Abstract
In computer-aided diagnosis (CAD) of mediolateral oblique (MLO) view of mammogram, the accuracy of tissue segmentation highly depends on the exclusion of pectoral muscle. Robust methods for such exclusions are essential as the normal presence of pectoral muscle can bias the decision of CAD. In this paper, a novel texture gradient-based approach for automatic segmentation of pectoral muscle is proposed. The pectoral edge is initially approximated to a straight line by applying Hough transform on Probable Texture Gradient (PTG) map of the mammogram followed by block averaging with the aid of approximated line. Furthermore, a smooth pectoral muscle curve is achieved with proposed Euclidean Distance Regression (EDR) technique and polynomial modeling. The algorithm is robust to texture and overlapping fibro glandular tissues. The method is validated with 340 MLO views from three databases—including 200 randomly selected scanned film images from miniMIAS, 100 computed radiography images and 40 full-field digital mammogram images. Qualitatively, 96.75 % of the pectoral muscles are segmented with an acceptable pectoral score index. The proposed method not only outperforms state-of-the-art approaches but also accurately quantifies the pectoral edge. Thus, its high accuracy and relatively quick processing time clearly justify its suitability for CAD.
Similar content being viewed by others
Introduction
In CAD system for mammography, three anatomical landmarks, i.e., pectoral muscle, breast border, and nipple have to be first extracted automatically before the mammogram is to be analyzed for cancer detection [1]. In a high-quality mammogram, the pectoral muscle should be seen at the level of the nipple or below [2]. Consequently, the pectoral muscle always appears as a high-intensity triangular region in the upper portion of the mediolateral oblique (MLO) view of mammogram.
Accurate segmentation of the pectoral muscle in MLO view is very useful in many areas of automated mammographic analysis. In an automated breast tissue density quantification method, the pectoral muscle must be excluded completely. If this step is not followed, fatty tissues will cause an overestimation of the breast density [3]. On the other hand, if the triangular pectoral region inadvertently includes some of the dense glandular area adjacent to the pectoral region, the mammogram density may be underestimated. Moreover, as mentioned by Gupta et al. [4], due to the similarity between pectoral muscles and the mammographic glandular parenchyma, a significantly high number of false positives can occur in mass detection procedures.
The importance of pectoral muscle segmentation makes it a great source of encouragement for researchers working on CAD. In the recent past, several works were proposed on automatic segmentation of pectoral muscle. Karssemeijer et al. [5] applied Hough transform and thresholding method to detect the pectoral muscle with an accuracy of 67 %. Aylward et al. [6] first approximated the pectoral muscle edge by a straight line. This was later refined using a voting scheme. Ferrari et al. [7] implemented and modified the Hough transform, and segmented 21.42 % of pectoral muscle with false positive (FP) and false negative (FN) <10 %. Kinoshita et al. [8] applied Radon transformation to segment muscle with an accuracy of 69.6 % with FP and FN <15 %. However, all these methods consider the muscle to be straight and exhibit a lack in considering its actual curved nature. The obvious outcome of this lacuna is the moderate accuracy of these methods.
Further, Ferrari et al. [9] implemented an efficient detection algorithm based on Gabor wavelet to obtain a smooth pectoral edge with an accuracy of 79.76 % for FP and FN <10 %. However, this method was very time-consuming. Kwok et al. [10] used an iterative threshold method to detect the pectoral muscle approximately and then applied an iterative cliff algorithm with an accuracy of 83.9 % with no quantitative evaluation. Also, Kwok’s algorithm cannot segment highly dense and textured muscle. Raba et al. [11] combined an adaptive histogram approach and a selective region growing method to obtain the pectoral muscle edge. However, in mammograms where the fibro glandular tissues appear near the pectoral muscle, this method may result in over-segmentation. Ma et al. [12] applied two graph-based detection methods to segment the muscle where 80 % images had average error less than 2 mm. Zhou et al. [13] realized that the texture-field orientation (TFO) method performs quantitative assessment with high accuracy but it shows a high false-positive region near textured muscle in mammograms of type BI-RADS-4. This region is important for the analysis of lymph node detection, density estimation, etc. Chakraborty et al. [14] applied an average gradient and shape-based feature method to delineate muscle with an accuracy of 71 % for FP and FN < 10 %. However, this method failed to detect small and low contrast pectoral muscle and, thus, left room for improvement in accuracy.
The aim of the proposed work is to develop a robust technique for the detection of pectoral muscle on MLO view which must consider curvature as well as the overlapping of muscle with dense tissues and variable textures accurately for any shape and size. The technique should be applicable to any kind of mammographic image. In the proposed method, Hough transform is used to obtain straight line approximation on a probable texture gradient map and Euclidean distance regression is applied to refine the detection results. The final segmented pectoral muscle region is assessed by two radiologists and compared with two state-of-the-art methods for pectoral muscle detection. A total of 340 MLO mammograms from screen film, digitized, and digital databases are used in this study.
Materials
The methodology presented in this work is applied on three types of database:
-
1.
Scanned film images
The images from miniMIAS database [15] originate from a film-screen mammographic imaging process in the United Kingdom National Breast Screening Program. These images have a spatial resolution of 1,024 × 1,024, pixel resolution of 200 μm and a gray value resolution of 8 bit. The experiment is conducted on 200 MLO mammograms from miniMIAS consisting of simple, curved, and complex muscles.
-
2.
Computed radiography images (CR):
The proposed algorithm is applied to 100 MLO mammograms collected from Central India Cancer Research Institute (CICRI) Hospital, Nagpur, India. The images have been digitized using scanner GE Senographe 600T. These images have a spatial resolution of 2,370 × 1,770, pixel resolution of 100 μm and a gray value resolution of 12 bit. All images are down sampled by a factor of 2, without loss of significant information to 1,185 × 885 pixels, with an effective resolution of 200-μm pixel and held at 8-bit gray level scale images in order to simplify calculations.
-
3.
Full-field digital mammograms images (FFDM):
The database consists of 40 digital MLO mammograms acquired with Hologic Selenia Dimensions Digital Tomosynthesis Mammography unit at Star Imaging and Research Centre, Pune, India. These images have a spatial resolution of 3,328 × 4,096, pixel resolution of 70 μm, and a gray value resolution of 12 bit. The images were down sampled by a factor of 4 which transforms the size to 832 × 1,024 and pixel resolution to 280 μm.
Method
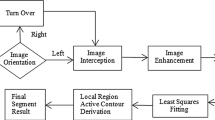
The flow of the proposed method for pectoral muscle detection for CAD is as shown in Fig. 1. The preprocessing steps are: median filtering, artifacts removal like labels, tags, etc., orientation correction, and contrast enhancement. Using morphological operation and connected component algorithm, artifacts were removed. As in mammography, there is no orientation standard and known orientation makes detection of muscle somewhat easier than unknown position [16], so muscles are correctly oriented with the help of pattern-based Radon transform [17]. As a result, each mammogram has chest wall on the left side and pectoral muscle in the upper left corner as shown in Fig. 2. The breast background is also cropped in order to reduce processing time and image file size without losing anatomical information. Furthermore, to improve image contrast and accuracy of muscle segmentation, enhancement [18, 19] is done.
Probable Texture Gradient Map and Straight Line Approximation
Shannon entropy [19] is used to characterize the texture of muscle. Because of the sharp intensity variation of the pectoral edge, the muscle edge pixels indicate high entropy. Instead of calculating global entropy, entropy in the 9 × 9-pixel neighborhood is calculated to extract the texture information locally. Furthermore, in properly oriented mammograms the pectoral muscle edge is oriented in the first quadrant and in the direction perpendicular to the gradient at each pixel location. With the help of the two above-mentioned features and intensity, a probable texture gradient (PTG) map is plotted as the collection of all pixels that have entropy above the threshold entropy, orientation between 30° and 80°, and intensities greater than 50 % of the maximum intensity.
The Hough Transform [19] can detect straight lines in an image. However, as dense tissues overlap the pectoral edge, accurate straight line approximation also becomes challenging. Given this, instead of applying the Hough transform to an intensity map, it is applied to a PTG map to obtain accurate straight approximated muscle. Out of n pixels from a PTG map, the Hough transform finds sets of pixels from the accumulator cells that form the strongest straight line as the approximated pectoral edge. Let the coordinates of the straight pectoral edge be P (P w, 1) and Q (1, P h), where P w is the width and P h is the height of the pectoral muscle as shown in Fig. 2.
Steps for straight line approximation of the muscle using Hough transform on PTG map are as follows:
-
1.
Consider oriented mammogram I(x,y) (Fig. 3a).
-
2.
Improve contrast of the oriented mammogram (Fig. 3b).
-
3.
Calculate entropy H(x,y) in the 9 × 9-pixel neighborhood.
-
4.
Apply Sobel operator to obtain gradient orientation angle α (x, y).
-
5.
Obtain probable texture gradient (PTG) map of the mammogram.
$$ \mathrm{P}\mathrm{T}\mathrm{G}\left(x,y\right)=\left\{1\Big|H\left(x,y\right)\ge {H}_T,30{}^{\circ}\le \beta \left(x,y\right)\le 80{}^{\circ},I\left(x,y\right)\ge 0.5\;{I}_{\max}\right\} $$(1)where H T , the entropy threshold is equal to H mean ; the mean value of the local entropy for miniMIAS images or is equal to H UQ ; upper quartile of the local entropy for CR and FFDM images. I max, the maximum intensity of the image. β, the pectoral edge orientation perpendicular to gradient given as β(x, y) = α(x, y) − 90° where α is the angle of the gradient (Fig. 3c).
-
6.
Consider the initial Region of interest (ROI) from PTG for straight line estimation.
$$ ROI=\left\{\left(x,y\right):0\le x\le M\&0\le y\le 3N\kern0.10em /\kern0.10em 4\right\} $$(2)where M and N are width and height of the mammogram, respectively (Fig. 3d).
-
7.
Apply Hough transform to obtain strongest straight line as approximated muscle edge (white line) (Fig. 3d, e).
-
8.
If the coordinates of the end point of the strongest line contain first row and first column, then the strongest line is the approximated straight pectoral edge. If not, change the ROI by 20 % and repeat steps 6–8.
Region of Interest for Pectoral Muscle and Block Averaging
After a straight line approximation, the Region of Interest for Pectoral Muscle (ROIPM) is obtained by shifting the approximated straight pectoral edge left at (P w − N p, 1) to (1, P h − N p) and right at (P w + N p, 1) to (1, P h + N p) on a PTG map. This approximation allows for accurate prediction of the pectoral edge and can remove distant outliers, where N p is the horizontal distance in terms of number of pixels. Furthermore, before muscle refining, m × m block averaging is done on ROIPM to obtain important clustered pixels. Blocks with a mean value of 0.51 or higher are considered as 1 while others 0. This step removes those pixels of the ROIPM that are least important or scattered.
Euclidean Distance Regression
Euclidean distance regression (EDR) is designed to refine the straight line approximation into a curve that can demarcate any type of pectoral edge more accurately. A statistical tool such as regression [20] is used to derive the relationship among variables. The regression function is given by y = f(x 1, x 2, ….. x n ), where x is the horizontal and y the vertical coordinates. Given the pectoral edge points (x 1, y 1), (x 2, y 2), …, (x n , y n ) the best fit \( y={a}_0+{a}_1x+\cdots +{a}_m{x}^m \) can be obtained as shown in Fig. 4a where (m ≤ n − 2).
The regression plot of the best fit curve can include the 95 % confidence band or 95 % prediction band. The confidence band indicates that a 95 % true best fit curve lies within the confidence band as shown in Fig. 4b. The prediction band indicates the scattering of the expected pixel of pectoral edge data and accounts for uncertainty in the curve itself as shown in Fig. 4c.
To obtain only those pixels that are more likely to lie on pectoral edge, EDR is applied. The initial value (P w, 1) will choose that pixel from the next non-zero row of block averaged ROIPM, which is at a minimum Euclidean distance until Q (1, P h) is reached as shown by solid line in Fig. 5. Thus the pixels of ROIPM are iterated for minimum Euclidean distance to obtain a set of pixels belonging to the confidence band Pc and is given by:
where D is the Euclidean distance between two pixels and given as \( {D}_{i\;}\left({x}_i,{y}_j\right)=\sqrt{{\left({x}_0-{x}_i\right)}^2+{\left({y}_0-{y}_j\right)}^2} \) where i ∈ M, j ∈ N & (x 0, y 0) = (P w , 1) is the initial value.
The curve obtained from EDR is ragged, and is smoothened using cubic polynomial fitting function with four coefficients. Cubic polynomial function has been chosen because of the pectoral edge shape which is usually curved, and is represented by cubic function, instead of linear function.
Steps of EDR for accurate pectoral muscle detection are as follows:
-
1.
Extract ROIPM from PTG map using approximated straight line (Fig. 6a).
Fig. 6 Pectoral muscle segmentation using Euclidean distance regression: a Region of interest for pectoral muscle (ROIPM); b 5 × 5 Block averaging on ROIPM; c Euclidean distance regression applied to ROIPM (dark gray); d Zoomed view of white rectangular area shown in part c; e Pectoral edge smoothened using cubic polynomial
-
2.
Calculate m × m block averaging (BAG) on ROIPM (Fig. 6b).
$$ \mathrm{BAG}\;\left[x:x+\left(m-1\right),\;y\;:y+\left(m-1\right)\right]=\frac{{\displaystyle \sum_{s=0}^{\left(m-1\right)}{\displaystyle \sum_{t=0}^{\left(m-1\right)}PTG\left(x+s,\;y+t\right)}}}{m^2} $$(5)where
$$ \begin{array}{l}x=1,m+1,2m+1,\dots \dots .M-\left(m-1\right).\kern0.24em \mathrm{and}\hfill \\ {}y=1,m+1,2m+1\dots ..N-\left(m-1\right)\hfill \end{array} $$ -
3.
Apply thresholding to block averaged image BAG as discussed previously.
-
4.
Apply Euclidean distance regression (Eq. 3) on ROIPM with initial value as (P w,1) and iterate till the point Q(1,P h ) is reached giving ragged boundary (Fig. 6c, d).
-
5.
Apply cubic polynomial modeling (Eq. 4) on ragged muscle to obtain smooth curve (Fig. 6e).
Need of Block Averaging on ROIPM
ROIPM is the region near the straight approximated pectoral muscle with pixels that have relatively high value of intensity, and entropy and gradient angle lying between 30° and 80°. EDR initializes with P (P w, 1) and chooses successive pixels with minimum Euclidean distance on ROIPM. When block averaging is not done, false-positive pixels are collected by EDR as shown in Fig. 7b. It gives inaccurate edge with FN of 9.65 % as shown in Fig. 7c. However, when block averaging is applied, false-positive pixels that are not significant are removed before applying EDR (Fig. 7d). This gives a more accurate pectoral edge as shown in Fig. 7e compare to EDR without block averaging (Fig. 7b, c).
Trimming of Sharp Convex Muscles
Due to sharp curvatures in some mammograms like mdb233, the approximated straight line covers false-positive region near lower left edge of the muscle as shown in Fig. 8a (white line). Furthermore, when EDR is applied (gray line) to the ROIPM, a few outliers near the lower left edge of the pectoral muscle bias the pectoral muscle shape. Hence, EDR needs trimming at lower left edge. Therefore, the algorithm has to be converged for the pixel selected first; close to the left chest wall as shown in Fig. 8b, c.
Choice of Parameters
An optimum size of N p is chosen to cover pectoral edge variation for highly concave- or convex-shaped muscle. If a small N p value is chosen, ROIPM will miss pixels that are important for accurate delineation of the pectoral edge. In contrast, if a large value of N p is chosen, unwanted pixels in the vicinity may get detected to form an inaccurate shape of the muscle. The empirical value of N p is chosen by varying it for extreme cases of muscle curvature, and corresponding FP and FN are calculated. The N p that gives the smallest FP and FN is calculated for extreme cases and selected as optimized N p = 70 pixels.
For block averaging, when the block size is large, e.g., 7 × 7, 9 × 9, and 15 × 15, details of the muscles are lost, thus leading to high FN. Conversely, when the block size is smaller, e.g., 3 × 3 pixels or 5 × 5 pixels, muscle details are retained properly. For block averaging in this paper, the value of m × m is chosen as 5 × 5.
Performance Metrics
Two radiologists visually inspect the enhanced mammograms to draw the standard pectoral muscle using a cursor for the evaluation of the performance of proposed automated pectoral muscle detection method. Let P = (p 1,p 2,……p x) be automated pectoral muscle boundary pixels, and Q 1 = (q 11,q 12,……q 1y) and Q 2 = (q 21,q 22,……q 2y) be each radiologist’s manually drawn standard pectoral muscle boundary. For each MLO view mammogram, the accuracy of algorithm is evaluated by three quantitative performance metrics and the two radiologists’ proposed Pectoral Score Index (PSI).
-
1.
Hausdorff Distance (HD): The Hausdorff distance is defined as
$$ HD\left(P,Q\right)= \max \left(h\left(P,Q\right),h\left(Q,P\right)\right) $$(6)where \( h\left(P,Q\right)=\underset{p\in P}{ \max}\underset{q\in Q}{ \min}\left\Vert p-q\right\Vert \)
‖.‖ is the Euclidean distance and where, P ∈ acquired pectoral muscle region, Q ∈ standard pectoral muscle region. The average of the HD with respect to two radiologists is taken.
-
2.
Pixel percentage of false positive (FP): A pixel inside the acquired pectoral edge but outside the reference region. Mathematically, pixel percentage of false positive is given as,
$$ \mathrm{Pixel}\;\mathrm{percentage}\;\mathrm{of}\;\mathrm{F}\mathrm{P}=\frac{\left|P\cup Q\right|-\left|Q\right|\;}{\left|Q\right|}\times 100\% $$(7) -
3.
Pixel Percentage of False Negative (FN):
A pixel outside the acquired pectoral edge but inside the reference region. Mathematically, pixel percentage of false negative is given as,
$$ \mathrm{Pixel}\;\mathrm{percentage}\;\mathrm{of}\;\mathrm{F}\mathrm{N}=\frac{\left|P\cup Q\right|-\left|P\right|\;}{\left|Q\right|}\times 100\% $$(8) -
4.
Pectoral Score Index (PSI):
The pectoral muscle is evaluated using four level pectoral score index recommended by two radiologists after obtaining segmentation results. Deviation, in the form of FP and FN, is mathematically assessed based on the result given by both radiologists. Mean score of the maximum tolerable deviation allowed by radiologists in terms of FP and FN is calculated and expressed as the Pectoral score index given in Table 1. Radiologists classify the observed images into four categories on a scale of pectoral score index: accurate, adequate, moderate and inadequate. Accurate (1) means the segmentation is almost perfect with a straight line or curve overlapping the pectoral muscle edge correctly. All such images have (FP and FN) < 5 %. Adequate (2) means the straight line or curve does not precisely overlap the pectoral muscle but is sufficiently accurate, and can be used for diagnosis. Such images have 5 % ≤ (FP and FN) < 10 %. Moderate (3) means some breast tissue is excluded or some non-breast tissue is included in the muscle segmented result. Images show 10 % ≤ (FP and FN) < 15 %. Inaccurate (4) means the straight line or curve is not present or completely inaccurate; images have (FP and FN) > 15 %.
Table 1 Pectoral score index
Results and Comparative Analysis
The proposed algorithm is applied to three different types of database. To evaluate the efficiency of the method, performance evaluation metrics mentioned earlier are used. Figures 9, 10, and 11 demonstrate the results obtained automatically from the proposed method. As is evident, the proposed algorithm is able to detect muscles which are curved and completely overlapped by dense tissues and texture.
The results of the proposed method are compared with two well-known pectoral muscle detection algorithms proposed by Kwok et al. [10] and Kinoshita et al. [8] on the same dataset of images and platform. Tables 2, 3, and 4 summarize the comparative analysis of all these methods for miniMIAS (200 images), CR (100 images) and FFDM (40 images), respectively. The table shows mean and standard deviation for all three metrics calculated between the automatically detected and manually drawn pectoral muscle with number of images marked as accurate, adequate, moderate and inaccurate with reference to PSI. In the present study, pectoral muscle segmentation is considered to be acceptable when (FP and FN) ≤ 10 %.
Algorithm Processing Time
The algorithm is implemented on MATLAB ver. R2013b. The computer used is a Dell PC with Intel core 2 Duo processor T6670 clocked at 2.2 GHZ with 2 GB RAM. The average processing time required for preprocessing step is 1.85 s, Hough texture based straight line estimation is 2.721 s and Euclidean distance regression with polynomial modeling for curve fitting is 2.089 s. Computational time for valid straight line estimation is longer than EDR. As for valid straight line estimation, if needed, ROI needs to be tuned in iterations. Thus the proposed method requires an average processing time of 4.81 s for muscle detection. The processing time required by Kwok method and Kinoshita method is 6.012 and 3.526 s, respectively. While the Kinoshita method requires less processing time, it is not efficient because it only detects straight edges without taking tissue occlusion into account.
Limitations
The proposed method sometimes fails in the detection of muscle that is small in size and low in contrast. Additionally, it must be noted that the accuracy of the algorithm depends on the accuracy of the straight line approximation of the pectoral edge and ROIPM. The proposed method will prove to be more accurate if the accuracy of the above steps increases.
Conclusion
The strength of the proposed method is that it can detect muscle even when completely obscured by dense tissues. In addition, another advantage is its robustness in dealing with multiple textures and artifacts present on the muscle and its edge. Experimentation also shows that preliminary extraction of ROIPM reduces the chance of false detection of muscle. Furthermore, it can be seen from Tables 2, 3, and 4 that for all types of database, the proposed method outperforms Kwok [10] and Kinoshita [8] method in terms of HD (in millimeter), FP (in percent) and FN (in percent). In Kwok’s method, because of extensive intensity overlap between pectoral and non-pectoral regions, the straight line estimation technique is not suitable as it is based on iterative threshold selection that affects curve segmentation [21]. In contrast, in the proposed work the Hough transform applied on a PTG map gives accurate straight line estimations even though muscle may be overlapped by dense tissues. The Kinoshita method demonstrates better performance when images have straight-line edges and dissatisfactory performance with other types of edges of the pectoral muscle region [8]. However, in the present study EDR helps to detect curved edges. Hence, the proposed method automatically mimics the decision making capability of a radiologist.
A novel texture gradient-based method for detection of the pectoral muscle has been presented in this work. The application of the Hough transform on a probable texture gradient map instead of intensity map, ROIPM extraction, block averaging and EDR with cubic modeling are used to detect pectoral muscle accurately. Furthermore, the algorithm relies on texture, gradient and intensity to detect pectoral muscle, thus overcoming the limitations of state-of-the-art methods. The proposed algorithm is tested on 340 mammograms from three databases, and in the case of 96.75 % of mammograms, pectoral muscle is segmented with an acceptable pectoral score index. With its high accuracy, robustness, and fast processing time, this method has the potential to be an efficient means of automatic pectoral muscle segmentation of MLO mammograms in CAD.
This method shows scope for future research. To overcome the limitations discussed previously, the present method may be added to a region-based contrast enhancement technique [22], and automatic assessment of mammographic positioning [21]. Moreover, when an image is properly exposed at optimal contrast, the pectoral muscle can be differentiated from other glandular tissue with relative ease.
In addition, while drawing a PTG map additional texture features could improve the accuracy of pectoral segmentation. Furthermore, polynomial modeling can be made adaptive with respect to variations in parameters like the sum of squares due to error (SSE) to reduce FPs and FNs in the detection of muscle.
References
Ferrari R, Rangayyan R, Borges R, Frere A: Segmentation of the fibro-glandular disc in mammograms using Gaussian mixture modeling. Med Biol Eng Comput 42(3):378–387, 2004
Eklund GW, Cardenosa G, Parsons W: Assessing adequacy of mammographic image quality. Radiology 190(2):297–307, 1994
Saha PK, Udupa JK, Conant EF, Chakraborty DP, Sullivan D: Breast tissue density quantification via digitized mammograms. IEEE Trans Med Imaging 20:792–803, 2001
Gupta R, Undrill PE: The use of texture analysis to delineate suspicious masses in mammography. Phys Med Biol 40:835–855, 1995
Karssemeijer N: Automated classification of parenchymal patterns in mammograms. Phys Med Biol 43:365–378, 1998
Aylward SR, Hemminger BM, Pisano ED: In: Karssemeijer N, et al Eds. Mixture modeling for digital mammogram display and analysis. Digital Mammography, Comput Imag and Vision Series, vol. 13. Kluwer Academic Publishers, Dordrecht, 1998, pp 305–312
Ferrari RJ, Rangayyan RM, Desautels JEL, Frere AF: Segmentation of mammograms: Identification of the skin—air boundary, pectoral muscle, and fibro-glandular disc. In: Yaffe MJ, Ed. IWDM 2000. Proc. 5th Int Workshop Digital Mammography, Toronto, ON, Canada, 573-579, 2001
Kinoshita SK, Azevedo Marques PM, Pereira Jr, RR, Rodrigues JA, Rangayyan RM: Radon-domain detection of the nipple and the pectoral muscle in mammograms. J Digit Imaging 21(1):37–49, 2008
Ferrari RJ, Rangayyan RM, Desautels JEL, Borges RA, Frere AF: Automatic identification of the pectoral muscle in mammograms. IEEE Trans Med Imaging 23(2):232–245, 2004
Kwok SM, Chandrasekhar R, Attikiouzel Y, Rickard MT: Automatic pectoral muscle segmentation on mediolateral oblique view mammograms. IEEE Trans Med Imaging 23(9):1129–1140, 2004
Raba D, Oliver A, Marti J, Peracaula M, Espunya J: Breast segmentation with pectoral muscle suppression on digital mammograms. Pattern Recog and Image Anal, Lecture Notes in Computer Science, vol. 3523. Springer, Berlin Heidelberg, 2005, pp 471–478
Ma F, Bajger M, Slavotinek JP, Bottema MJ: Two graph theory based methods for identifying the pectoral muscle in mammograms. Pattern Recogn 40(9):2592–2602, 2007
Zhou C, Wei J, Chan H, Paramagul C, Hadjiiski L, Sahiner B, Douglas J: Computerized image analysis: texture-field orientation method for pectoral muscle identification on MLO-view mammograms. Med Phys 37(5):2289–2299, 2010
Chakraborty J, Mukhopadhyay S, Singla V, Khandelwal N, Bhattacharyya P: Computerized image analysis: texture-field orientation method for pectoral muscle identification on MLO-view mammograms. J Digit Imaging 25:387–399, 2012
Suckling J, Parker J, Dance D, Astley S, Hutt I, Boggis C, Ricketts I, Stamatakis E, Cerneaz N, Kok S, et al: The mammographic image analysis society digital mammogram database, Exerpta Medical. Int Congr Ser 1069:375–378, 1994
Mustra M, Grgic M: Robust automatic breast and pectoral muscle segmentation from scanned mammograms. Signal Process 93(10):2817–2827, 2013
Jain AK: Fundamentals of Digital Image Processing Prentice Hall Information and System Sciences Series. Prentice Hall, NJ, USA, 1989
Zuiderveld K: Contrast limited adaptive histogram equalization. Graphic gems IV. Academic Press Professional, San Diego, 1994, pp 474–485
Gonzalez RC, Woods RE: Digital image processing, 3rd edition. Pearson Prentice Hall, New Jersey, 2008
Motulsky HJ, Christopoulos A: Fitting models to biological data using linear and nonlinear regression. A practical Guide to Curve Fitting. GraphPad Software Inc., San Diego CA, www.graphpad.com, 2003
Kwok SM: Attribute-driven segmentation and analysis of mammograms. Ph.D. dissertation: University of Western Australia- School of Electrical, Electronic and Computer Engineering, and Centre for Intelligent Information Processing Systems. Ch 4-5, 2004
Morrow WM, Paranjape RB, Rangayyan R, Desautels JEL: Region-based contrast enhancement of mammograms. IEEE Trans Med Imaging 11(3):392–406, 1992
Acknowledgments
The authors would like to thank Dr. Varsha S. Hardas, Consultant Radiologist of Star Imaging & Research Centre, Pune, India, for providing FFDM images and assessing the segmentation results. Thanks is also given to Dr Suchitra Mehta, Central India Cancer Research Institute (CICRI), Nagpur, India, for providing CR Mammograms and also for assessing segmentation results. We also acknowledge the financial support provided by Centre of Excellence on Combedded System: Hybridization of Communication and Embedded Systems under TEQIP 1.2.1 at VNIT, Nagpur.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bora, V.B., Kothari, A.G. & Keskar, A.G. Robust Automatic Pectoral Muscle Segmentation from Mammograms Using Texture Gradient and Euclidean Distance Regression. J Digit Imaging 29, 115–125 (2016). https://doi.org/10.1007/s10278-015-9813-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-015-9813-5