Abstract
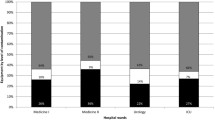
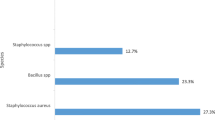
Hospital environmental conditions, human occupancy, and the characteristics of the equipment influence the survival of microbial communities and raise a concern with regard to nosocomial infections. The objective of the present work was to use the monitoring of Pseudomonas aeruginosa, Klebsiella spp. and non-tuberculous mycobacteria as a strategy to improve knowledge on microbial colonization of non-critical equipment and surfaces, in a tertiary hospital from Central Portugal. A 3-month microbiological survey was performed in a district teaching hospital. A total of 173 samples were obtained from the wards Hematology, Urology, Medicine, and Renal Transplants, and 102 presumptive strains recovered. Per sampling, Pseudomonas Isolation agar showed 42.8 to 73.3% of presumptive P. aeruginosa colonies and MacConkey agar recovered mostly Staphylococcus. Most of the colonies recovered in Middlebrook 7H10-PANTA belonged to the genus Methylobacterium. Taps and WC shower curtains carry high bacterial species diversity. The Redundancy Analysis grouped the samples in those mostly handled by patients, and those mostly handled by healthcare staff or of mixed use. This study shows that the preferential users of the space and equipment seem to be important contributors to the microbial community. The most recovered genus was Methylobacterium, known as colonizer of the water distribution system therefore, it is possible that the water points and biofilms in taps also contribute as dispersion hotspots.



Similar content being viewed by others
References
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W et al (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Angenent LT, Kelley ST, St Amand A, Pace NR, Hernandez MT (2005) Molecular identification of potential pathogens in water and air of a hospital therapy pool. Proc Natl Acad Sci USA 102:4860–4865. doi:10.1073/pnas.0501235102
Anzai Y, Kim H, Park JY, Wakabayashi H, Oyaizu H (2000) Phylogenetic affiliation of the pseudomonads based on 16S rRNA sequence. Int J Syst Evol Microbiol 50:1563–1589. doi:10.1099/00207713-50-4-1563
Bessonneau V, Mosqueron L, Berrubé A, Mukensturm G, Buffet-Bataillon S, Gangneux JP et al (2013) VOC contamination in hospital, from stationary sampling of a large panel of compounds, in view of healthcare workers and patients exposure assessment. PLoS ONE 8:e55535. doi:10.1371/journal.pone.0055535
Cateau E, Maisonneuve E, Peguilhan S, Quellard N, Hechard Y, Rodier MH (2014) Stenotrophomonas maltophilia and Vermamoeba vermiformis relationships: bacterial multiplication and protection in amoebal-derived structures. Res Microbiol 165:847–851. doi:10.1016/j.resmic.2014.10.004
Covert TC, Rodgers MR, Reyes AL, Stelma GN. (1999) Occurrence of nontuberculous mycobacteria in environmental samples. Appl Environ Microbiol 65:2492–2496. http://www.ncbi.nlm.nih.gov/pubmed/10347032
de Abreu PM, Farias PG, Paiva GS, Almeida AM, Morais PV (2014) Persistence of microbial communities including Pseudomonas aeruginosa in a hospital environment: a potential health hazard. BMC Microbiol 14:118. doi:10.1186/1471-2180-14-118
Falkinham J, Hilborn E, Arduino M, Pruden A, Edwards MA (2015) Epidemiology and ecology of opportunistic premise plumbing pathogens: Legionella pneumophila, Mycobacterium avium, and Pseudomonas aeruginosa. Environ Health Perspect 110:A174. doi:10.1289/ehp.1409385
Fernandez-Rendon E, Cerna-Cortes JF, Ramirez-Medina M, Helguera-Repetto C, Rivera-Gutierrez S, Estrada-Garcia T et al (2012) Mycobacterium mucogenicum and other non-tuberculous mycobacteria in potable water of a trauma hospital: a potential source for human infection. J Hosp Infect 80:74–76. doi:10.1016/j.jhin.2011.10.003
Høiby N (2011) Recent advances in the treatment of Pseudomonas aeruginosa infections in cystic fibrosis. BMC Med 9:32. doi:10.1186/1741-7015-9-32
Jukes TH, Cantor CR (1990) Evolution of protein molecules, vol 33. Academic Press, New York, pp 21–132
Kovaleva J, Degener JE, van der Mei HC (2014) Methylobacterium and its role in health care-associated infection. J Clin Microbiol 52:1317–1321. doi:10.1128/JCM.03561-13
Lai CC, Cheng A, Liu WL, Tan CK, Huang YT, Chung KP et al (2011) Infections caused by unusual Methylobacterium species. J Clin Microbiol 49:3329–3331. doi:10.1128/JCM.01241-11
Leverstein-Van Hall M, Blok HEM, Paauw A, Fluit AC, Troelstra A, Mascini EM et al (2006) Extensive hospital-wide spread of a multidrug-resistant Enterobacter cloacae clone, with late detection due to a variable antibiogram and frequent patient transfer. J Clin Microbiol 44:518–524. doi:10.1128/JCM.44.2.518-524.2006
Neely AN (2000) A survey of gram-negative bacteria survival on hospital fabrics and plastics. J Burn Care Res 21:523–527
Nielsen P, Fritze D, Priest FG (1995) Phenetic diversity of alkaliphilic Bacillus strains: proposal for nine new species. Microbiology 141:1745–1761. doi:10.1099/13500872-141-7-1745
Panagiotou M, Papaioannou AI, Kostikas K, Paraskeua M, Velentza E, Kanellopoulou M et al (2014) The epidemiology of pulmonary nontuberculous mycobacteria: data from a general hospital in Athens, Greece, 2007–2013. Pulm Med 2014:1–9. doi:10.1155/2014/894976
Pendleton JN, Gorman SP, Gilmore BF (2013) Clinical relevance of the ESKAPE pathogens. Expert Rev Anti Infect Ther 11:297–308. doi:10.1586/eri.13.12
Pitcher DG, Saunders N, Owen RJ (1989) Rapid extraction of bacterial genomic DNA with guanidium thiocyanate. Lett Appl Microbiol 8:151–156. doi:10.1111/j.1472-765X.1989.tb00262.x
Radomski N, Cambau E, Moulin L, Haenn S, Moilleron R, Lucas FS (2010) Comparison of culture methods for isolation of nontuberculous mycobacteria from surface waters. Appl Environ Microbiol 76:3514–3520. doi:10.1128/AEM.02659-09
Rainey FA, Ward-Rainey N, Kroppenstedt RM, Stackebrandt E (1996) The genus Nocardiopsis represents a phylogenetically coherent taxon and a distinct actinomycete lineage: proposal of Nocardiopsaceae fam. nov. Int J Syst Bacteriol 46:1088–1092. doi:10.1099/00207713-46-4-1088
Ramos T, Dedesko S, Siegel J, Gilbert J, Stephens B (2015) Spatial and temporal variations in indoor environmental conditions, human occupancy, and operational characteristics in a new hospital building. PLoS ONE 10:e0118207. doi:10.1371/journal.pone.0118207
Richardson ET, Morrow CD, Kalil DB, Bekker LG, Wood R (2014) Shared air: a renewed focus on ventilation for the prevention of tuberculosis transmission. PLoS ONE 9:e96334. doi:10.1371/journal.pone.0096334
Schau HP (1986) I. Gylstorff (Herausgeber), Infektionen durchMycoplasmatales. 568 S., 39 Abb., 72 Tab. Jena 1985. VEB Gustav Fischer Verlag. M 260,00. J Basic Microbiol 26:224. doi:10.1002/jobm.3620260410
Struve C, Krogfelt KA (2004) Pathogenic potential of environmental Klebsiella pneumoniae isolates. Environ Microbiol 6:584–590. doi:10.1111/j.1462-
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599. doi:10.1093/molbev/msm092
Tan TY, Tan JSM, Tay H, Chua GH, Yong Ng LS, Syahidah N (2013) Multidrug-resistant organisms in a routine ward environment: differential propensity for environmental dissemination and implications for infection control. J Med Microbiol 62:766–772. doi:10.1099/jmm.0.052860-0
Tang JW (2009) The effect of environmental parameters on the survival of airborne infectious agents. J R Soc Interface 6:S737–S746. doi:10.1098/rsif.2009.0227.focus
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Umscheid C, Mitchell MD, Doshi J, Agarwal R, Williams K, Brennan PJ (2011) Estimating the proportion of healthcare-associated infections that are reasonably preventable and the related mortality and costs. Infect Control Hosp Epidemiol 32:101–114. doi:10.1086/657912
Van den Mooter M, Swings J (1990) Numerical analysis of 295 phenotypic features of 266 Xanthomonas strains and related strains and an improved taxonomy of the genus. Int J Syst Bacteriol 40:348–369. doi:10.1099/00207713-40-4-348
Wiener-Well Y, Galuty M, Rudensky B, Schlesinger Y, Attias D, Yinnon AM (2011) Nursing and physician attire as possible source of nosocomial infections. Am J Infect Control 39:555–559. doi:10.1016/j.ajic.2010.12.016
Acknowledgements
This work was financial supported by Instituto Piaget, Portugal, under the project “Estudo da prevalência e da variabilidade genética de estirpes de Pseudomonas aeruginosa e de Mycobacterium sp. em ambiente hospitalar” and by Fundação para a Ciência e Tecnologia, Portugal under the project Projecto PTDC/BIA-MIC/114958/2009- “ArsTOOL-Construção de um bioacumulador para biorremediação de arsénio” Pedro Farias was supported by an Instituto Piaget fellowship. Susana Alarico was supported by FCT fellowship SFRH/BPD/43321/2088. We thank the directing office and staff of the Hematology, Medicine A, and Urology/Renal Transplants wings as well as Dr. Filomena Coelho from the Centre for Infection Control Office.
Author’s Contributions
Conception of the project and responsible for obtaining funding support PVM, AFA, NE. Bacterial work performed by PF, FG, CM, PVM, mycobacterium work performed by DR, SA, FG, NE. First draft produced by PVM, PF, writing and discussing of the manuscript by PF, PVM, SA, NE. All authors agree to act as guarantors of the work, ensuring that questions related to any part of the work are appropriately investigated and resolved.
Conflicts of interest
All authors report no conflicts of interest relevant to this article.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Geadas Farias, P., Gama, F., Reis, D. et al. Hospital microbial surface colonization revealed during monitoring of Klebsiella spp., Pseudomonas aeruginosa, and non-tuberculous mycobacteria. Antonie van Leeuwenhoek 110, 863–876 (2017). https://doi.org/10.1007/s10482-017-0857-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-017-0857-z