Abstract
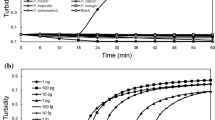
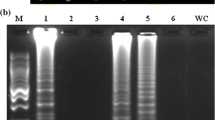
To improve the diagnosis of Stewart’s wilt, we developed a loop-mediated isothermal amplification (LAMP) assay, based on two conserved sequences of the cpsD and pstS–glmS gene regions. The detection limit of the LAMP assay was 104 colony-forming units/mL. No cross reaction was observed with other Pantoea spp., other genera associated with sweet corn diseases, or strains isolated from the surface of maize leaves in Japan. The LAMP reaction was not inhibited by leaf contents. The simplicity of sample preparation and short processing time make this LAMP assay useful for field surveillance of Stewart’s wilt.


Similar content being viewed by others
References
Coplin DL, Majerczak DR (1990) Extracellular polysaccharide genes in Erwinia stewartii: directed mutagenesis and complementation analysis. Mol Plant-Microbe Interact 3:286–292
Coplin DL, Majerczak DR, Zhang Y, Kim WS, Jock S, Geider K (2002) Identification of Pantoea stewartii subsp. stewartii by PCR and strain differentiation by PFGE. Plant Dis 86:304–311
Gehring I, Wensing A, Gernold M, Wiedemann W, Coplin DL, Geider K (2014) Molecular differentiation of Pantoea stewartii subsp. indologenes from subspecies stewartii and identification of new isolates from maize seeds. J Appl Microbiol 116:1553–1562
Harper SJ, Ward LI, Clover GRG (2010) Development of LAMP and real-time PCR methods for the rapid detection of Xylella fastidiosa for quarantine and field applications. Phytopathology 100:1282–1288
IPPC (International Plant Protection Convention) (2002) International Standards for Phytosanitary Measures: Diagnostic protocols for regulated pests. Pub No 27. Food and Agriculture Organization of the United Nations, Rome. http://www.ippc.int/sites/default/files/documents/20140428/ispm_27_2006_en_2014-04-28_201404281447–141.32%20KB.pdf. Cited 5 Nov 2014
Kaneko H, Kawana T, Fukushima E, Suzutani T (2007) Tolerance of loop-mediated isothermal amplification to a culture medium and biological substances. J Biochem Biophys Methods 70:499–501
Kubota R, Vine BG, Alvarez AM, Jenkins DM (2008) Detection of Ralstonia solanacearum by loop-mediated isothermal amplification. Phytopathology 98:1045–1051
Lamka GL, Hill JH, McGee DC, Braun EJ (1991) Development of an immunosorbent assay for seedborne Erwinia stewartii in corn seeds. Phytopathology 81:839–846
MAFF Plant Protection Station (2014) Plants and plant products subject to phytosanitary inspection at growing site. Internet Resource: http://www.pps.go.jp/english/faq/import/seeds.html. Cited 19 Sep 2014
Mergaert J, Verdonck L, Kersters K (1993) Transfer of Erwinia ananas (synonym, Erwinia uredovora) and Erwinia stewartii to the genus Pantoea emend. as Pantoea ananas (Serrano 1928) comb. nov. and Pantoea stewartii (Smith 1898) comb, nov., respectively, and description of Pantoea stewartii subsp. indologenes subsp. nov. Int J Syst Evol Microbiol 43:162–173
Nagamine K, Hase T, Notomi T (2002) Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol Cell Probes 16:223–229
Notomi T, Okayama H, Masubuchi H, Yonekawa T, Watanabe K, Amino N, Hase T (2000) Loop-mediated isothermal amplification of DNA. Nucleic Acids Res 28:E63
Okuda M, Matsumoto M, Tanaka Y, Subandiyah S, Iwanami T (2005) Characterization of the tufB-secE-nusG-rplKAJL-rpoB gene cluster of the citrus greening organism and detection by loop-mediated isothermal amplification. Plant Dis 89:705–711
Okuda M, Kawano S, Murayama Y, Iwanami T (2008) Conditions for loop-mediated isothermal amplification (LAMP) and a nonmacerating DNA extraction method to assay for huanglongbing (citrus greening) disease (in Japanese with English summary). Jpn J Phytopathol 74:316–320
Oya H, Nakagawa H, Saito N, Uematsu H, Ohara T (2008) Detection of Acidovorax avenae subsp. citrulli from seed using LAMP method (in Japanese with English summary). Jpn J Phytopathol 74:304–310
Pataky J, Ikin R (2003) Pest risk analysis: the risk of introducing Erwinia stewartii in maize seed. International Seed Federation, Geneva. http://www.worldseed.org/cms/medias/file/TradeIssues/PhytosanitaryMatters/PestRiskAnalysis/Erwinia_stewartii.pdf. Cited 5 Nov 2014
Tambong JT, Mwange KN, Bergeron M, Ding T, Mandy F, Reid LM, Zhu X (2008) Rapid detection and identification of the bacterium Pantoea stewartii in maize by TaqMan real-time PCR assay targeting the cpsD gene. J Appl Microbiol 104:1525–1537
Wensing A, Zimmermann S, Geider K (2010) Identification of the corn pathogen Pantoea stewartii by mass spectrometry of whole-cell extracts and its detection with novel PCR primers. Appl Environ Microbiol 76:6248–6256
Xu R, Chen Q, Djama ZR, Tambong JT (2010) Miniprimer PCR assay targeting multiple genes: a new rapid and reliable tool for genotyping Pantoea stewartii subsp. stewartii. Lett Appl Microbiol 50:216–222
Acknowledgments
We thank the National Institute of Agrobiological Sciences for providing the bacterial strains.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Uematsu, H., Inoue, Y. & Ohto, Y. Detection of Pantoea stewartii from sweet corn leaves by loop-mediated isothermal amplification (LAMP). J Gen Plant Pathol 81, 173–179 (2015). https://doi.org/10.1007/s10327-015-0580-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10327-015-0580-4