Abstract
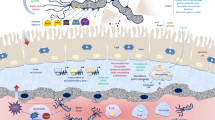
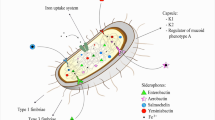
Pseudomonas aeruginosa can penetrate the extracellular mucin barrier formed by the intestinal epithelial cell layer and establish gut-derived sepsis in immunocompromised patients. We found that two efficient mechanisms, flagellar motility and mucin degradation, are needed for penetration of P. aeruginosa through the mucin barrier. Deletion of the flagellar motility-related gene, the filament protein gene fliC, the cap protein gene fliD, and the motor complex protein genes motABCD from P. aeruginosa PAO1 decreased association of P. aeruginosa with the apical surface of human epithelial colorectal adenocarcinoma (Caco-2) cells. A penetration experiment using an artificial mucin layer suggested that the decreased penetration is caused by attenuation of mucin penetration ability. Additionally, the presence of P. aeruginosa decreased the total mucin, including the secreted mucin protein MUC2, on the surface of the Caco-2 cell monolayer, regardless of flagellar motility. Construction of the PAO1 mutant series knocked out 12 putative serine protease genes and identified the mucD gene, which participated in degradation of total mucin, including MUC2. Furthermore, decreased association with the surface of the Caco-2 cell monolayer was observed in the mucD mutant, and the decrease was synergistically amplified by double knockout with fliC. We conclude that P. aeruginosa can penetrate the mucin layer using flagellar motility and mucin degradation, which is dependent on the MucD protease or the mucD gene-related protease.






Similar content being viewed by others
References
Vincent JL, Rello J, Marshall J, Silva E, Anzueto A, Martin CD, et al. International study of the prevalence and outcomes of infection in intensive care units. JAMA. 2009;302:2323–9.
Shimizu K, Ogura H, Goto M, Asahara T, Nomoto K, Morotomi M, et al. Altered gut flora and environment in patients with severe SIRS. J Trauma. 2006;60:126–33.
Tancrede CH, Andremont AO. Bacterial translocation and gram-negative bacteremia in patients with hematological malignancies. J Infect Dis. 1985;152:99–103.
Bertrand X, Thouverez M, Talon D, Boillot A, Capellier G, Floriot C, et al. Endemicity, molecular diversity and colonisation routes of Pseudomonas aeruginosa in intensive care units. Intensive Care Med. 2001;27:1263–8.
Marshall JC, Christou NV, Meakins JL. The gastrointestinal tract. The “undrained abscess” of multiple organ failure. Ann Surg. 1993;218:111–9.
Koh AY, Mikkelsen PJ, Smith RS, Coggshall KT, Kamei A, Givskov M, et al. Utility of in vivo transcription profiling for identifying Pseudomonas aeruginosa genes needed for gastrointestinal colonization and dissemination. PLoS ONE. 2010;5:e15131.
Dharmani P, Srivastava V, Kissoon-Singh V, Chadee K. Role of intestinal mucins in innate host defense mechanisms against pathogens. J Innate Immun. 2009;1:123–35.
McGuckin MA, Lindén SK, Sutton P, Florin TH. Mucin dynamics and enteric pathogens. Nat Rev Microbiol. 2011;9:265–78.
Tlaskalová-Hogenová H, Stěpánková R, Kozáková H, Hudcovic T, Vannucci L, Tučková L, et al. The role of gut microbiota (commensal bacteria) and the mucosal barrier in the pathogenesis of inflammatory and autoimmune diseases and cancer: contribution of germ-free and gnotobiotic animal models of human diseases. Cell Mol Immunol. 2011;8:110–20.
Hirakata Y, Izumikawa K, Yamaguchi T, Igimi S, Furuya N, Maesaki S, et al. Adherence to and penetration of human intestinal Caco-2 epithelial cell monolayers by Pseudomonas aeruginosa. Infect Immun. 1998;66:1748–51.
Hirakata Y, Finlay BB, Simpson DA, Kohno S, Kamihira S, Speert DP. Penetration of clinical isolates of Pseudomonas aeruginosa through MDCK epithelial cell monolayers. J Infect Dis. 2000;181:765–9.
Conrad JC, Gibiansky ML, Jin F, Gordon VD, Motto DA, Mathewson MA, et al. Flagella and pili-mediated near-surface single-cell motility mechanisms in P. aeruginosa. Biophys J. 2011;100:1608–16.
Okuda J, Hayashi H, Okamoto M, Sawada S, Minagawa S, Yano Y, et al. Translocation of Pseudomonas aeruginosa from the intestinal tract is mediated by the binding of ExoS to an Na,K-ATPase regulator, FXYD3. Infect Immun. 2010;78:4511–22.
Herrmann A, Davies JR, Lindell G, Mårtensson S, Packer NH, Swallow DM, et al. Studies on the “insoluble” glycoprotein complex from human colon. Identification of reduction-insensitive MUC2 oligomers and C-terminal cleavage. J Biol Chem. 1999;274:15828–36.
Johansson ME, Phillipson M, Petersson J, Velcich A, Holm L, Hansson GC. The inner of the two Muc2 mucin-dependent mucus layers in colon is devoid of bacteria. Proc Natl Acad Sci USA. 2008;105:15064–9.
Göttke MU, Keller K, Belley A, Garcia RM, Hollingsworth MA, Mack DR, et al. Functional heterogeneity of colonic adenocarcinoma mucins for inhibition of Entamoeba histolytica adherence to target cells. J Eukaryot Microbiol. 1998;45:17S–23S.
Mattar AF, Teitelbaum DH, Drongowski RA, Yongyi F, Harmon CM, Coran AG. Probiotics up-regulate MUC-2 mucin gene expression in a Caco-2 cell-culture model. Pediatr Surg Int. 2002;18:586–90.
Silva AJ, Pham K, Benitez JA. Haemagglutinin/protease expression and mucin gel penetration in El Tor biotype Vibrio cholerae. Microbiology. 2003;149:1883–91.
Sheikh J, Czeczulin JR, Harrington S, Hicks S, Henderson IR, Le Bouguénec C, et al. A novel dispersin protein in enteroaggregative Escherichia coli. J Clin Invest. 2002;110:1329–37.
Aristoteli LP, Willcox MD. Mucin degradation mechanisms by distinct Pseudomonas aeruginosa isolates in vitro. Infect Immun. 2003;71:5565–75.
Henke MO, John G, Rheineck C, Chillappagari S, Naehrlich L, Rubin BK. Serine proteases degrade airway mucins in cystic fibrosis. Infect Immun. 2011;79:3438–44.
Rashid MH, Kornberg A. Inorganic polyphosphate is needed for swimming, swarming, and twitching motilities of Pseudomonas aeruginosa. Proc Natl Acad Sci USA. 2000;97:4885–90.
Toutain CM, Zegans ME, O’Toole GA. Evidence for two flagellar stators and their role in the motility of Pseudomonas aeruginosa. J Bacteriol. 2005;187:771–7.
Doyle TB, Hawkins AC, McCarter LL. The complex flagellar torque generator of Pseudomonas aeruginosa. J Bacteriol. 2004;186:6341–50.
Hatano K, Matsumoto T, Furuya N, Hirakata Y, Tateda K. Role of motility in the endogenous Pseudomonas aeruginosa sepsis after burn. J Infect Chemother. 1996;2:240–6.
Fleiszig SM, Arora SK, Van R, Ramphal R. FlhA, a component of the flagellum assembly apparatus of Pseudomonas aeruginosa, plays a role in internalization by corneal epithelial cells. Infect Immun. 2001;69:4931–7.
Bucior I, Pielage JF, Engel JN. Pseudomonas aeruginosa pili and flagella mediate distinct binding and signaling events at the apical and basolateral surface of airway epithelium. PLoS Pathog. 2012;8:e1002616.
Liu Z, Miyashiro T, Tsou A, Hsiao A, Goulian M, Zhu J. Mucosal penetration primes Vibrio cholerae for host colonization by repressing quorum sensing. Proc Natl Acad Sci USA. 2008;105:9769–74.
Stover CK, Pham XQ, Erwin AL, Mizoguchi SD, Warrener P, Hickey MJ, et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature (Lond). 2000;406:959–64.
Okuda J, Hayashi N, Tanabe S, Minagawa S, Gotoh N. Degradation of interleukin 8 by the serine protease MucD of Pseudomonas aeruginosa. J Infect Chemother. 2011;17:782–92.
Schäfer A, Tauch A, Jäger W, Kalinowski J, Thierbach G, Pühler A. Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene (Amst). 1994;145:69–73.
Hoang TT, Karkhoff-Schweizer RR, Kutchma AJ, Schweizer HP. A broad-host-range Flp–FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene (Amst). 1998;212:77–86.
Kuwayama H, Obara S, Morio T, Katoh M, Urushihara H, Tanaka Y. PCR mediated generation of a gene disruption construct without the use of DNA ligase and plasmid vectors. Nucleic Acids Res. 2002;30:E2.
Heeb S, Itoh Y, Nishiyo T, Schnider U, Keel C, Wade J, et al. Small, stable shuttle vectors based on the minimal pVS1 replicon for use in gram-negative, plant-associated bacteria. Mol Plant-Microbe Interact. 1996;13:232–7.
Hirakata Y, Srikumar R, Poole K, Gotoh N, Suematsu T, Kohno S, et al. Multidrug efflux systems play an important role in the invasiveness of Pseudomonas aeruginosa. J Exp Med. 2002;196:109–18.
Hilgendorf C, Spahn-Langguth H, Regårdh CG, Lipka E, Amidon GL, Langguth P. Caco-2 versus Caco-2/HT29-MTX co-cultured cell lines: permeabilities via diffusion, inside- and outside-directed carrier-mediated transport. J Pharm Sci. 2000;89:63–75.
Song S, Byrd JC, Koo JS, Bresalier RS. Bile acids induce MUC2 overexpression in human colon carcinoma cells. Cancer (Phila). 2005;103:1606–14.
Sambrook J, Fritsch EF, Maniatis TA. Molecular cloning: a laboratory manual. 2nd ed. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989.
Simon R, Priefer U, Plihler A. A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram-negative bacteria. Biotechnology. 1983;1:784–94.
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, et al. The Pfam protein families database. Nucleic Acids Res. 2012;40:D290–301.
Damron FH, Goldberg JB. Proteolytic regulation of alginate overproduction in Pseudomonas aeruginosa. Mol Microbiol. 2012;84:595–607.
Boucher JC, Martinez-Salazar J, Schurr MJ, Mudd MH, Yu H, Deretic V. Two distinct loci affecting conversion to mucoidy in Pseudomonas aeruginosa in cystic fibrosis encode homologs of the serine protease HtrA. J Bacteriol. 1996;178:511–23.
Boucher JC, Yu H, Mudd MH, Deretic V. Mucoid Pseudomonas aeruginosa in cystic fibrosis: characterization of muc mutations in clinical isolates and analysis of clearance in a mouse model of respiratory infection. Infect Immun. 1997;65:3838–46.
Firoved AM, Boucher JC, Deretic V. Global genomic analysis of AlgU (σE)-dependent promoters (sigmulon) in Pseudomonas aeruginosa and implications for inflammatory processes in cystic fibrosis. J Bacteriol. 2002;184:1057–64.
Mathee K, McPherson CJ, Ohman DE. Posttranslational control of the algT (algU)-encoded σ22 for expression of the alginate regulon in Pseudomonas aeruginosa and localization of its antagonist proteins MucA and MucB (AlgN). J Bacteriol. 1997;179:3711–20.
Murray TS, Kazmierczak BI. Pseudomonas aeruginosa exhibits sliding motility in the absence of type IV pili and flagella. J Bacteriol. 2007;190:2700–8.
Imperi F, Ciccosanti F, Perdomo AB, Tiburzi F, Mancone C, Alonzi T, et al. Analysis of the periplasmic proteome of Pseudomonas aeruginosa, a metabolically versatile opportunistic pathogen. Proteomics. 2009;9:1901–15.
Silva AJ, Pham K, Benitez JA. Haemagglutinin/protease expression and mucin gel penetration in El Tor biotype Vibrio cholerae. Microbiology. 2003;149:883–91.
Szabady RL, Yanta JH, Halladin DK, Schofield MJ, Welch RA. TagA is a secreted protease of Vibrio cholerae that specifically cleaves mucin glycoproteins. Microbiology. 2010;157:516–25.
Henderson IR, Czeczulin J, Eslava C, Noriega F, Nataro JP. Characterization of Pic, a secreted protease of Shigella flexneri and enteroaggregative Escherichia coli. Infect Immun. 1999;67:5587–96.
Grys TE, Siegel MB, Lathem WW, Welch RA. The StcE protease contributes to intimate adherence of enterohemorrhagic Escherichia coli O157:H7 to host cells. Infect Immun. 2005;73:1295–303.
Acknowledgments
This research was supported by a Grant-in-Aid for Scientific Research (C) from the Japan Society for the Promotion of Science (JSPS), and a Grant from the “Academic Frontier” Project for Private Universities from the Japanese Ministry of Education, Culture, Sport, Science and Technology (MEXT) to N.G. N.H. was the recipient of a scholarship from The Japan Scholarship Foundation.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Hayashi, N., Matsukawa, M., Horinishi, Y. et al. Interplay of flagellar motility and mucin degradation stimulates the association of Pseudomonas aeruginosa with human epithelial colorectal adenocarcinoma (Caco-2) cells. J Infect Chemother 19, 305–315 (2013). https://doi.org/10.1007/s10156-013-0554-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10156-013-0554-4