Abstract
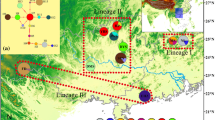
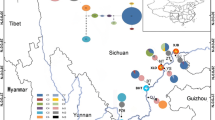
Belonging to the genus Dendrobium of Orchidaceae, Dendrobium moniliforme is an endangered species with disjunct distribution in East Asia, possessing significant medicinal value. To investigate its phylogeography, this study compared sequences of two mitochondrial DNA (mtDNA) fragments (nad 1/b-c and nad 7/2-3) from 143 samples of 18 natural populations of D. moniliforme almost covering the entire range of the Sino-Japanese Floristic Region (SJFR) of East Asia. As a result, a total of 30 mtDNA haplotypes were identified in these populations which revealed high levels of haplotype diversity (H d = 0.8733) and total genetic diversity (H T = 0.8886). Additionally, G ST value being significantly lower (0.451) than N ST value (0.722) (P < 0.05) indicated the presence of strong phylogeographic structure in these populations, and the network of mtDNA haplotypes showed that all haplotypes were divided into two clades (A and B). Haplotype overlap was observed among several D. moniliforme population groups in mainland China, suggesting the occurrence of ongoing and/or historical gene flow among them. No common haplotypes were shared by D. moniliforme populations from mainland China and the CKJ Islands (representing Taiwan [China], South Korea, and Japan collectively), pointing to their allopatric evolution in the two regions. Moreover, mismatch distribution analysis and neutral test of mtDNA data rejected the population expansion model. According to the mtDNA-based results, we infer that multiple refugia for D. moniliforme existed in the SJFR of East Asia during the Quaternary glacial period.




Similar content being viewed by others
References
Arditti J, Ghani AKA (2000a) Numerical and physical properties of orchid seeds and their biological implications (vol 145, pg 367, 2000). New Phytol 146:569
Arditti J, Ghani AKA (2000b) Tansley review No. 110—numerical and physical properties of orchid seeds and their biological implications. New Phytol 145:367–421
Avise JC (2009) Phylogeography: retrospect and prospect. J Biogeogr 36:3–15
Bai WN, Liao WJ, Zhang DY (2010) Nuclear and chloroplast DNA phylogeography reveal two refuge areas with asymmetrical gene flow in a temperate walnut tree from East Asia. New Phytol 188:892–901
Bandelt HJ, Forster P, Sykes BC, Richards MB (1995) Mitochondrial portraits of human populations using median networks. Genetics 141:743–753
Bandelt HJ, Forster P, Rohl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Bao XS, Shun QS, Ye YQ, Gu HF (1999) History and current status of Dendrobium medicinal ‘fengdou‘. Chin Tradit Herb Drugs 22:540–542
Bulpitt CJ, Li Y, Bulpitt PF, Wang J (2007) The use of orchids in Chinese medicine. J R Soc Med 100:558–563
Caicedo AL, Schaal BA (2004) Heterogeneous evolutionary processes affect R gene diversity in natural populations of Solanum pimpinellifolium. Proc Natl Acad Sci USA 101:17444–17449
Caieedo AL, Sehaal BA (2004) Population structure and phylogeography of Solanum Pimpinellifolium inferred from anueleargene. Mol Ecol 13(7):1871–1882
Cameron KM (2009) On the value of nuclear and mitochondrial gene sequences for reconstructing the phylogeny of vanilloid orchids (Vanilloideae, Orchidaceae). Ann Bot 104:377–385
Demesure B, Sodzi N, Petit RJ (1995) A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol Ecol 4:129–131
Deng KJ, Yang ZJ, Liu C, Zhao W, Liu C, Feng J, Ren ZL (2007) Identification and phylogenetic application of unique nucleotide sequence of nad 7 intron2 in Rhodiola (Crassulaceae) species. Hereditas 29:371–375
Ding G, Zhang DZ, Feng ZY, Fan WJ, Ding XY, Li XX (2008) SNP, ARMS and SSH authentication of medicinal Dendrobium officinale KIMURA et MIGo and application for identification of Fengdou drugs. Biol Pharm Bull 31:553–557
Duminil J, Pemonge MH, Petit RJ (2002) A set of 35 consensus primer pairs amplifying genes and introns of plant mitochondrial DNA. Mol Ecol Notes 2:428–430
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2005) Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Geng LX, Zheng R, Ren J, Niu ZT, Sun YL, Xue QY, Liu W, Ding XY (2015) Applylication of new type combined fragments: nrDNA ITS+ nad 1-intron 2 for identification of Dendrobium species of Fengdous. Acta Pharm Sin 50:1060–1067
Gu RQ, Chen Z, Li J, Yang XL, Li S, Zhu YG, Zhang WJ, Chen JK (2010) Identification of four medicines of Panax L. revealed by mitochondrial nad 1 gene b/c intron. J Chin Med Mater 33:45–48
Guo YL, Ge S (2004) The utility of mitochondrial nad 1 intron in phylogenetic study of Oryzeae with reference to the systematic position of Porteresia. Acta Phytotaxon Sin 42:333–344
Harpending HC (1994) Signature of ancient population growth in a low-resolution mitochondrial DNA mismatch distribution. Hum Biol 66:591–600
Holder K, Montgomerie R, Friesen VL (1999) A test of the glacial refugium hypothesis using patterns of mitochondrial and nuclear DNA sequence variation in Rock Ptarmigan (Lagopus mutus). Evolution 53:1936–1950
Hu SY (1970) Dendrobium in Chinese medicine. Econ Bot 24:165–170
Kadyrova GD, Ryzhovaa NN, Kochievaa EZ (2010) Phylogenetic relationships among Fagopyrum species based on analysis of the nad1 gene b/c intron. Mosc Univ Biol Sci Bull 65:161–163
Li EX, Yi S, Qiu YX, Guo JT, Comes HP, Fu CX (2008a) Phylogeography of two East Asian species in Croomia (Stemonaceae) inferred from chloroplast DNA and ISSR fingerprinting variation. Mol Phylogenet Evol 49:702–714
Li X, Ding X, Chu B, Zhou Q, Ding G, Gu S (2008b) Genetic diversity analysis and conservation of the endangered Chinese endemic herb Dendrobium officinale Kimura et Migo (Orchidaceae) based on AFLP. Genetica 133:159–166
Librado P, Rozas J (2009) Dnasp v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Nei M, Takezaki N. (1983) Estimation of genetic distances and phylogenetic trees from DNA analysis. Proceeding of 5th World Congress on Genetics Applied to Livestock Production 21:405–412
Palmer JD (1992) Mitochondrial DNA in plant systematics: applications and limitations. In: Soltis PS, Soltis DE, Doyle JJ (eds) Molecular systematics of plants. Chapman and Hall, New York, pp 36–49
Palmer JD, Herbon LA (1988) Plant mitochondrial DNA evolves rapidly in structure, but slowly in sequence. J Mol Evol 28:87–97
Petit RJ, Brewer S, Bordács S (2002) Identification of refugia and post-glacial colonisation routes of European white oaks based on chloroplast DNA and fossil pollen evidence. For Ecol Manag 156:49–74
Pfeifer M, Schatz B, Pico FX (2009) Phylogeography and genetic structure of the orchid Himantoglossum hircinum (L.) Spreng. across its European central-marginal gradient. J Biogeogr 36:2353–2365
Pons O, Petit RJ (1996) Measuring and testing genetic differentiation with ordered versus unordered alleles. Genetics 144:1237–1245
Qi XS, Chen C, Comes HP, Sakaguchi S, Liu YH, Tanaka N, Sakio H, Qiu YX (2012) Molecular data and ecological niche modelling reveal a highly dynamic evolutionary history of the East Asian Tertiary relict Cercidiphyllum (Cercidiphyllaceae). New Phytol 196:617–630
Qiu YX, Qi XS, Jin XF, Tao XY, Fu CX, Naiki A, Comes HP (2009a) Population genetic structure, phylogeography, and demographic history of Platycrater arguta (Hydrangeaceae) endemic to East China and South Japan, inferred from chloroplast DNA sequence variation. Taxon 58:1226–1241
Qiu YX, Sun Y, Zhang XP, Lee J, Fu CX, Comes HP (2009b) Molecular phylogeography of East Asian Kirengeshoma (Hydrangeaceae) in relation to quaternary climate change and landbridge configurations. New Phytol 183:480–495
Qiu YX, Fu CX, Comes HP (2011) Plant molecular phylogeography in China and adjacent regions: Tracing the genetic imprints of Quaternary climate and environmental change in the world’s most diverse temperate flora. Mol Phylogenet Evol 59:225–244
Rogers AR (1995) Genetic evidence for a Pleistocene population explosion. Evolution 49:608–615
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Sakaguchi S, Qiu YX, Liu YH, Qi XS, Kim SH, Han J, Takeuchi Y, Worth JR, Yamasaki M, Sakurai S, Isagi Y (2012) Climate oscillation during the Quaternary associated with landscape heterogeneity promoted allopatric lineage divergence of a temperate tree Kalopanax septemlobus (Araliaceae) in East Asia. Mol Ecol 21:3823–3838
Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S, Kumar S (2011) MEGA 5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tan GX, Tian ML, Huang YB, Rong TZ (2007) The systematic position of waxy corn and utility of mitochondrial nad 1 intron in the phylogenetic study of the Gemus Zea. J Sichuan Agric Univ 25:39–43
Tsi ZH (1999) Flora of China (Chinese Edition). Science Press, Beijing
Wang L, Abbott RJ, Zheng W, Chen P, Wang Y, Liu J (2009) History and evolution of alpine plants endemic to the Qinghai-Tibetan Plateau: Aconitum gymnandrum (Ranunculaceae). Mol Ecol 18:709–721
Wang JF, Gong X, Chiang YC, Kuroda C (2013) Phylogenetic patterns and disjunct distribution in Ligularia hodgsonii Hook. (Asteraceae). J Biogeogr 40:1741–1754
Xiang XG, Hu H, Wang W, Jin XH (2011) DNA barcoding of the recently evolved genus Holcoglossum (Orchidaceae: Aeridinae): a test of DNA barcode candidates. Mol Ecol Resour 11:1012–1021
Xie GW (1997) Phytogeographical affinities of forest floras between China and Japan. J For Res 8:87–90
Yamane K, Yano K, Kawahara T (2006) Pattern and rate of indel evolution inferred from whole chloroplast intergenic regions in sugarcane, maize and rice. DNA Res 13:197–204
Yang R, Meng LH, Zhang X, Liu JQ (2005) The mitochondrial DNA nad1 sequence variation of Picea crassifolia (Pinaceae) in the Qinghai-Tibetan Plateau platform and adjacent populations. Acta Ecol Sin 25:3307–3313
Ye MR, Hou BW, Luo J, Yan WJ, Liu W, Ding XY (2015) Genetic diversity and conservation of the endangered herb dendrobium moniliforme based on amplified fragment length polymorphism markers. Sci Hortic-Amsterda 189:51–58
Zhang T, Wang ZT, Xu LS, Zhou KY (2005) Application of mitochondrial nad 1 intron 2 sequences to molecular identification of some species of Dendrobium Sw. Chin Tradit Herb Drugs 36:1059–1062
Zhang QQ, Liu SJ, Fang CW, He XL (2011) Analysis of chemical constituents of essential oil from flowers of Dendrobium moniliforme (L.) Sw. by GC-MS. Modern Chin Med 13:34–36
Acknowledgements
We are grateful to Ms. Chun Gu for revising the manuscript, Dr. Joongku Lee and Dr. Kamezo Saito for providing samples from South Korea and Japan, respectively. This work was financed by the National Natural Science Foundation of China (No. 31170300) and also by the Priority Academic Program Development of Jiangsu Higher Education Institutions. This work was also supported by the Stable Talent Fundation of Anhui Science and Technology University (No. 830130).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Daniel Sanchez Mata.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ye, M., Liu, W., Xue, Q. et al. Phylogeography of endangered Dendrobium moniliforme in East Asia based on mitochondrial DNA sequence variations. Biodivers Conserv 26, 1659–1674 (2017). https://doi.org/10.1007/s10531-017-1324-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10531-017-1324-x