genetic and genotypic diversity. AMOVA analysis showed that majority of the total genetic diversity was distributed within the vineyards while moderate to high genetic differentiation was also observed among some P. viticola populations. Three populations (BJ, GX, and GS2) were clearly separated from the rest of the populations by using PCA and Nei’s genetic distance analysis. Mantel test showed no significant correlation between unbiased Nei’s genetic distance and geographical distance.



Similar content being viewed by others
References
Altizer, S., Harvell, D., & Friedle, E. (2003). Rapid evolutionary dynamics and disease threats to biodiversity. Trends in Ecology & Evolution, 18(11), 589–596.
Blum, M., Waldner, M., & Gisi, U. (2010). A single point mutation in the novel PvCesA3 gene confers resistance to the carboxylic acid amide fungicide mandipropamid in Plasmopara viticola. Fungal. Genetics and Biology, 47(6), 499–510.
Chen, W. J., Delmotte, F., Cervera, S. R., Douence, L., Greif, C., & Corio-Costet, M. F. (2007). At least two origins of fungicide resistance in grapevine downy mildew populations. Applied and Environmental Microbiology, 73(16), 5162–5172.
Delmotte, F., Chen, W., Richard-Cervera, S., Greif, C., Papura, D., Giresse, X., Mondor-Genson, G., & Corio-Costet, M. (2006). Microsatellite DNA markers for Plasmopara viticola, the causal agent of downy mildew of grapes. Molecular Ecology Notes, 6(2), 379–381.
Delmotte, F., Machefer, V., Giresse, X., Richard-Cervera, S., Latorse, M., & Beffa, R. (2011). Characterization of single-nucleotide-polymorphism markers for Plasmopara viticola, the causal agent of grapevine downy mildew. Applied and Environmental Microbiology, 77(21), 7861–7863.
Dewar, R. (1907). The vine mildew—Plasmopara viticola. Agricultural Journal of the Cape of Good Hope, 31, 324–329.
Drenth, A., & Goodwin, S. B. (1999). Population structure of Oomycetes. In J. J. Worral (Ed.), Structure and dynamics of fungal populations (pp. 195–224). Amstedam: Springer.
Dutech, C., Enjalbert, J., Fournier, E., Delmotte, F., Barres, B., Carlier, J., Tharreau, D., & Giraud, T. (2007). Challenges of microsatellite isolation in fungi. Fungal Genetics and Biology, 44(10), 933–949.
Fontaine, M. C., Austerlitz, F., Giraud, T., Labbe, F., Papura, D., Richard-Cervera, S., & Delmotte, F. (2013). Genetic signature of a range expansion and leap-frog event after the recent invasion of Europe by the grapevine downy mildew pathogen Plasmopara viticola. Molecular Ecology, 22, 2771–2286.
FRAC. (2008). Retrieved October 15, 2009, from http://www.frac.info.
Gennadios, P. G. (1889). About the downy mildew of the grapevine. Greek Agriculture, 8, 297–307.
Gessler, C., Pertot, I., & Perazzolli, M. (2011). Plasmopara viticola: a review of knowledge on downy mildew of grapevine and effective disease management. Phytopathologia Mediterranea, 50(1), 3–44.
Gobbin, D., Bleyer, G., Keil, S., Kassemeyer, H., & Gessler, C. (2007). Evidence for sporangial dispersal leading to a single infection event and a sudden high incidence of grapevine downy mildew. Plant Pathology, 56(5), 843–847.
Gobbin, D., Jermini, M., Loskill, B., Pertot, I., Raynal, M., & Gessler, C. (2005). Importance of secondary inoculum of Plasmopara viticola to epidemics of grapevine downy mildew. Plant Pathology, 54(4), 522–534.
Gobbin, D., Pertot, I., & Gessler, C. (2003a). Genetic structure of a Plasmopara viticola population in an isolated Italian mountain vineyard. Journal of Phytopathology, 151(11–12), 636–646.
Gobbin, D., Pertot, I., & Gessler, C. (2003b). Identification of microsatellite markers for Plasmopara viticola and establishment of high throughput method for SSR analysis. European Journal of Plant Pathology, 109(2), 153–164.
Gobbin, D., Rumbou, A., Linde, C. C., & Gessler, C. (2006). Population genetic structure of Plasmopara viticola after 125 years of colonization in European vineyards. Molecular plant pathology, 7(6), 519–531.
Hug, F. (2005). Genetic structure and epidemiology of Plasmopara viticola populations from Australian grape growing regions. Dissertation, ETH Zurich, Switzerland.
Ji, L. L., Li, H. Q., Ren, Y. Z., Qi, L. M., Zhao, B. L., Li, G. Y., & Li, C. (2012). Dynamics of sporangium diffusion of Plasmopara viticola and its correlation with disease incidence in vineyard. Journal of Fruit Science, 29(1), 94–98.
Kamvar, Z. N., & Grünwald, N. J. (2013). Poppr 1.0. 0: An R package for genetic analysis of populations with mixed (clonal/sexual) reproduction.http://www.inside-r.org/packages/poppr/versions/1-0-0.
Koopman, T., Linde, C. C., Fourie, P. H., & McLeod, A. (2007). Population genetic structure of Plasmopara viticola in the Western Cape Province of South Africa. Molecular Plant Pathology, 8(6), 723–736.
Kump, I., Blaise, P., & Gessler, C. (1998). The use of RAPD-markers to estimate genetic diversity of Plasmopara viticola in a single vineyard. In Third Int. Workshop on Grapevine Downy and Powdery Mildew, 50(176), 72–73.
Lafon, R., & Bulit, J. (1981). Downy mildew of the vine. In I. D. M. Spencer (Ed.), The Downy Mildews: London: Academic press.
Lafon, R., & Clerjeau, M. (1988). Downy mildew. In R. Pearson & A. Goheen (Eds.), Compendium of grape diseases (pp. 11–13). St. Paul: APS Press.
Lei, B. Z. (2004). The investigation and study of the main fungus diseases on grape in Xinjiang. Dissertation, Shihezi University.
Liu, L., & Zhang, Z. (2010). Studies on Grapevine Diseases in Weibei Rainfed Highland (Jingyang). Northern Horticulture, 18, 174–177.
Müller, K., & Sleumer, H. (1934). Biologische Untersuchungen über die Peronosporakrankheit des Weinstocks. Landwirtschaftliche Jahrbücher, 4, 509–576.
Matasci, C. L., Jermini, M., Gobbin, D., & Gessler, C. (2010). Microsatellite based population structure of Plasmopara viticola at single vine scale. European Journal of Plant Pathology, 127(4), 501–508.
McRitchie, J. J. (1978). Downy mildew of grape. Plant Pathology Circular, 193.
Peakall, R., & Smouse, P. E. (2012). GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research—an update. Bioinformatics, 28(19), 2537–2539.
Peakall, R., Smouse, P. E., & Huff, D. (1995). Evolutionary implications of allozyme and RAPD variation in diploid populations of dioecious buffalograss Buchloe dactyloides. Molecular Ecology, 4(2), 135–148.
Qi, H. X., Qi, Y. S., Zhang, J. Z., Wang, T. K., & Li, S. M. (2009). Outbreak Causes and Seasonal Epidemic Dynamics of Wine Grape Downy Mildew. Nonwood Forest Research, 27(1), 57–60.
Rouxel, M., Mestre, P., Baudoin, A., Carisse, O., Delière, L., Ellis, M. A., Gadoury,D., Lu,J., Nita,M., Richard-Cervera,S., Schilder,A., Wise,A. & Delmotte,F. (2013a). Geographic Distribution of Species of Plasmopara viticola Causing Downy Mildew on Wild and Cultivated Grapes in Eastern North America. Phytopathology (in press).
Rouxel, M., Mestre, P., Comont, G., Lehman, B. L., Schilder, A., & Delmotte, F. (2013b). Phylogenetic and experimental evidence for host‐specialized cryptic species in a biotrophic oomycete. New Phytologist, 197(1), 251–263.
Rouxel, M., Papura, D., Nogueira, M., Machefer, V., Dezette, D., Richard-Cervera, S., Carrere, S., Mestre, P., & Delmotte, F. (2012). Microsatellite markers for characterization of native and introduced populations of Plasmopara viticola, the causal agent of grapevine downy mildew. Applied and Environmental Microbiology, 78(17), 6337–6340.
Rumbou, A., & Gessler, C. (2004). Genetic dissection of Plasmopara viticola population from a Greek vineyard in two consecutive years. European Journal of Plant Pathology, 110(4), 379–392.
Rumbou, A., & Gessler, C. (2006). Particular structure of Plasmopara viticola populations evolved under Greek island conditions. Phytopathology, 96(5), 501–509.
Scherer, E., & Gisi, U. (2006). Characterization of genotype and mating type in European isolates of Plasmopara viticola. Journal of Phytopathology, 154(7–8), 489–495.
Schröder, S., Telle, S., Nick, P., & Thines, M. (2011). Cryptic diversity of Plasmopara viticola (Oomycota, Peronosporaceae) in North America. Organisms Diversity & Evolution, 11(1), 3–7.
Somasundaram, S., & Kalaiselvam, M. (2011). Molecular tools for assessing genetic diversity. International Training Course on Mangroves and Biodiversity, Annamalai University, India, 82–91.
Stark-Urnau, M., Seidel, M., Kast, W., & Gemmrich, A. (2000). Studies on the genetic diversity of primary and secondary infections of Plasmopara viticola using RAPD/PCR. Vitis, 39(4), 163–166.
Sun, H. Y., Wang, H. C., Stammler, G., Ma, J. X., Liu, J. M., & Zhou, M. G. (2010). Sensitivity of Chinese Isolates of Plasmopara viticola to Metalaxyl and Dimethomorph. Journal of Phytopathology, 158(6), 450–452.
Wan, Y. Z., Schwaninger, H., Li, D., Simon, C., Wang, Y. J., & Zhang, C. H. (2008). A review of taxonomic research in Chinese wild grapes. Vitis, 47(2), 81–88.
Weir, B. S., & Cockerham, C. C. (1984). Estimating F-statistics for the analysis of population structure. evolution, 1358–1370.
Williams, M. G. (2005). Impact of environmental conditions on the infection behaviour of western australian strains of plasmopara viticola, causal agent of downy mildew in grapevines. Dissertation, University of Western Australia.
Wong, F., Burr, H., & Wilcox, W. (2001). Heterothallism in Plasmopara viticola. Plant Pathology, 50(4), 427–432.
Wright, S. (1978). Variability Within and Among Populations. Evolution and the genetics of populations. Chicago: University of Chicago Press.
Yeh, F. C., Yang, R., Boyle, T. J., Ye, Z., & Xiyan, J. M. (2000). POPGENE 32, Microsoft Window-based Freeware for Population Genetic Analysis, Version 1.32. Edmonton: Molecular Biology and Biotechnology Centre, University of Alberta.
Yin, L., Zhang, Y. L., & Lu, J. (2010). Comparision of different DNA extraction methods for Plasmopara viticola. Sino-overseas Grapevine and Wine, 3, 4–7.
Yu, Y., Zhang, Y., Yin, L., & Lu, J. (2012). The Mode of Host Resistance to Plasmopara viticola Infection of Grapevines. Phytopathology, 102(11), 1094–1101.
Acknowledgments
We are very grateful to many colleagues who assisted in the collection of P. viticolasamples from various locations throughout China. Dr. Delmotte (INRA, France) is gratefully acknowledged for providing the reference P. viticola isolates and valuable comments on preparation of the manuscript. Dr. Dry (CSIRO) kindly provides language polishing. This work was supported by the “948” Program, Ministry of Agriculture, China (Grant no. 2006-G26) and the China Agriculture Research System (Grant no. CARS-30-yz-2).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yin, L., Zhang, Y., Hao, Y. et al. Genetic diversity and population structure of Plasmopara viticola in China. Eur J Plant Pathol 140, 365–376 (2014). https://doi.org/10.1007/s10658-014-0470-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-014-0470-1