Abstract
We hypothesized that a subset of sporadic triple negative (TN) breast cancer patients whose tumors have defective DNA repair similar to BRCA1-associated tumors are more likely to exhibit up-regulation of DNA repair-related genes, anthracycline-sensitivity, and taxane-resistance. We derived a defective DNA repair gene expression signature of 334 genes by applying a previously published BRCA1-associated expression pattern to three datasets of sporadic TN breast cancers. We confirmed a subset of 69 of the most differentially expressed genes by quantitative RT-PCR, using a low density custom array (LDA). Next, we tested the association of this DNA repair microarray signature expression with pathologic response in neoadjuvant anthracycline trials of FEC (n = 50) and AC (n = 16), or taxane-based TET chemotherapy (n = 39). Finally, we collected paraffin-fixed, formalin-embedded biopsies from TN patients who had received neoadjuvant AC (n = 28), and tested the utility of the LDA to discriminate response. Correlation between RNA expression measured by the microarrays and 69-gene LDA was ascertained. This defective DNA repair microarray gene expression pattern was significantly associated with anthracycline response and taxane resistance, with the area under the ordinary receiver operating characteristic curve (AUC) of 0.61 (95% CI = 0.45–0.77), and 0.65 (95% CI = 0.46–0.85), respectively. From the FFPE samples, the 69-gene LDA could discriminate AC responders, with AUC of 0.79 (95% CI = 0.59–0.98). In conclusion, a promising defective DNA repair gene expression signature appears to differentiate TN breast cancers that are sensitive to anthracyclines and resistant to taxane-based chemotherapy, and should be tested in clinical trials with other DNA-damaging agents and PARP-1 inhibitors.
Similar content being viewed by others
Introduction
Despite all of the advances in understanding breast cancer biology, there has been little change in survival for women with metastatic breast cancer over the last several decades. There were approximately 180,000 new cases of breast cancer diagnosed in the United States in 2007. About 15% of these, accounting for 25,000 to 30,000 cases each year, are “triple-negative” (TN) in that the tumors do not express the estrogen receptor (ER), progesterone receptor (PR), or HER-2/neu. Currently, there are no approved targeted agents for the treatment of TN cancers [1, 2]. The identification of new therapeutic targets and predictive markers is crucial to improving the outcome of this subtype (1, 2).
Germline BRCA1 gene mutation carriers have a 50–80% lifetime risk of developing breast cancers, represent less than 3% of all women with breast cancer, and are generally TN [3]. Functionally, BRCA1 protein is involved in the repair of double-stranded DNA breaks (DSBs) by homologous recombination [4]. DNA-damaging drugs, such as anthracyclines and platinums, can cause DSBs and cancer cells lacking functional BRCA1 have been shown in preclinical studies to be highly sensitive to these DNA damaging agents [5–8]. Clinically, BRCA1-associated tumors have also been demonstrated to be more sensitive to these agents than sporadic-matched controls [9–13]. Because of the phenotypic and genotypic similarities, therapeutically exploiting defective DNA repair in BRCA1-associated tumors has been extended to the larger subset of sporadic TN tumors [14–19]. For example, the use of targeted small molecules like the polyadenosine 5′-diphosphoribose [poly-(ADP-ribose)] polymerase (PARP-1) inhibitors, first tested in germline cancers has also recently been expanded to include sporadic TN tumors.
In this study, we first set out to identify a gene expression signature that can potentially distinguish TN breast cancers into those that exhibit DNA repair defects similar to tumors with BRCA1 mutations from TN breast cancers that may not carry a deficiency in homologous-recombination DNA repair. Second, we confirmed these expression results by two different RNA platforms (gene expression microarray vs. 69-gene low density custom array, LDA). Third, we tested this defective DNA repair microarray gene expression signature and its association with treatment response in TN breast cancers with the hypothesis that patients with this signature will demonstrate sensitivity to agents that affect DNA repair like anthracyclines, but not to non-DNA damaging agents, like taxanes. Finally, we tested the 69-gene LDA on formalin-fixed, paraffin-embedded (FFPE) core biopsies obtained from women with received neoadjuvant anthracycline chemotherapy (n = 28).
Methods
We used six gene expression datasets obtained by microarray analysis of tumor specimens from a total of 307 patients with primary triple-negative breast cancer.
The training sets used to obtain the candidate genes were the Baylor College of Medicine (BCM) dataset 1 (BCM1), the Nederlands Kanker Instituut (NKI2) [20], and the Wang dataset (GSE2034) [21]. The two anthracycline-treated validation sets used were from Baylor College of Medicine dataset 2 (BCM2), and EORTC (GSE6861) [22, 23]. The BCM1 and BCM2 consist of data from a total of 84 patients with TN breast cancer, whose frozen tumor specimens were archived at BCM. The other four datasets are publically available. Microarray and clinical data for the Wang and EORTC patients are available at the Gene Expression Omnibus database http://www.ncbi.nlm.nih.gov/geo), using the associated GSE accession codes, GSE2034 and GSE6861, respectively. The NKI2 dataset was downloaded from the Rosetta Web site (http://www.rii.com). The BCM1 and BCM2 dataset contained 68 and 16 triple negative breast cancer samples, as defined by IHC. The Wang, NKI2, and EORTC datasets contained data from 49, 57, and 89 primary breast tumor samples, respectively, and were ER-negative and PR-negative by IHC. As HER2 status was unavailable in the Wang and NKI2 dataset, HER2-negative patients were identified by microarray data, excluding those samples with ERBB2 and GRB7 overexpression. As such, from the 69 ER-negative and PR-negative samples in the NKI2 dataset, 20 samples were excluded due to overexpression of ERBB2 and GRB7 and 19 out of 76 samples were excluded from the Wang dataset.
The validation neoadjuvant gene expression microarray studies were conducted on two datasets: BCM2 and EORTC contained data from 16 and 89 triple-negative breast tumor samples, respectively. The treatment received by patients in the BCM2 dataset was four cycles of doxorubicin and cyclophosphamide, 60 mg/m2 and 600 mg/m2, respectively, every 3 weeks (AC). The patients in the EORTC dataset were randomized to receive anthracycline chemotherapy of FEC (6 cycles of 500 mg/m2 fluorouracil, 100 mg/m2 epirubicin, and 500 mg/m2 cyclophosphamide every 3 weeks), or primarily taxane-based chemotherapy of TET (3 cycles of 100 mg/m2 docetaxel, followed by 3 cycles of 90 mg/m2 epirubicin plus 70 mg/m2 docetaxel). Pathologic response (pCR) was defined as the complete disappearance of all tumor in the breast in all data sets except BCM2 which also included minute foci of residual disease (<0.1 cm).
Gene expression analysis
For BCM1 and BCM2 datasets, microarray analysis was performed with Affymetrix U133A GeneChips (Affymetrix, Santa Clara, CA), as previously published [24, 25], obtained from samples from BCM, Houston, TX, and Mt Vernon Hospital, United Kingdom. The publicly available datasets consisted of both Affymetrix (Wang and EORTC) and Agilent arrays (NKI2), with several different chip designs. To simplify analysis, we used only the gene probes that were common in all datasets (Supplementary Data).
Identification of samples with a high likelihood of having defective DNA repair
BRCA1-associated triple-negative tumors are more likely to have a deficiency in homologous recombination and DNA repair deficiency than sporadic triple-negative tumors. Van’t Veer et al. published a set of 430 genes found by microarray data to be differentially expressed between BRCA1-associated ER-negative tumors and sporadic ER-negative tumors, and an optimal set of 100 genes was found to discriminate between BRCA1 and sporadic cases. Although these results have not been externally validated or disproven, we hypothesized that using the set of 430 genes, one could identify a subset of triple-negative tumors likely to have defective DNA repair, similar to BRCA1-associated tumors and hence are more likely to exhibit anthracycline-sensitivity, taxane-resistance, and up-regulation of DNA repair-related genes [26]. The list of 430 genes and their gene ID are listed in Supplementary Data Conversion from gene ID to Affymetrix probe set ID was done using Netaffx.net online tool (Supplementary Data). We used these candidate genes and applied it to the BCM1, NKI2, and Wang training datasets which included 68, 49, and 57 samples of triple negative tumors.
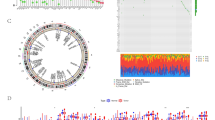
An algorithm was then introduced to rank the samples in each heat map (BCM1, Wang, and NKI). The genes for each sample were computed as the standardized gene-wise z-scores (underexpressed gene were multiplied by −1), and a total score was determined as the sum. The samples were then ranked according to the total score. The samples with the highest overall score have the gene expression pattern most similar to BRCA1-associated tumors, and those with the lowest score similar to “sporadic” tumors (Fig. 1, BCM1 Dataset). This ranking system was used in order to classify the samples in an objective manner. This algorithm was chosen, rather than metagene analysis, as a straightforward ranking system of differentially expressed genes equally, instead of metagene analysis where complex combinations of many genes and pathways are factored into the analysis. The ranked samples were then divided into high and low expression of genes with DNA repair signature based on the heat-map generated.
Identification of samples with BRCA1-like signature. a Heat map of 68 triple negative tumors from BCM ranked according to previously published BRCA1 gene signature. The samples are ranked according to an algorithm which places the tumors with a gene expression pattern most similar to that of sporadic tumors to the left, labeled with a green S, and the tumors with a BRCA1-like gene expression pattern to the right, labeled with a red B. b Three gene lists form each datasets (BCM1, Wang, NKI) were obtained. They were composed of the most differentially expressed genes between sporadic triple negative tumors with “BRCA1-like” gene expression pattern versus “sporadic” pattern. The final signature of 334 genes is derived from overlap of these three gene lists
This same algorithm was then applied to the Wang and NKI datasets (N = 57, and N = 49 samples, respectively). For each of the datasets the samples were ranked from low score to high score. A sample with a high score had a gene expression profile most similar to BRCA1-associated tumors, and thus was considered to have a high likelihood of having defective DNA repair signature. Three gene lists form each dataset were obtained. They were composed of the most differentially expressed genes between sporadic triple negative tumors with “BRCA1-like” gene expression pattern versus “sporadic” pattern using a false discovery rate of <5%, P < 0.01, 1.5-fold change. The final signature of 334 genes is derived from overlap of these three gene lists, with 136 genes overexpressed in and 198 underexpressed genes (Fig. 1b).
Receiver operating characteristic (ROC) curves were used to assess the accuracy of predictions. The association between expression and pathological complete response was examined by Fisher’s exact test. All statistical tests were two-sided. Sensitivity and specificity were calculated based on the optimal cut-off value as the shortest Euclidean distance obtained from the ROC curves. The Youden index (sensitivity + specificity − 1) was used to select a threshold for estimation of sensitivity and specificity.
Confirmation of expression measurements by single gene Q-RTPCR and by low density QPCR array (LDA)
To confirm measurement of RNA levels, expression values derived from normalized Affymetrix data were correlated with values from semi-quantitative RT-PCR for six genes normalized to 18S. Next, we confirmed measurements of these microarray RNA levels by low density arrays (LDA), based on real time quantitative RT-PCR (QRT-PCR) of 69 most differentially expressed genes.
Confirmation study in neoadjuvant AC patients with 69-gene LDA
The validation neoadjuvant AC study was conducted with the 69-gene LDA was conducted by identifying triple negative patients (n = 28) from the database of 145 patients from the University of Louisville, Kentucky, USA, who had received 6 cycles of standard AC chemotherapy. Pathologic response was assessed by a breast pathologist (SS) without prior knowledge of patient outcome, and pCR was defined as the complete disappearance of all invasive cancer in the breast. The LDA was then applied to RNA extracted form the pretreatment FFPE core biopsies. The AUC, sensitivity, and specificity were then calculated, as above.
Results
We have derived a gene expression profile that is associated with DNA repair deficiency in sporadic TN breast cancers. Van’t Veer et al. published a gene expression signature that can potentially distinguish breast tumors from germline BRCA1 mutation carriers from sporadic tumors. Using this gene signature and the genetic profiles of sporadic TN from three datasets, the overlap yielded a final signature of 334 with 136 genes overexpressed in and 198 underexpressed genes (Fig. 1).
Increased expression of known DNA repair genes in “BRCA1-like” tumors
We selected four known and commonly cited DNA repair genes (PARP-1, RAD51, FANCA, and CHK1) and measured the expression levels of these genes in TN breast cancers. By microarray, all four genes had increased expression in “BRCA1-like” tumors (Fig. 2a). Additionally, we confirmed the expression of PARP-1, RAD51, and CHK1 by single gene QRT-PCR, of which PARP1 and CHEK1 were significantly increased (P < 0.05), while RAD51 showed a trend towards increased expression in “BRCA1-like” tumors (P = 0.056) (Fig. 2b). These data are consistent with up-regulation of known DNA repair genes in these sporadic TN cancers that bear the “BRCA1-like” signature [26].
Increased expression of known DNA repair genes in “BRCA1-like” tumors versus other TN cancers. a By microarray—known DNA repair pathway genes (PARP1, RAD51, FANCA, CHK1) have increased gene expression in tumors identified as having defective DNA repair signature. BCM1, Wang, NKI2 Datasets combined. b By QRT-PCRNA—DNA repair-related genes (PARP1, CHEK1, and RAD51) had higher RNA expression in tumors identified as having defective DNA repair signature. High: tumors with BRCA1-like signature: Low: tumors without BRCA1-like signature. BCM1 Dataset
Confirmation of expression measurements by single gene Q-RTPCR and by low density QPCR array
To confirm measurement of RNA levels, expression values derived from normalized Affymetrix data were correlated with values from semi-quantitative RT-PCR for six genes normalized to 18S. Spearman rank correlations were positive for all 6 genes (SERPINF1, PDGRA, HSP14, EFEMP2, COL15A1, and CDH5), and significantly positive for 5 of 6 genes (P < 0.05).
Next, we confirmed measurements of these microarray RNA levels by the correlation of normalized Affymetrix data versus a 69-gene low density array (LDAs). Low density arrays (LDAs), based on real time quantitative RT-PCR (QRT-PCR), enable a more focused and sensitive approach to the study of gene expression than gene chips, while offering higher throughput than single gene RT-PCR. To compare expression profiles between specimens, normalization based on three reference genes was used. An average of three references genes was used for normalization in a manner previously described [27, 28]. Relative mRNA was expressed as \( 2^{{\Updelta C_{\text{T}} }} \) + 7.1, where ∆C T = C T (test gene) − C T (mean of three reference genes). The average expression of the mean of the three reference genes is 10, corresponding to a C T of 29.6. We confirmed the expression of 69 most differentially expressed genes normalized to ACTB, IPO8, and POLR2A at P < 0.05. The correlation coefficients between the two methods were significantly positive for 45 of 69, 65.2% of the genes (P < 0.05).
Defective DNA repair microarray gene expression signature is associated with anthracycline response and suggests taxane resistance
We hypothesized that those tumors exhibiting the presumptive defective DNA repair pattern would be most sensitive to DNA-damaging drugs, particularly doxorubicin, and would show relative resistance to taxanes. We then confirmed the value of this signature in association with response to neoadjuvant chemotherapy in independent clinical trials.
Consistent with these tumors having defective DNA repair, a higher pathologic response rate (pCR) to anthracycline chemotherapy was observed in those tumors that exhibited the defective DNA repair pattern (Fig. 3). In the first data set, 80 patients were enrolled in a prospective trial at BCM (BCM2 dataset) who were treated with neoadjuvant AC. Evaluating patients (N = 16) with TN breast cancer, a higher pCR or near pCR rate (vs. non-pCR) was observed in patients in patients with high likelihood of defective DNA repair (7/8 vs. 2/8), P = 0.04.
In the second validation data set involving 50 TN patients receiving neoadjuvant FEC chemotherapy and again, a higher pCR to FEC was observed in patients with high likelihood of defective DNA repair. The area under the ordinary receiver operating characteristic (ROC) curve is 0.61, 95% CI = 0.45–0.77 (Fig. 3a), with a sensitivity and specificity of 0.62and 0.62, respectively (Table 1).
Interestingly, this second validation neoadjuvant trial randomized patients to FEC versus a primarily taxane-based regimen, TET. Here, patients (n = 39) received six full cycles of docetaxel, while epirubicin was given for only three cycles at a low dose of 90 mg/m2, which is in total less than half the usually prescribed adjuvant dose. The defective DNA repair signature was associated, conversely, with relative taxane resistance. The area under the ordinary receiver operating characteristic (ROC) curve is 0.65, 95% CI = 0.46–0.85 (Fig. 3b), and the sensitivity and specificity of 0.61 and 0.76, respectively, indicating that this expression pattern was not representative of general chemosensitivity.
The utility of the 69-gene LDA in predicting anthracycline response
Of the 28 TN patients, 25% (7/28) achieved pathologic complete response. From FFPE core biopsies, sufficient RNA was isolated from 21 samples, which were then used to interrogate the 69-gene low density array (LDA). This 69-gene LDA could predict anthracycline response, with an AUC of 0.79 (95% CI = 0.59–0.98), with a sensitivity of 0.86, and a specificity of 0.64 (Fig. 4).
Discussion
There are no currently approved targeted therapies in TN breast cancer patients, who traditionally have a poor prognosis. Patients with chemotherapy-refractory disease after neoadjuvant treatment have a high chance of distant relapse and death [29]. In this study, we have identified a gene expression pattern that identifies patients whose tumors may have defective DNA repair similar to BRCA1-related breast cancer. This expression pattern was confirmed with two other RNA platforms, QRT-PCR and a 69-gene low density array (LDA). This signature was associated with sensitivity to DNA-damaging chemotherapy (anthracyclines) and relative taxane resistance, consistent with published preclinical data in BRCA1-deficient tumors [9, 30–32].
In neoadjuvant chemotherapy studies, pathologic complete response (pCR) is associated with improved patient outcome. Despite TN cancers as a whole having poor prognosis, paradoxically, TN breast cancer patients generally achieve a higher rate of pCR. Additionally, BRCA1 mutation carriers with breast cancer achieve a higher rate of pCR. A plausible explanation is that TN breast cancer is a heterogeneous disease [33–35] with some tumors characterized by defective DNA repair similar to BCA1 tumors, a defect that can be therapeutically exploited as these have an enhanced response to DNA-damaging agents. We sought to recognize this expression pattern in sporadic TN breast cancers that may have a deficiency in DNA repair, and hence, show a differential improved response to agents like anthracyclines, and possibly other DNA-damaging agents.
In a hereditary mouse model of breast cancer where mice spontaneously develop mammary tumors in which BRCA1 protein has been lost, differential responses to chemotherapy (doxorubicin, docetaxel, and cisplatin) have been observed [5–8, 31, 32, 36]. These mice demonstrated resistance to docetaxel, yet were highly sensitive to DNA-damaging drugs like cisplatin and doxorubicin. Additionally, sensitivity to PARP-1 inhibitors has also been shown [37, 38]. PARP-1 is a group of proteins that contribute to the survival of both proliferating and non-proliferating cells following DNA damage. It is involved in the first immediate cellular response to DNA damage, and its activation leads to DNA repair through the base excision repair (BER) pathway. Based on these observations, PARP-1 inhibitors have been reported to have high single agent activity in germline BRCA mutation carriers [39]. These findings have recently been extrapolated to sporadic TN breast cancer patients in combination with chemotherapy in metastatic triple negative patients [40].
Low density arrays (LDAs) have recently been introduced as a novel approach to confirm gene expression profiling results [41]. Based on QRT-PCR, these LDAs can be used on routinely processed, formalin-fixed, paraffin-embedded (FFPE) tissue and represent a valuable approach for sensitive and quantitative gene expression profiling of multiple genes. In this study, we confirmed with the gene expression pattern with small amounts of FFPE tissue. Successful application of these LDAs in breast cancer may assist in the selection of patients who might, or more importantly, might not benefit from anthracycline chemotherapy and other DNA damaging agents like PARP-1 inhibitors, and who might be better treated with taxane-based chemotherapy.
Limitations in this study would include the relatively small patient numbers in these analyses, as triple negative tumors account for only 15% of all breast cancers, thus increasing the difficulty in acquiring large datasets. Nonetheless, we have demonstrated a defective DNA repair signature that is associated with anthracycline response and taxane resistance in TN breast cancer patients. Further prospective validation in separate cohorts is underway.
References
Carey LA, Dees EC, Sawyer L, Gatti L, Moore DT, Collichio F et al (2007) The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res 13(8):2329–2334
von Minckwitz G KM, Kümmel S, Fasching P, Eiermann W, Blohmer J-U, Costa SD, Sibylle L, Dietmar V, Untch M (2008) Integrated meta-analysis on 6402 patients with early breast cancer receiving neoadjuvant anthracycline-taxane ± trastuzumab containing chemotherapy. SABCS Abstract 792008
Sorlie T, Tibshirani R, Parker J, Hastie T, Marron JS, Nobel A et al (2003) Repeated observation of breast tumor subtypes in independent gene expression data sets. Proc Natl Acad Sci USA 100(14):8418–8423
Bernstein C, Bernstein H, Payne CM, Garewal H (2002) DNA repair/pro-apoptotic dual-role proteins in five major DNA repair pathways: fail-safe protection against carcinogenesis. Mutat Res 511(2):145–178
Tassone P, Di Martino MT, Ventura M, Pietragalla A, Cucinotto I, Calimeri T et al (2009) Loss of BRCA1 function increases the antitumor activity of cisplatin against human breast cancer xenografts in vivo. Cancer Biol Ther 8(7):648–653
Murray MM, Mullan PB, Harkin DP (2007) Role played by BRCA1 in transcriptional regulation in response to therapy. Biochem Soc Trans 35(5):1342–1346
Kennedy RD, Quinn JE, Mullan PB, Johnston PG, Harkin DP (2004) The role of BRCA1 in the cellular response to chemotherapy. J Natl Cancer Inst 96(22):1659–1668
Tassone P, Tagliaferri P, Perricelli A, Blotta S, Quaresima B, Martelli ML et al (2003) BRCA1 expression modulates chemosensitivity of BRCA1-defective HCC1937 human breast cancer cells. Br J Cancer 88(8):1285–1291
Delaloge S, Bidard F, El Masmoudi Y, Bressac de Paillerets B, Caron O, Bourgier C, Garbay J, Spielmann M, André F. BRCA1 germ-line mutation: Predictive of sensitivity to anthracyclin alkylating agents regimens but not to taxanes? J Clin Oncol. Abstract 5742008 May 20, 26
Chappuis PO, Goffin J, Wong N, Perret C, Ghadirian P, Tonin PN et al (2002) A significant response to neoadjuvant chemotherapy in BRCA1/2 related breast cancer. J Med Genet 39(8):608–610
Hubert A, Mali B, Hamburger T, Rottenberg Y, Uziely B, Peretz T et al (2009) Response to neo-adjuvant chemotherapy in BRCA1 and BRCA2 related stage III breast cancer. Fam Cancer 8:173–177
Byrski T, Huzarski T, Dent R, Gronwald J, Zuziak D, Cybulski C et al (2009) Response to neoadjuvant therapy with cisplatin in BRCA1-positive breast cancer patients. Breast Cancer Res Treat 115:359–363
Byrski T, Gronwald J, Huzarski T, Grzybowska E, Budryk M, Stawicka M et al (2008) Response to neo-adjuvant chemotherapy in women with BRCA1-positive breast cancers. Breast Cancer Res Treat 108(2):289–296
Turner NC, Reis-Filho JS, Russell AM, Springall RJ, Ryder K, Steele D et al (2007) BRCA1 dysfunction in sporadic basal-like breast cancer. Oncogene 26(14):2126–2132
James CR, Quinn JE, Mullan PB, Johnston PG, Harkin DP (2007) BRCA1, a potential predictive biomarker in the treatment of breast cancer. Oncologist 12(2):142–150
Turner NC, Reis-Filho JS (2006) Basal-like breast cancer and the BRCA1 phenotype. Oncogene 25(43):5846–5853
Brody LC (2005) Treating cancer by targeting a weakness. N Engl J Med 353(9):949–950
Turner N, Tutt A, Ashworth A (2004) Hallmarks of ‘BRCAness’ in sporadic cancers. Nat Rev Cancer 4(10):814–819
Mueller CR, Roskelley CD (2003) Regulation of BRCA1 expression and its relationship to sporadic breast cancer. Breast Cancer Res 5(1):45–52
van de Vijver MJ, He YD, van’ t Veer LJ, Dai H, Hart AA, Voskuil DW et al (2002) A gene-expression signature as a predictor of survival in breast cancer. N Engl J Med 347(25):1999–2009
Mohsin SK, Weiss HL, Gutierrez MC, Chamness GC, Schiff R, Digiovanna MP et al (2005) Neoadjuvant trastuzumab induces apoptosis in primary breast cancers. J Clin Oncol 23(11):2460–2468
Farmer P, Bonnefoi H, Anderle P, Cameron D, Wirapati P, Becette V et al (2009) A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nat Med 15(1):68–74
Bonnefoi H, Potti A, Delorenzi M, Mauriac L, Campone M, Tubiana-Hulin M et al (2007) Validation of gene signatures that predict the response of breast cancer to neoadjuvant chemotherapy: a substudy of the EORTC 10994/BIG 00–01 clinical trial. Lancet Oncol 8(12):1071–1078
Chang JC, Wooten EC, Tsimelzon A, Hilsenbeck SG, Gutierrez MC, Elledge R et al (2003) Gene expression profiling for the prediction of therapeutic response to docetaxel in patients with breast cancer. Lancet 362(9381):362–369
Chang JC, Wooten EC, Tsimelzon A, Hilsenbeck SG, Gutierrez MC, Tham YL et al (2005) Patterns of resistance and incomplete response to docetaxel by gene expression profiling in breast cancer patients. J Clin Oncol 23(6):1169–1177
Martin RW, Orelli BJ, Yamazoe M, Minn AJ, Takeda S, Bishop DK (2007) RAD51 up-regulation bypasses BRCA1 function and is a common feature of BRCA1-deficient breast tumors. Cancer Res 67(20):9658–9665
Cronin M, Pho M, Dutta D, Stephans JC, Shak S, Kiefer MC et al (2004) Measurement of gene expression in archival paraffin-embedded tissues: development and performance of a 92-gene reverse transcriptase-polymerase chain reaction assay. Am J Pathol 164(1):35–42
Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, et al (2002) Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol 3(7):RESEARCH0034
Liedtke C, Mazouni C, Hess KR, Andre F, Tordai A, Mejia JA et al (2008) Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol 26(8):1275–1281
Wysocki PJ, Korski K, Lamperska K, Zaluski J, Mackiewicz A (2008) Primary resistance to docetaxel-based chemotherapy in metastatic breast cancer patients correlates with a high frequency of BRCA1 mutations. Med Sci Monit 14(7):SC7–SC10
Tassone P, Blotta S, Palmieri C, Masciari S, Quaresima B, Montagna M et al (2005) Differential sensitivity of BRCA1-mutated HCC1937 human breast cancer cells to microtubule-interfering agents. Int J Oncol 26(5):1257–1263
Gilmore PM, McCabe N, Quinn JE, Kennedy RD, Gorski JJ, Andrews HN et al (2004) BRCA1 interacts with and is required for paclitaxel-induced activation of mitogen-activated protein kinase kinase kinase 3. Cancer Res 64(12):4148–4154
Teschendorff AE, Miremadi A, Pinder SE, Ellis IO, Caldas C (2007) An immune response gene expression module identifies a good prognosis subtype in estrogen receptor negative breast cancer. Genome Biol 8(8):R157
Kreike B, van Kouwenhove M, Horlings H, Weigelt B, Peterse H, Bartelink H et al (2007) Gene expression profiling and histopathological characterization of triple-negative/basal-like breast carcinomas. Breast Cancer Res 9(5):R65
Schneider BP, Winer EP, Foulkes WD, Garber J, Perou CM, Richardson A et al (2008) Triple-negative breast cancer: risk factors to potential targets. Clin Cancer Res 14(24):8010–8018
Sgagias MK, Wagner KU, Hamik B, Stoeger S, Spieker R, Huber LJ et al (2004) Brca1-deficient murine mammary epithelial cells have increased sensitivity to CDDP and MMS. Cell Cycle 3(11):1451–1456
Ashworth A (2008) A synthetic lethal therapeutic approach: poly(ADP) ribose polymerase inhibitors for the treatment of cancers deficient in DNA double-strand break repair. J Clin Oncol 26(22):3785–3790
Farmer H, McCabe N, Lord CJ, Tutt AN, Johnson DA, Richardson TB et al (2005) Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434(7035):917–921
Fong PC, Boss DS, Yap TA, Tutt A, Wu P, Mergui-Roelvink M et al (2009) Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med 361(2):123–134
O’Shaughnessy J, Osborne C, Pippen J, Yoffe M, Patt D, Monaghan G, Rocha C, Ossovskaya V, Sherman B, Bradley C (2009) Efficacy of BSI-201, a poly (ADP-ribose) polymerase-1 (PARP1) inhibitor, in combination with gemcitabine/carboplatin (G/C) in patients with metastatic triple-negative breast cancer (TNBC): results of a randomized phase II trial. J Clin Oncol. Abstract 32009; 27
Abruzzo LV, Lee KY, Fuller A, Silverman A, Keating MJ, Medeiros LJ et al (2005) Validation of oligonucleotide microarray data using microfluidic low-density arrays: a new statistical method to normalize real-time RT-PCR data. Biotechniques 38(5):785–792
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rodriguez, A.A., Makris, A., Wu, M.F. et al. DNA repair signature is associated with anthracycline response in triple negative breast cancer patients. Breast Cancer Res Treat 123, 189–196 (2010). https://doi.org/10.1007/s10549-010-0983-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-010-0983-z