Abstract
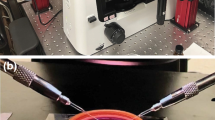
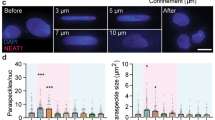
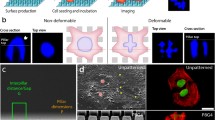
Limited space for cell division, perhaps similar to the compressed microenvironment of a growing tumor, has been shown to induce phenotypic and karyotypic changes to a cell during mitosis. To expand understanding of this missegregation of chromosomes in aberrant multi-daughter or asymmetric cell divisions, we present a simple technique for subjecting mammalian cells to adjustable levels of confinement which allows subsequent interrogation of intracellular molecular components using high resolution confocal imaging. PDMS micropatterned confinement structures of subcellular height with neighboring taller media reservoir channels were secured on top of confluent cells with a custom compression well-plate system. The system improved ease of use over previous devices since confined cells could be initially grown on glass coverslips in a 12-well plate, and subsequently be imaged by high resolution confocal imaging, or during compression by live cell imaging. Live cell imaging showed a significant increase in abnormal divisions of confined cells across three different cell lines (HeLa, A375, and A549). Immunofluoresecence stains revealed a significant increase in cell diameter and chromosome area of confined cells, but no significant increase in centrosome-centromere distance upon division when compared to unconfined cells. The developed system could open up studies more broadly on confinement effects on mitotic processes, and increase the throughput of such studies.






Similar content being viewed by others
References
G.B. Alsop, D. Zhang, J. Cell Biol. 162, 383 (2003)
C. Antonio, I. Ferby, H. Wilhelm, M. Jones, E. Karsenti, A.R. Nebreda, I. Vernos, Cell 102, 425 (2000)
F. Borel, O.D. Lohez, F.B. Lacroix, R.L. Margolis, Proc. Natl. Acad. Sci. U. S. A. 99, 9819 (2002)
D.R. Burgess, F. Chang, Trends Cell Biol. 15, 156 (2005)
K.F. DeLuca, S.M.A. Lens, J.G. DeLuca, J. Cell Sci. 124, 622 (2011)
C. Ditchfield, V.L. Johnson, A. Tighe, R. Ellston, C. Haworth, T. Johnson, A. Mortlock, N. Keen, S.S. Taylor, J. Cell Biol. 161, 267 (2003)
N.J. Ganem, S.A. Godinho, D. Pellman, Nature 460, 278 (2009)
M.C. Good, M.D. Vahey, A. Skandarajah, D.A. Fletcher, R. Heald, Science 342, 856 (2013)
D.J. Gordon, B. Resio, D. Pellman, Nat. Rev. Genet. 13, 189 (2012)
J. Hazel, K. Krutkramelis, P. Mooney, M. Tomschik, K. Gerow, J. Oakey, J.C. Gatlin, Science 342, 853 (2013)
M. Hetzer, O.J. Gruss, I.W. Mattaj, Nat. Cell Biol. 4, E177 (2002)
H.M.J. Hut, W. Lemstra, E.H. Blaauw, G.W.A. Van Cappellen, H.H. Kampinga, O.C.M. Sibon, Mol. Biol. Cell 14, 1993 (2003)
T. Itabashi, Y. Terada, K. Kuwana, T. Kan, I. Shimoyama, Proc. Natl. Acad. Sci. U. S. A. 109, 7320 (2012)
F. Jiang, N.P. Caraway, R. Li, R.L. Katz, Oncogene 24, 3409 (2005)
A. Krämer, B. Maier, J. Bartek, Mol. Oncol. 5, 324 (2011)
O.M. Lancaster, M. Le Berre, A. Dimitracopoulos, D. Bonazzi, E. Zlotek-Zlotkiewicz, R. Picone, T. Duke, M. Piel, B. Baum, Dev. Cell 25, 270 (2013)
M. Le Berre, J. Aubertin, M. Piel, Integr. Biol. 4, 1406 (2012)
C.L. Limoli, M.I. Kaplan, E. Giedzinski, W.F. Morgan, Free Radic. Biol. Med. 31, 10 (2001)
W.L. Lingle, S.L. Barrett, V.C. Negron, A.B. D’Assoro, K. Boeneman, W. Liu, C.M. Whitehead, C. Reynolds, J.L. Salisbury, Proc. Natl. Acad. Sci. U. S. A. 99, 1978 (2002)
D. Liu, G. Vader, M.J.M. Vromans, M.A. Lampson, S.M.A. Lens, Science 323, 1350 (2009)
X. Lu, Y. Kang, Cancer Res. 69, 8536 (2009)
C. Luk, M.S. Tsao, J. Bayani, F. Shepherd, J.A. Squire, Cancer Genet. Cytogenet. 125, 87 (2001)
M. Macville, E. Schröck, H. Padilla-Nash, C. Keck, B.M. Ghadimi, D. Zimonjic, N. Popescu, T. Ried, Cancer Res. 59, 141 (1999)
L. Menendez, B.B. Benigno, J.F. Mcdonald, Mol. Cancer 3, 1 (2004)
M. Nishigaki, K. Aoyagi, I. Danjoh, Cancer Res. 65, 2115 (2005)
G.A. Pihan, J. Wallace, Y. Zhou, P. Carcinomas, Cancer Res. 63, 1398 (2003)
C.L. Rieder, E.D. Salmon, J. Cell Biol. 124, 223 (1994)
I. Sánchez, L. Goya, A.K. Vallerga, G.L. Firestone, Cell Growth Differ. 4, 215 (1993)
M.P. Stewart, J. Helenius, Y. Toyoda, S.P. Ramanathan, D.J. Muller, A.A. Hyman, Nature 469, 226 (2011)
M. Théry, V. Racine, A. Pépin, M. Piel, Y. Chen, J.-B. Sibarita, M. Bornens, Nat. Cell Biol. 7, 947 (2005)
H.T.K. Tse, W.M. Weaver, D. Di Carlo, PLoS One 7, e38986 (2012)
P. Tseng, J.W. Judy, D. Di Carlo, Nat. Methods 9, 1113 (2012)
S.P. Wheatley, Y. Wang, J. Cell Biol. 135, 981 (1996)
A.S. Wilson, B.E. Power, P.L. Molloy, Biochim. Biophys. Acta 1775, 138 (2007)
M. Wühr, Y. Chen, S. Dumont, A.C. Groen, D.J. Needleman, A. Salic, T.J. Mitchison, Curr. Biol. 18, 1256 (2008)
S. Yasuda, T. Oceguera-Yanez, F. Kato, M. Okamoto, Y. Yonemura, S. Terada, T. Ishizaki, N. Shuh, Nature 428, 767 (2004)
Acknowledgments
The authors would like to thank Yeganeh Amini, Coleman Murray, Dr. Matthew Schibler, and Nikhil Kalluri for their assistance in this work. Confocal laser scanning microscopy was performed at the CNSI Advanced Light Microscopy/Spectroscopy Shared Resource Facility at UCLA, supported with funding from NIH-NCRR shared resources grant (CJX1-443835-WS-29646) and NSF Major Research Instrumentation grant (CHE-0722519).
Ethics statement
The manuscript does not contain clinical studies or patient data. The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online Resource 1
(PDF 1.27 mb)
Online Resource 2
(AVI 52 kb)
Online Resource 3
(AVI 28 kb)
Online Resource 4
(AVI 1849 kb)
Online Resource 5
(AVI 522 kb)
Online Resource 6
(PDF 5.35 MB)
Rights and permissions
About this article
Cite this article
Kittur, H., Weaver, W. & Di Carlo, D. Well-plate mechanical confinement platform for studies of mechanical mutagenesis. Biomed Microdevices 16, 439–447 (2014). https://doi.org/10.1007/s10544-014-9846-4
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10544-014-9846-4