Abstract
The aquatic environment is generally affected by the presence of environmental xenobiotic compounds. One of the major xenobiotic detoxifying enzymes is glutathione S-transferase (GST), which belongs to a family of multifunctional enzymes involved in catalyzing nucleophilic attack of the sulfur atom of glutathione (γ-glutamyl-cysteinylglycine) to an electrophilic group on metabolic products or xenobiotic compounds. Because of the unique nature of the aquatic environment and the possible pollution therein, the biochemical evolution in terms of the nature of GSTs could by uniquely expressed. The full complement of GSTs has not been studied in marine organisms, as very few aquatic GSTs have been fully characterized. The focus of this article is to present an overview of the GST superfamily and their critical role in the survival of organisms in the marine environment, emphasizing the critical roles of GSTs in the detoxification of marine organisms and the unique characteristics of their GSTs compared to those from non-marine organisms.
Similar content being viewed by others
Introduction
Glutathione S-transferases (E.C.2.5.1.18) are a family of multifunctional enzymes, which have widely varying catalytic roles. First identified some 40 years ago in corn, GSTs have subsequently been isolated in many other plants and higher organisms (both terrestrial and aquatic) (Frear and Swanson, 1970). In their most prominent role, the GSTs belong to a group of xenobiotic metabolizing phase II detoxification enzymes which represent approximately 1% of the total cellular proteins in eukaryotes and some prokaryotes (Salinas and Wong, 1999). The major role of the detoxification GSTs is cellular defense against chemically induced toxicity. The GST defensive apparatus deactivates hydrophobic xenobiotic compounds, which may otherwise result in cytotoxic and or genotoxic species. The aforementioned deactivation is achieved by the nucleophilic attack of the glutathione (GSH) sulfur to electrophilic substrates, resulting in either nucleophilic substitution or addition depending on the hydrophobic substrate (Armstrong, 1991). The resulting GS-hydrophobic conjugate is exported from animal cells via a membrane-bound ATP-dependent pump followed by metabolism to mercapturates and excretion from the animals (Mathews and van Holde, 1990). A similar system in plants is used for transport of hydrophobic pigmentation compounds such as anthocyanins to their resident vacuole after conjugation with GSH (Marrs, 1996).
Some members of GST superfamily play a prominent role in other physiological processes in both plants and animals. One such major role is the so-called ligandin (transport) functions. The GSTs operate in the isolation and transport of endogenous hydrophobic compounds such as steroids, bilirubin, heme, and bile salts in animal cells and auxins in plants (Marrs, 1996; Salinas and Wong, 1999). The most important of the ligandin functions may be in the synthesis of the localized hormones prostaglandins and the leukotrienes (Salinas and Wong, 1999). The binding of exogenous ligandin compounds has generally been found to take place at sites other than the active site and may directly result in the loss of catalytic activity due to conformational change upon binding (Nishiihira et al., 1993; Dirr et al., 1994).
The cellular levels of the GSTs are controlled by gene promoter regions, which are specific to each of the GST classes. Induction of the genes coding for the GSTs can be accomplished by either a bifunctional or a monofunctional xenobiotic compound (Fernandes et al., 1996). The two groups of inducers are grouped by their ability to spur the phase I or II portions of the detoxification pathway. In general, bifunctional inducers can be characterized as compounds, which have a strong affinity for the aryl hydrocarbon receptor (AhR) such as the group of polycyclic aromatic hydrocarbons (PAHs) (Figure 1: pathway I). The aforementioned receptor triggers both the phase I and II levels of the detoxification mechanism. Monofunctional inducers do not share the same structural features as their bifunctional counterparts. In fact, members of the family of monofunctional compounds share little structural similarity and ultimately have little effect on the AhR. To understand how the monofunctional inducers operate it is necessary to describe a number of other genetic features unique to the phase II enzymes. Other genetic structures of note in the GST regulatory mechanism include an enhancer of the phase II enzymes. Some of the earliest research into the structure-function relationships within the detoxification mechanism revealed that a dioxin response element (DRE) was responsible for increasing the concentration of the phase II enzymes when the phase I enzyme production is triggered by the AhR (Prochaska et al., 1985). Moreover bifunctional compounds are metabolized to compounds resembling monofunctional species by the phase I enzymes and are thus disposed of by the second phase. The phase II enzyme production via the induced phase I pathway is accomplished through a sequence termed as the antioxidant or electrophilic response element (ARE or EpRE). It is here at the DRE, ARE, and EpRE that the aforementioned monofunctional inducers stimulate phase II enzyme production thus bypassing the induction of the phase I enzymes and triggering only phase II production (Figure 1: pathway II; Prochaska et al., 1985).
A simplified representation of the phase I and II metabolic pathways in a typical liver cell (detoxification center). Route I is a mechanism representative of phase I and II enzymes induction by a bifunctional inducer (xenobiotic compound). Route II is a possible mechanism for phase II enzyme induction by monofunctional inducers (xenobiotic compound) as proposed by Prochaska et al. (1985). In route II, the phase II enzymes only are induced by the presence of the monofunctional inducers. Bifunctional inducers form complexes with the AhR and are thus metabolized to compounds resembling the monofunctional inducers. Uptake by the dioxin response element (DRE) follows metabolism of the bifunctional inducer.
The puzzle of the evolution of the GSTs is to date incomplete. Several classes of GST isoenzymes have been distinguished, such as alpha, kappa, mu, omega, pi, sigma, zeta, etc. based on a variety of criteria, including nucleotide and amino acid sequences, substrate specificities, immunological properties and tertiary/quaternary structural characteristics. As research into the structure—function relationship of the GSTs moves forward, the lines between the classes have become blurred. In fact, many of the so-called class-defining substrates show some form of cross reactivity between the major classes. Discovery of a GST with certain characteristics belonging to a particular group by one classification procedure and substrate specificity analysis, which indicates that the same GST belongs to a different class, is not uncommon. The above phenomena beg the question: does the classification system for the GST superfamily need reform? The answer to this question is “yes” when two possibilities are brought into context. First, the existence of the aforementioned class intermediates could go a long way in explaining the cross reactivity. Second, the response of a GST to a specific substrate may be a “learned” response, that is, exposure to one type of substrate may trigger the production of a GST with a specific response for that substrate. A combination of the above possibilities could create a versatile and powerful tool for response to a broad range of xenobiotic compounds. Over time, the response mounted by the GST genetic mechanism to the environmental contaminants can become a permanent fixture of the organism’s genetic makeup. The aforementioned organism still retains the genetic power to any other assault by a second xenobiotic compound. With the possibilities listed above in mind, it would seem logical that a classification system that does not afford the possibility that an organism’s GST could show a multiple response to a group of characterizing substrates would need replacement. It would seem most logical to either group the GSTs by only the substrates to which they show activity or incorporate the response substrate types into the name of the GST, thus creating another characterizing criteria.
Classification
The GST superfamily includes cytosolic and mitochondrial GSTs. The cytosolic GSTs from mammals are all dimeric with subunit molecular masses from 23 kDa to 27 kDa. Based on amino acid sequence similarities, seven class cytosolic GSTs are recognized in mammalian organisms, designated alpha, mu, pi, sigma, theta, omega, and zeta (Hayes and Pulford, 1995; Board et al., 1997; Armstrong, 1997; Tong et al., 1998; Hayes and McLellan, 1999; Board et al., 2000; Sheehan et al., 2001; Hayes et al., 2005). Further, other class cytosolic GSTs, named beta, delta, epsilon, lambda, phi, rho, and tau, have been identified in nonmammalian organisms, such as plants, insects, bacteria, fungi, helminth, fish, etc. (Sheehan et al., 2001; Dixon et al., 2002; Ding et al., 2003; Konishi et al., 2005a, b). On the other hand, the mitochondrial GST isoenzymes designated kappa have been investigated in mammalian species (Pemble et al., 1996; Jowsey et al., 2003; Ladner et al., 2004; Morel et al., 2004; Robinson et al., 2004). The kappa class GSTs are dimeric with subunits containing 226 amino acid residues. The kappa class GSTs are quite distinct from cytosolic GSTs.
The classification of the GST superfamily has been an evolving process. The most successful early attempt at classification found GSTs placed in one of three categories (alpha, mu, and pi) based on partial amino acid sequences, xenobiotic substrate recognition, and antibody cross reactivity (Mannervik et al., 1985). Following the early classification of mouse, rat, and human cytosolic isozymes by the Mannervik group, work on the fourth (theta) and fifth (omega) GST classifications was taking place nearly simultaneously in 1991 by Meyer et al. (theta) and in 1993 by Tomarev et al. (omega). The theta class was proposed on the basis of a partial amino acid homology of human and rat cytosolic isozymes. While amino acid sequence homology of octopus and squid lens S-crystallins comprised the omega class. In addition, in 1996 Pemble et al. cloned and identified a new GST gene from rat mitochondria that belongs to the kappa class.
In the meantime, the classification of plant GST isozymes was also established independently (Marrs et al., 1995; Marrs, 1996; Droog et al., 1995; Droog, 1997; Neuefeind et al., 1997a, b; Edward et al., 2000; Dixon et al., 2002). The current classification system for plant GSTs recognizes four main classes, two of which are plant specific and the other two are more phylogenetically widespread. The plant-specific classes are phi and tau. Phi class GSTs with herbicide-detoxifying activity are also called type I GSTs and contain three exons and two introns (Shah et al., 1986). Tau class is type III GSTs and contains two exons and one intron (Droog, 1997). The other two classes are theta and zeta classes. Zeta class is called type II GSTs, which have 10 exons and 9 introns (Edward et al., 2000).
In addition, there is a separate GST classification for bacteria and insects. Beta class GSTs in a wide variety of bacteria were reviewed in Vuilleumier (1997). The most generally accepted classification system of insect GSTs recognizes two classes, class I or delta class, which is insect specific, and class II which consists of the omega/zeta/sigma/theta classes, based on sequence alignments and immunological properties (Fournier et al., 1992; Syvanen et al., 1994; Yu, 1996; Ranson et al., 1997; Prapanthadara et al., 1998; Lougarre et al., 1999; Hemingway, 2000). As current research has shown with GSTs from different organisms, a classification process is necessary in cases where a family of closely related GSTs exists. Therefore, a process may need to be developed for the marine GSTs.
So far, there are no clearly established criteria to classify GSTs from marine organisms. Efforts in the classification of marine organisms have been hampered somewhat by the intense focus placed on the determination of the role of GSTs in the detoxification process of marine organism. In fact, only a limited number of marine GSTs have been characterized kinetically and by full amino acid sequencing (Figures 2, 3, 4, 5, 6 and 7; Table 1; George and Young, 1988; Fitzpatrick and Sheehan, 1993; Lin and Chuang, 1993; Fitzpatrick et al., 1995a, b; Kaaya et al., 1999; Angelucci et al., 2000; Gallagher et al., 2000; Gallagher and Sheehy, 2000; Vidal and Narbonne, 2000; Hoarau et al., 2002; Vidal et al., 2002; Yang et al., 2002; 2003). In addition to those fully characterized marine GSTs mentioned above, a number of GSTs have been characterized by immunochemical techniques (Table 2; George and Young, 1988; Lin and Chuang, 1993; Fitzpatrick and Sheehan, 1993; Tomarev et al., 1993; Fitzpatrick et al., 1995a; Gallagher et al., 1996b; James et al., 1998; Perez-Lopez et al., 1998; Angelucci et al., 2000; Gallagher et al., 2000; Perez-Lopez et al., 2000; Melgar-Riol et al., 2001; Hoarau et al., 2002; Vidal et al., 2002). Even though antibodies made against mammalian GSTs may not cross react GSTs from marine organisms, immunological cross reactivity may provide helpful information for the classification of marine GSTs. In spite of the fact that the characterizations of marine GSTs cover alpha, mu, pi, sigma, and theta classes based on the classification criteria of mammalian GSTs, certain marine GSTs may belong to different classes when distinct characteristics are considered. It is evident from the classification of those fully characterized marine GSTs, and the response of the partially characterized GSTs to immunochemical reactivity, that the marine GSTs in general must constitute a dissimilar branch in the GST evolutionary process.
Comparison of the alpha class GSTs from marine animals. The sequence alignment of seven alpha class GSTs from marine animals is available in GenBank. Amino acids highlighted in green are identical in all species. Amino acids highlighted in yellow are identical in at least four species. Dashes show the gaps that were introduced to maximize similarity. 1: Zebrafish (Danio rerio, GenBank accession no. NM_213394); 2: red sea bream (Pagrus major, GenBank accession no. AB158410); 3: red sea bream (P. major, GenBank accession no. AB158411); 4: rock bream (Oplegnathus fasciatus, GenBank accession no. AY734529); 5: silver carp (Hypophthalmichthys molitrix, GenBank accession no. EF100904); 6: bighead carp (Hypophthalmichthys nobilis, GenBank accession no. EF100902); 7: mangrove rivulus (Kryptolebias marmoratus, GenBank accession no. AY626242).
Comparison of the mu class GSTs from marine animals. Sequence alignment of two mu class GSTs from marine animals available in GenBank. Identical amino acid sequences are indicated by highlighting. Dots denote amino acids defining the G-site (GSH binding). 1: Pacific white shrimp (Litopenaeus vannamei, GenBank accession no. AY573381); 2: zebrafish (Danio rerio, GenBank accession no. NM_212676).
Comparison of the pi class GSTs from marine animals. The sequence alignment of various pi class GSTs from marine animals is available in GenBank. Amino acids highlighted in green are identical in all species. Amino acids highlighted in yellow are identical in at least five species. Dots and stars indicate amino acids defining the G-site (GSH binding) and H-site (substrate binding), respectively. Dashes show the gaps which were introduced to maximize similarity. 1: Sockeye salmon (Oncorhynchus nerka, GenBank accession no. AB026119); 2: blue mussel (Mytilus edulis, GenBank accession no. AY557404); 3: Mediterranean mussel (Mytilus galloprocincialis, GenBank accession no. AF527010); 4: Asian clam (Corbicula fluminea, GenBank accession no. AY885667); 5: mussel (Unio tumidus, GenBank accession no. AY885666); 6: zebrafish (Danio rerio, GenBank accession no. AB194127); 7: zebrafish (D. rerio, GenBank accession no. AB194128); 8: European eel (Anguilla anguilla, GenBank accession no. AY530199).
Comparison of the theta class GSTs from marine animals. Sequence alignment of four Theta class GSTs from marine animals available in GenBank. Amino acids highlighted in green are identical in all species. Amino acids highlighted in yellow are identical in three species. Dashes show the gaps that were introduced to maximize similarity. 1: Plaice (Pleuronectes platessa, GenBank accession no. X95199); 2: plaice (P. platessa, GenBank accession no. X95200); 3: largemouth bass (Micropterus salmoides, GenBank accession no. AY335905); 4: mangrove rivulus (Kryptolebias marmoratus, GenBank accession no. DQ525602).
Catalytic Mechanism, General Structure, and Class Structure
The GSTs that have been used as research tools thus far are predominantly from non-marine organisms. Therefore, the intricacies of some of GST structure and function of marine organisms can be derived only from cytosolic GSTs of non-marine organisms. Fortunately most GSTs, marine or non-marine, exhibit variations on a basic theme. There are four basic functions in which a particular GST participates (detoxification, targeting for transmembrane transport, protection of tissues from oxidative damage, and ligandin binding, the nonenzymatic binding for intracellular transport). Ultimately, this means that the structural cues from those non-marine GSTs that have been intensively studied can be used to derive the function of a lesser studied marine GST. Because a majority of the marine GSTs involved are detoxifiers, it is appropriate to highlight this function of the GSTs.
Catalytic Mechanism and General Structure
The most predominant catalytic mechanism for GSTs in relation to the detoxification function is the nucleophilic attack of the GSH sulfur to the electrophilic center of a xenobiotic compound. The quaternary structure for cytosolic GSTs in most organisms is that of a dimer. The subunit molecular weights range from 21 to 29 kDa (Mannervik and Danielson, 1988). Usually, both homodimeric and heterodimeric GSTs are reported within the same organism, but unknown heterodimers exist between differing GST classes (Habig et al., 1974, 1976; Jakoby et al., 1976). In terrestrial organisms, trimers and tetramers have been found to exist in the microsomal forms of GST. The microsomal GSTs have been shown to contain subunits with molecular masses of approximately 14 kDa (Morgenstern et al., 1982). A small number of monomeric GST forms have been found to exist (Overbaugh et al., 1988; Dean et al., 1995). There has been no report of marine GSTs in the aforementioned monomeric or polymeric forms. This does not mean that the exceptional GST forms do not exist in marine organisms, only that there has been none discovered thus far.
Each GST is known to contain a G-site capable of binding the GSH substrate and an H-site that has xenobiotic compound binding capabilities. Because the common factor to all in the highly diverse GST superfamily is the ability to bind GSH, it is logical that the structural features comprising the G-site share a highly conserved amino acid sequence. In fact, the N-terminal region (amino acid residue 1–81) is recognized as such a structural domain (Reinemer et al., 1996; Armstrong, 1997). The bedrock distinction among the GSH sites among different GST classes is the interaction of the protein with the sulfur of GSH. The aforementioned structural differences are discussed further in the following section.
The xenobiotic hydrophobic H-site is the main structure accounting for the variability of GST enzyme activity. As opposed to the highly conserved nature of the G-site, the H-site generally shares little sequence homology between classes. In general, interclass amino acid sequence identity is rarely greater than 35% in the H-site region. The wide range of xenobiotic compounds accepted for catalysis by the GST superfamily seems to have a number of features in common. The most important structural feature required of the xenobiotic compound for binding to the H-site seems to be the carbon to carbon double bond bordering the electron-withdrawing group. The above mentioned feature is characteristic of a Michael acceptor and can be obtained through a phase I (cytochrome P450) detoxification metabolism if the substrate lacks them naturally (van der Oost et al., 1996).
The kinetic mechanisms of GSTs have been studied, and several catalytic mechanisms, including random, ping-pong, and sequential, have been proposed (Mannervik, 1985; Mannervik and Danielson, 1988; Pickett, 1989; Tang and Chang, 1996; Caccuri et al., 1997; Polekhina et al., 2001; Labrou et al., 2001; 2005). The kinetic mechanism of the GST-catalyzed conjugation reaction is very complex and class dependent. For instance, the maize GST I-catalyzed conjugation reaction between GSH and CDNB follows a rapid equilibrium random sequential Bi Bi kinetic mechanism (Labrou et al., 2001; 2005), whereas a steady-state sequential rapid Bi Bi mechanism was proposed for octopus GST, rat GSTs M1-1, M1-2, and A3-3 (Jakobson et al., 1979; Ivanetich et al., 1990; Tang and Chang, 1996).Thus, the structural differences within the active sites (ball and socket, for example) may influence the type of mechanism the GST molecule uses to catalyze its reaction. The catalytic assay universally recognized for the evaluation of GST activity requires 1-chloro-2,4-dinitrobenzene (CDNB) as the hydrophobic substrate (Figure 8). The chlorosubstitution involving CDNB and GSH yields a product that is easily identifiable by spectrophotometric measurement at 340 nm and is the only characteristic GST assay with a crystal structure of a transition state analogue (Armstrong, 1997). Unfortunately not all GSTs have the ability to catalyze CDNB conjugation with GSH. The majority of the GSTs that do not recognize CDNB as a substrate are ignored and thus underestimated in the representation of total GSTs present within an organism. The rate-limiting step in the enzymatic conjugation of CDNB with GSH is in the physical release of product in some GST classes (Johnson et al., 1993). Herein lies the structural importance in the catalytic abilities of the GST classes. A closer examination of the prominent structural features of the GST classes will assist in determination of the possible catalytic capabilities of previously unstudied marine GSTs.
The commonly accepted biological assay for the evaluation of the enzymatic characteristics of the GSTs. The chlorosubstitution involves conjugation of the tripeptide, glutathione (γ-glutamylcystienyl glycine) to the xenobiotic substrate 1-chloro-2,4-dinitrobenzene. The assay product is easily identifiable by spectrophotometric measurement at 340 nm.
Class Structural Features
The catalytic mechanism/general structural features above have stressed the significance of the structural and catalytic similarities and differences in the GST classes. As was noted in the preceding text, the GST classes share some remarkable similarities in their G-site homology and mechanisms. At the same time, the identical classes exhibit a high degree of variability in their H-site homology, while performing similar functions. In this section we highlight the specific structural features that are responsible for the functional aspects of each GST class. A summary of these features is provided in Table 3 (Ji et al., 1992; Sinning et al., 1993; Armstrong et al., 1996; Wilce et al., 1996), and crystal structures of five class GSTs are presented in Figure 9 (Sinning et al., 1993; Ji et al., 1995; Xiao et al., 1996; Oakley et al., 1997; Rossjohn et al., 1998).
Crystal structure of five class GSTs: (a) human alpha GST, (b) rat mu GST, (c) human pi GST, (d) squid sigma GST, and (e) human theta GST. Regions showing ball and the socket areas in alpha, pi, and mu class have been encircled. Source: a, b, c, and e: Protein Database Bank (http://www.rcsb.org/pdb/home/home.do). c: Ji et al., (1995).
Alpha Class GSTs
The subunit interface for the alpha class GSTs is that of a ball-and-socket type joint with the Phe52 serving as the ball and the hydrophobic socket residing between α4 and α5 helices of domain II (Armstrong, 1997) where the GSH and xenobiotic binding domains are referred to as domain I and II, respectively. All alpha, mu, and pi class GSTs have such a ball and socket style interface (Figure 9). Phenylalanine serves as the ball in all three classes and the position of Phe is fairly conserved (Figure 10). The three-dimensional structure of a number of class alpha GSTs revealed a typical subunit molecular mass of approximately 26 kDa (Sinning et al., 1993; Dirr et al., 1994; Cameron et al., 1995), which exists as a dimer in the cytosolic conditions. GSH stabilization in the alpha class GSTs is predominantly due to two amino acid residues. Tyr9 for alpha class GSTs is the stabilization key in the G-site. Site-directed mutagenesis experiments in which Tyr9 was replaced with a Phe residue resulted in a loss of approximately 90% of the enzymatic activity in all except the alpha class GSTs (Rushmore and Pickett, 1993). The reduced loss of activity in the alpha class (mutant alpha class activity was nearly 10-fold higher than the mutants from either the mu or the pi class) was the result of a second major stabilization residue found in the G-site, which is the Arg15 (Rushmore and Pickett, 1993). Compared to the pi and mu GSTs the C-terminal of the alpha class GSTs is longer by some 4 to 8 amino acid residues (Salinas and Wong, 1999). In the remaining ball-and-socket style cytosolic GSTs the shorter C-terminal restricts access to the G-site. The longer alpha C-terminal also forms an α-helix (α9), which comprises a portion of the smaller H-site (Sinning et al., 1993). This helix is thought to be important to dimer stabilization and affects both the GSH-binding rate and the ionization state of the catalytically essential residue Tyr9 (Dirr and Wallace, 1999; Gustafsson et al., 1999). Thus the alpha class C-terminal has a positive effect on catalytic activity by not blocking access to the G-site and forming a portion of the H-site.
Alignment of partial protein sequences of alpha, mu, and pi class GSTs showing the homologous phenylalanine (F) involved in the ball region of the protein. Sequence alignment was performed by ClustalW using Mega 3.1 software. Residue phenylalanine in ball is highlighted by a red circle. 1: Human (Homo sapiens, GenBank accession no. S49975), Phe52; 2: red sea bream (Pagrus major, GenBank accession no. AB158410), Phe52; 3: rock bream (Oplegnathus fasciatus, GenBank accession no. AY734529), Phe52; 4: Norway rat (Rattus norvegicus, GenBank accession no. BC063172), Phe57; 5: Pacific white shrimp (Litopenaeus vannamei, GenBank accession no. AY573381), Phe57; 6: zebrafish (Danio rerio, GenBank accession no. NM_212676), Phe57; 7: human (Homo sapiens, GenBank accession no. NM_000852), Phe56; 8: blue mussel (Mytilus edulis, GenBank accession no. AY557404), Phe48; 9: Asian clam (Corbicula fluminea, GenBank accession no. AY885667), Phe48.
Mu Class GSTs
The mu class GSTs contain many of the same structural attributes as their alpha class counterparts but with slight variations (Ji et al., 1992; Sinning et al., 1993; Dirr et al., 1994; Raghunathan et al., 1994). For example, the representative GSTs for the mu class also have subunit molecular mass of approximately 26 kDa and a subunit interaction for the dimer consisting of ball-and-socket type interface. A highly recognizable feature of the mu class GSTs is the so-called mu loop, which is the result of an insertion in domain I. Another structural attribute, which is recognizable in both mu and pi forms of GST, is the C-terminal wall (Sinning et al., 1993). The wall ultimately restricts xenobiotic access to the H-site. GSH stabilization in the mu class also differs by residue placement and type. The major stabilization factor in the G-site of mu class GSTs is Tyr6. In contrast to the alpha class, there is no assistance provided in the mu class G-site by an Arg residue.
Pi Class GSTs
The pi class GSTs also have the ball-and-socket style interface (Armstrong, 1997). The three-dimensional structure of the pi GSTs reveals that it also has a GSH stabilization segment consisting of a lone Tyr (Tyr7) like mu GSTs. Among the ball-and-socket style the pi class seems to be closest in structural relation to the mu class with a subunit molecular mass of slightly smaller 23 kDa (Reinemer et al., 1992; Dirr et al., 1994; Garcia-Saez et al., 1994). Beyond the similarities listed in the preceding text, the pi and mu classes share C-terminal structural similarities. On average, the C-terminal end is 4 to 8 amino acid residues shorter than their alpha counterpart and forms the previously mentioned wall that results in a partially blocked access to the xenobiotic binding site (Salinas and Wong, 1999). Lastly, the pi and mu classes share an H-site form that is larger and more open to solvent entry than the alpha class GSTs (Sinning et al., 1993).
Theta Class GSTs
The theta class GSTs are quite distinct from alpha, mu, or pi class GSTs. In theta class GSTs the main subunit interface lacks a ball-and-socket style and the cleft between the two subunits is significantly less noticeable (Reinemer et al., 1996; Rossjohn et al., 1998). Structurally it seems that the theta class GSTs are missing the loop feature that holds the Phe residue for interaction with the α4 and α5 helices and the hydrophobic pockets within the helices themselves (Armstrong, 1997). Therefore, the hydrophilic GSTs rely mainly on electrostatic and hydrogen bond interactions for binding at the dimer interface. The theta class GSTs show typical cytosolic GST subunit molecular mass with the average at 27 kDa. A unique structural feature in the theta class GSTs is the involvement of a Ser residue as the GSH stabilization residue, in contrast to the involvement of a Tyr residue in alpha, mu, and pi class GSTs. Also, the G-site of the theta class GSTs is, in general, much deeper than that of alpha, mu, and pi class GSTs (Wilce et al., 1996).
Sigma Class GSTs
Like the theta class GSTs, the sigma class GSTs are also lacking in both components of the ball-and-socket interface present in the alpha, mu, and pi class GSTs (Armstrong, 1997), and electrostatic interactions near the active site which play a key role in protein stability in this group of GSTs (Stevens et al., 2000). However, the sigma and theta class GSTs differ in certain characteristics. The sigma class GSTs show subunit molecular mass with the average at 23 kDa, in contrast to 27 kD for the theta class. The sigma class GSTs rely on a Tyr residue, instead of Ser, for GSH stabilization in the G-site. This stabilization residue is a feature that the sigma class shares with the ball-and-socket style cytosolic GSTs (Kanaoka et al., 1997). Other structural comparisons between the sigma class GSTs and the alpha, mu, or pi class GSTs reveal some surprising similarities. For example, the squid sigma class GST has a largely open hydrophobic active site cavity as opposed to the partial blockage of the alpha and mu class active site cavities (Armstrong et al., 1996). The hydrogen bonding interaction between the glutathione and the squid class sigma GST is very similar to that of the pi and alpha class GSTs but differ from the mu class in the hydrogen bonding pattern to the cystein carbon to oxygen bond in GSH (Ji et al., 1995).
The aforementioned features provide a snapshot of the family of reported marine GSTs as a structurally diverse group of multifunctional enzymes. It is the multifunction nature of the GST superfamily that may yield some promising uses from certain members of the GST family. Among the most promising and interesting roles proposed for marine GSTs is that of biomarker and/or detoxification tool.
Phylogenic Relationship
The evolutionary relationships of the cytosolic GST isozymes have been the subject of some debate. At the forefront of the debate is the order in which the GST classes began to divide from the archetype GST and how an organism of advanced evolutionary age could contain activity from what is thought to be a relatively young GST class. To understand the evolutionary process we must first understand the beginning of GST evolution. The consensus developed in the literature thus far is that the archetype GST is the theta class (Buetler and Eaton, 1992; Armstrong et al., 1996; Marrs, 1996; Salinas and Wong, 1999). Ancient roots of the theta class are evident in its distribution among a wide variety of species (bacteria, plants, mammals, etc.). Another good indication to the primitive roots of the theta class is in the simplicity of its gene structure. For example, when a comparison is made of the plaice GST-A gene structure to its most similar cousin belonging to Drosophila GST 1.1 (Toung et al., 1991; Leaver and George, 1995; Armstrong et al., 1996), research has shown that both of the above GST genes are intronless. This is in opposition to the GST genes of younger organisms, where multiple introns are the rule.
The debate as to which of the GST classes diverged from the theta archetype is generally confined to a few proposed evolutionary pathways (Buetler and Eaton, 1992; Armstrong et al., 1996; Salinas and Wong, 1999). In the evolutionary pathway proposed by Buetler and Eaton (1992; Figure 11) each of the individual classes, as they are known, diversified from a gene duplication in the theta archetype starting with the pi and mu classes and ending with the alpha class. In the above scheme, the evidence for the initial divergence of the pi class is the presence of ethacrynic acid activity in the relatively ancient aquatic species Cnidaria (Stenersen et al., 1987). The Buetler and Eaton (1992) evolutionary pattern does not pinpoint the step from which the sigma class has sprung, only that this class “may be the result of an evolutionary event limited to relatively young mu or alpha classes in old organisms, such as the parasite nematode, was possible through a transfer of genetic information from host to parasite.” The presence of activity such as the aforemetioned is explained much more precisely by the remaining evolutionary proposals. The two remaining proposed pathways share similar intermediates with only small differences in the order of the individual class divergence.
Evolutionary relationship among the alpha (α), mu (μ), pi (π), theta (θ), S-crystallin, and sigma (σ) class GSTs as proposed by Buetler and Eaton (1992). In the evolution of modern GST classes, the archetype GST is proposed to be the enzyme from the θ class. A dual divergence from the θ class is proposed, with the π and σ classes being first to split. All divergence in the Buetler and Eaton pathway springs directly from the θ class without class intermediates.
In the pathways proposed by Salinas and Wong (1999; Figure 12) the individual GST classifications, as they are known today, did not spring directly from the archetype theta class as proposed by Buetler and Eaton (1992). Instead, an intermediate class containing the characteristics of all three of the modern alpha/mu/pi classes was first to diverge. It was also proposed that the sigma class diverged directly from the theta progenitor prior to intermediate bifurcation. From the proposed intermediate the pi class was first to diverge, thus leaving a second alpha/mu intermediate. From the secondary intermediate the mu class branched therefore leaving the alpha class as the final step in the evolutionary pathway.
GST evolutionary relationship among the alpha (α), mu (μ), pi (π), theta (θ), S-crystallin, and sigma (σ) class GSTs, as proposed by Salinas and Wong (1999). In the evolution of modern GST classes, the archetype GST is proposed to be the enzyme from the θ class. A dual divergence from the θ class was proposed with a α/μ/π intermediate and σ classes being first to split. All divergence in proposed the pathway do not evolve directly from the θ class. Instead, class intermediates are proposed, which bear the characteristics of all the classes comprising the intermediate.
The GST evolutionary pathway as proposed by Armstrong (1997), while similar to the pathway above proposed by Salinas and Wong (1999), has a few major differences (Figure 13). First, and foremost, Armstrong (1997) relied on three-dimensional and catalytic residue structural differences to highlight the evolutionary distinctions between the GST classes. For example, for all classes the catalytic amino acid residue in theta class is tyrosine. Three dimensional structures, on the other hand, indicated a clear distinction between the theta/sigma classes and the alpha/mu/pi classes, with the latter containing a “ball-and-socket” style dimer interface using a phenylalanine residue (ball) on one monomer and a hydrophobic socket located between two helices on the opposing monomer. The theta and sigma GST classifications are missing the residues required for the “ball-and-socket” interaction and therefore rely solely on electrostatic interactions across the dimer interface. Armstrong (1997) proposed an evolutionary pathway, suggesting that the sigma class could not have evolved directly from a theta progenitor since the sigma class contains a catalytic residue more closely related to the newer sigma/alpha/mu/pi intermediate. This proposed pathway could explain what would be a rather strange evolutionary track taken by the sigma class in the Salinas and Wong (1999) pathway. Second, Armstrong et al. (1996) proposed that the ultimate progenitor of the eukaryotic theta is thought to be the microsomal kappa class, of which little is functionally known. The kappa class is preceded in the evolutionary chain by the bacterial theta class. The final difference between the evolutionary tracks proposed by Armstrong (1997) and Salinas and Wong (1999) is in the branching of the alpha, mu, and pi individual classes from the aforementioned triple intermediate (Figures 12 and 13). While Salinas and Wong (1999) have proposed a pi class deviation from the triple intermediate followed by the mu class, Armstrong (1997) envisions the deviation of the mu class followed by the pi class. Both of the review articles (Armstrong, 1997; Salinas and Wong, 1999) rely on previously published research (Pemble and Taylor, 1992; Dirr et al., 1994) to draw their evolutionary conclusions. Currently it seems less important as to which modern class split from the alpha/mu/pi intermediate first than the presence of this intermediate itself.
The evolutionary relationship among the alpha (α), mu (μ), pi (π), theta (θ), S-crystallin, kappa (κ), and sigma (σ) class GSTs as proposed by Armstrong (1997). In this proposal the progenitor of the modern GST classes is the microsomal ê class, from which the θ class is believed to have evolved. A dual divergence from a σ/α/μ/π intermediate is proposed, with further intermediates bearing the enzymatic characteristics of successively fewer modern GST classes. Divergence in the proposed pathway means that the modern GST classes do not evolve directly from the θ class.
It is believed that the Buetler and Eaton theory, while having some merit, is not a full explanation for the divergence of the GST classes. It is unlikely to think that all ancient organisms (marine organisms in particular) that contain activity of younger GSTs do so because of a relationship with either an evolutionary younger organism or vice versa. Therefore, the evolutionary pathway, as proposed by Buetler and Eaton (1992), may not be appropriate for marine GST classification. A more simplistic and therefore plausible explanation would be the existence of class intermediates as proposed by both Salinas and Wong (1999) and Armstrong (1997). When data are compiled on aquatic species which have been characterized by amino acid sequencing, substrate specificity, or immunochemical properties, it is apparent that the proposed evolutionary pathways containing class intermediates are the most feasible. However, modifications must be made to the evolutionary pathways because some marine GSTs seem to fit the criteria for a number of the accepted GST classifications. The simplest answer to the contradictory nature of the classification of marine organisms may be that evolutionarily older marine organisms have retained the characteristics of the sigma/alpha/mu/pi, alpha/mu/pi, alpha/pi, or the alpha/mu intermediates and thus do not fit a single class criterion. For example, characterization of channel catfish (Ictalurus punctatus) yielded immunochemical cross reactivity to alpha and pi classes with limited reactivity toward the mu class (Gallagher et al., 1996b). Similar results were obtained when GSTs from English sole (Pleuronectes vetulus) were characterized. The sole GSTs showed cross reactivity with both mu and pi classes (Gallagher et al., 1996a). Sequencing experiments also indicate conflicting results. On comparison of differing signpost sequences it is possible to place a GST into a number of classes (Fitzpatrick et al., 1995a). Therefore, we feel that a new classification criterion should be used to designate those GSTs that fit into multiple classes, especially those from marine organisms. Based on the reported sequences of marine GSTs (Figures 2, 3, 4, 5, 6 and 7, Leaver et al., 1993; Tomarev et al., 1993; Leaver et al., 1997; Kudo et al., 1999; Strausberg and Feingold, 2002; Contreras-Vergara et al., 2004; Doi et al., 2004; Yang et al., 2004; Doyen et al., 2005; Konishi et al., 2005a, b; Lee et al., 2005; Nam et al., 2005; Suzuki et al., 2005; Woods et al., 2005; Hoarau et al., 2006; Lee et al., 2006; Liao et al., 2006), a phylogenic tree was constructed via the NJ (neighbor-joining) method using the Mega 3.1 software which shows the evolutionary relationship of various classes of GST (Figure 14). A new evolutionary relationship of GSTs from marine organisms has been proposed (Figure 15) based on the evolutionary tree shown in Figure 14. In this proposal the pi/sigma intermediate split off early into pi and sigma. Then mu split off from alpha/mu/rho/theta intermediate followed by alpha and then rho/theta split off into rho and theta. The most prominent difference in this proposal from the other three proposals mentioned in the preceing text is that theta is located at the end rather than at the beginning of the evolutionary track. Although some GSTs belong to different classes, they have closer evolutionary relationship than those in the same class. Therefore, it may not be appropriate to use only sequence information for the classification of marine GSTs. However, sequencing data in addition to other important characteristics, such as substrate specificities, immunological properties, three-dimensional structures, and biological functions, could be utilized to develop a new classification criterion for marine organism GSTs.
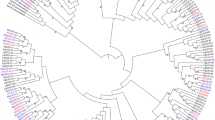
Phylogenic tree of reported GSTs from marine organisms. The tree was constructed by the NJ method using the Mega 3.1 software. Selected bootstrap values from 100 traditional trees are shown in each internal branch of the phylogenetic tree nodes. Branch length is proportional to estimates of evolutionary change. The amino acid sequences are zebrafish (Danio rerio, GenBank accession no. NM_213394); 2: red sea bream (Pagrus major, GenBank accession no. AB158410); 3: red sea bream (P. major, GenBank accession no. AB158411); 4: rock bream (Oplegnathus fasciatus, GenBank accession no. AY734529); 5: silver carp (Hypophthalmichthys molitrix); 6: bighead carp (Hypophthalmichthys nobilis); 7: mangrove rivulus (Kryptolebias marmoratus, GenBank accession no. AY626242); 8: Pacific white shrimp (Litopenaeus vannamei, GenBank accession no. AY573381); 9: zebrafish (Danio rerio, GenBank accession no. NM_212676); 10: sockeye salmon (Oncorhynchus nerka, GenBank accession no. AB026119); 11: blue mussel (Mytilus edulis, GenBank accession no. AY557404); 12: Mediterranean mussel (Mytilus galloprocincialis, GenBank accession no. AF527010); 13: Asian clam (Corbicula fluminea, GenBank accession no. AY885667); 14: mussel (Unio tumidus, GenBank accession no. AY885666); 15: zebrafish (Danio rerio, GenBank accession no. AB194127); 16: zebrafish (D. rerio, GenBank accession no. AB194128); 17: European eel (Anguilla anguilla, GenBank accession no. AY530199); 18: red sea bream (Pagrus major, GenBank accession no. AB158412); 19: arrow squid (Ommastrephes sloanei pacificus, GenBank accession no. AH003423); 20: plaice (Pleuronectes platessa, GenBank accession no. X95199); 21: plaice (P. platessa, GenBank accession no. X95200); 22: largemouth bass (Micropterus salmoides, GenBank accession no. AY335905); 23: mangrove rivulus (Kryptolebias marmoratus, GenBank accession no. DQ525602).
The evolutionary relationship among alpha (α), mu (μ), pi (π), sigma (σ), rho (ρ), and theta (θ) class GSTs from marine organisms proposed by Blanchette et al. (2007). In this proposal the π/σ intermediate split off early into π and σ. Then μ split off from α/μ/ρ/θ intermediate followed by α and then ρ/θ split off into ρ and θ. The most prominent difference in this proposal from the other three proposals is that θ is located at the end rather than at the beginning of the evolutionary track.
A new classification protocol, which incorporates the characteristics of the aforementioned intermediates, should include the functional characteristics, such as ball-and-socket style, of the GST isozymes under analysis. Along with the structural attributes, which are explained in the following sections, the functional characteristics will allow for further distinction between similar GSTs. The functional quality is referred to the specialized substrate activity inherent in each GST for a specific type of xenobiotic compound. The specialized substrate activity referred to above revolved around enzymes of each class (alpha, mu, pi, theta, and sigma) and the unique enzymatic activity of each class has been said to have a particular xenobiotic compound. For instance, class alpha GSTs are said to be highly active with cumene hydroperoxide, while class mu GSTs show highest activity with epoxides such as trans-stilbene oxide. Finally, class pi GSTs are said to be hyperactive with ethacrynic acid ([2,3-di-chloro-4-(2-methylenebutryl)-phenoxy] acetic acid) (Mannervik and Danielson, 1988; Blanchette and Singh, 2007). Therefore, a new classification criterion has been proposed in this article based on the functional characteristics (ball-and-socket) and substrate specificities of GSTs from marine organisms (Tables 1 and 4). In this classification, BS (ball-and-socket)-I, BS-II, BS-III, and BS-IV show four classes of marine GSTs whose structures possess ball-and-socket style interfaces, and that catalyze alpha/mu, alpha/pi, mu/pi, and alpha/mu/pi class-specific substrates, repectively. The problem with the so-called “characterizing substrates” and their application to the classification of GSTs is that there are many examples of crossover activity. Crossover activity can be defined as a GST of one class having activity with a substrate that has been designated for another class. A more appropriate application of the characterizing substrate designations may be a modification of the method proposed in the early days of research into the functionality of the glutathione transferases (Boyland and Chasseaud, 1969), which recommended that GSTs should be classified by the general substrate class with which they are most active. For example, GSTs would be named glutathione aryl transferase, epoxide transferase, alkyl transferase, etc. The aforementioned method of GST nomenclature was deemed not applicable when it was discovered that GSTs of varying classes may contain overlapping activities. Inclusion of the Boyland and Chasseaud (1969) classification method into the currently used classification method, which refers mainly to GST structural attribute, will allow a human GST from the pi class, showing hyperactivity with aryl compounds, to be termed as a human-aryl-glutathione S-transferase (h-aryl-GST-pi). The functional aspects of each GST would be the easiest characteristic to determine and would provide potential users of the enzyme a more informed choice for experimentation. For example, if a researcher was in need of an enzyme for use in a biomarker/detoxification role for a study at a polluted site and that site is contaminated by an epoxide xenobiotic compound, the obvious choice of GSTs to act in a biomarker role for the aforementioned site would be a glutathione epoxide transferase. In general, to those who use marine GSTs in a biomarker role the main concern is the functionality of the enzyme, and it is this biomarker/detoxification role that dominates the utility of marine GSTs. Therefore, whether a GST is from the pi or mu class makes little difference. As long as the GST is active with the xenobiotic substrate of choice, the ultimate goal is accomplished. An in depth structural, catalytic and sequencing analyses are still extremely important for a better understanding of the biophysical characterization of the marine glutathione transferases. However, a classification scheme based on these aforementioned techniques may be a time-consuming prospect. Therefore, addition of a functional aspect to the GST nomenclature may be a great leap forward in the use of the GSTs as a biomarker/detoxification tool.
Characterized Marine GSTs
The river of knowledge on marine GSTs does not run very deep. In general, the marine GSTs have been the focus of many studies involving environmental pollutants such as polychlorinated biphenyls (PCBs), dichlorodiphenyltrichloroethane (DDT), polycyclic aromatic hydrocarbons (PAHs), etc. Involved in these detoxification /biomarker studies, characterization of GSTs customarily provides a basic foundation of knowledge about the GST. Although the characteristics resulting from these studies can provide a general picture of marine GSTs under investigation, these rarely provide the complete story. In fact, it is not uncommon to find a group of studies focusing on a GST from a single marine organism and each focusing on a different set of characteristics of a given GST.
Unlike the marine GSTs, the group of nonaquatic GSTs has at least one member of each of the major classes fully characterized. This makes classification of a nonaquatic GST with only a small number of attributes known much easier. Unfortunately the group of marine GSTs has a limited number of enzymes fully characterized (Figures 2, 3, 4, 5, 6 and 7, Tables 1 and 2), which makes classification of a partially characterized GST extremely difficult. A fully characterized marine GST is defined here as one that can be placed into a class by either full amino acid sequencing or positive immunological cross reactivity with the same class of GSTs. While many other classification approaches can be employed in the characterization of GSTs, the other techniques, such as substrate/inhibitor enzymatic activity, are known to yield highly variable results.
Among the group of marine GSTs studied, thus far, the majority are from highly mobile fish species such as that of the sea bass, Dicentrarchus labrax (Angelucci et al., 2000). The characteristics of the sea bass GSTs are typical of most research studies. D. labrax contains multiple forms of GST (two isozymes) having isoelectric points of 6.7 and 8.2. The major isozyme (75% of total GST purified) is a homodimer with subunit molecular mass of 26.5 kDa and a pI of 6.7. The basic enzyme (25% of total GST purified) is also a homodimer with smaller subunit molecular mass of 23.5 kDa and a pI of 8.2. The immunological properties of D. labrax GSTs yield some interesting results. Immunoblotting analysis indicates that the D. labrax isozymes have no cross reactivity with each other and therefore have different immunological determinants. Isozyme 6.7 displayed negative cross reactivity with various GSTs from mammalian, amphibian, and bacterial sources. The isozyme 8.2, on the other hand, showed positive cross reactivity with GST 2.2 from the rat alpha class (Angelucci et al., 2000).
The two other characteristics elucidated in the D. labrax investigation yielded some interesting results. Because the N-terminal portions of both isozymes were blocked, sequencing was performed on trypsin-digested segments of the sea bass GSTs. Sequencing of two small segments of D. labrax isozyme 6.7 was found to be an 85% match with that of plaice liver GST-A (Figure 7). The plaice liver GST-A was ultimately determined to be the theta class (Leaver et al., 1993). In addition, a small segment (∼30 amino acids) of the basic sea bass GST was found to be approximately 50% homologous with rabbit alpha class GST II. Characterizing substrate enzymatic activity showed similar results to that of the segment sequencing. Both the acidic and basic D. labrax isozymes showed increased activity with trans-non-2-enal. Trans-non-2-enal is a substrate that has been known to show increased enzymatic activity with GSTs from both the alpha and mu classes (Brophy et al., 1989). Interestingly, the substrates that carried the second highest activity differed for each of the sea bass GSTs. D. labrax GST 8.2 also had high activity with ethacrynic acid, which shows increased activity with pi class GSTs. In contrast, sea bass GST 6.7 exhibited high enzymatic activity with the alpha class substrate Δ5-androstene-3,17-dione. These results present an example of the confusion that can result when conflicting data are the outcome of partial characterization. Sequence data that match with one GST class and substrate characterization results sometimes indicate that the same GST may belong to another class. This observation is common to most partial and even some full characterization experiments involving GSTs.
Another example of fish GSTs is from the marine flat fish plaice (Pleuronectes platessa). Initial identification was made with the near neutral homodimeric GST (pI 6.7) with subunit mass of 23.5 kDa. The major GST comprised about 80% of the plaice liver cytosolic GST activity with CDNB (George and Young, 1988) (Figure 7; Tables 1 and 2). Along with the major subunit, other cytosolic GSTs were also separated. The other homodimeric GST carrying approximately 20% of the remaining enzymatic activity has subunit molecular mass of 26.5 kD and a pI of 5.7. The remaining P. platessa GST activity, less than 1% of the total CDNB activity, is divided between a homodimeric enzyme with subunit molecular mass of 24.5 kDa and a heterodimer with subunit masses of 24.5 kDa and 26.5 kDa. Enzymatic activity with the GST characterizing substrates indicates that the major GST isozyme from P. platessa GST shows high levels of activity with the alpha/mu class substrate p-nitrobenzyl chloride. The homodimer comprising 26.5-kDa subunits showed increased activity with the mu class substrate DCNB.
Further characterization of the major plaice isozyme determined that the cytosolic GST was most closely related to the plant, insect, and mammalian theta classes of GSTs (Leaver et al., 1993). Two Plaice GST cDNA clones were found to encode for 224-amino-acid polypeptides (Figure 7) of 25,723 Da and 25,603 Da, respectively. In an immunochemical experiment with a polyclonal antibody raised in rabbit against the major P. platessa isozyme, it was found that the highest concentration of GST-A is in the liver and the bile duct (Gallagher et al., 1995). Further screening of the plaice genomic DNA library with GST cDNA resulted in the discovery of two overlapping clones (Leaver and George, 1996). The clones were genes coding for GST-A, A1, and A2. Analysis of the first two gene products indicated that each had activity with a different group of compounds and that the sequences of the three GSTs share 60% to 70% homology. The GST-A promoter was shown to be induced by xenobiotic compounds similar to those which induce the alpha class GSTs. The GST-A promoter was noted to respond to oxidative stress in a manner similar to that of the aryl response elements (AREs) in mammals (Leaver and George, 1996). The AREs have been noted to play a role in the detoxification of xenobiotic compounds. The A1 promoter, on the other hand, was shown to contain estrogen and peroxisomal proliferators response elements (EREs and PPREs), which are known to play a role in female sexual maturity and in fatty acid metabolism (Issemann and Green, 1990; Martinez et al., 1991).
Among the well characterized marine GSTs, the sigma class GST of the squid (Ommastrephes sloani pacificus-arrow/Sloane’s squid) digestive gland is one of the most interesting. The O. pacificus GST became quite significant when characterization studies indicated that it might be the prototype for a new class of GSTs. It was later designated as the sigma class, and its sequence serves as the archetype for the new class (Figure 6). The digestive gland GST was chosen for analysis in O. pacificus owing to the gland’s functional similarities to that of the liver in vertebrate species. The purified O. pacificus GST displayed subunit characteristics typical of that of most aquatic and nonaquatic GSTs. The squid digestive gland GST exhibited both major and minor subunits with molecular masses near 25 kDa (Tomarev et al., 1993). The enzymatic activity of O. pacificus GST was among its most unique characteristics, comprising one of the highest GST activities ever recorded with the general CDNB substrate (Ji et al., 1995). Similar results were obtained from the enzymatic testing of digestive gland GSTs from another squid, Loligo vulgaris (Harris et al., 1991). The sigma class hyperactivity with CDNB is possibly due to the extreme open nature of the H-site, as discussed earlier. The open nature of the aforementioned active site is also proposed to hinder the catalytic addition of GSH to enones and/or epoxides (Armstrong et al., 1996). The reduced activity with enones and epoxides can be attributed to the lack of an electrophilic residue at amino acid position 106. In sigma class GSTs the Phe residing at position 106 corresponds to a Tyr residue at position 115 in GSTs, with increased enone/epoxide activity such as in the mu class (Armstrong et al., 1996).
The amino acid sequence of O. pacificus digestive gland GST shared little homology with GSTs from other major classes. It was only 32% to 34% identical to the GSTs from porcine pi class. Similar comparison results were obtained with vertebrate GSTs from the alpha class (29% to 32%). The squid GST was only 19% to 23% homologous to the GSTs of selected vertebrates and insects, which were more closely related to the GSTs of the mu or theta class (Tomarev et al., 1993). Comparison of the sequence data indicates the unique nature of the GSTs from the sigma class. Comparison of the intron-exon structure of the 25-kb squid GST gene yielded results similar to that of the sequence comparison, with the pi class gene most similar in structure. Finally, the tissue distribution of the squid GST mRNA was examined. Northern blot hybridization analysis was used to determine the sigma class distribution in differing tissue samples from O. pacificus (Tomarev et al., 1993). As was the case with P. platessa, the homodimeric GST found in highest concentration in the detoxification organ (liver-P. platessa; digestive gland-O. pacificus) was found to be tissue nonspecific (Tomarev et al., 1993; Gallagher et al., 1995).
There are two distinct groups of marine organisms in relation to GSTs. These groups of marine creatures are distinguished by their relative mobility within the ocean environment. The mobile nature of marine organisms ultimately dictates whether an organism must live within a localized pollution pool or if it has the ability to move out of the pool, thus lessening exposure to harmful toxins. So far, we have described the GST characteristics of only one of the above classes, those of the “highly mobile fish species.” Sedentary organisms consisting of various types of shellfish, on the other hand, must endure, adapt and survive the rigors of a polluted environment. The majority of GSTs from this second group of organisms listed above are investigated in regards to their ability to lessen the effects of any toxic xenobiotic compounds found in their immediate environment. Their capacity to detoxify xenobiotic compounds is not discussed in this review. We will, however, list characteristics of a few characterized GSTs from sedentary marine organisms, such as blue mussel (Mytilus edulis) and quahog (Mercinaria mercinaria (Figure 4; Tables 1, 2, 4 and Table 5; Fitzpatrick and Sheehan, 1993; Fitzpatrick et al., 1995a; Yang et al., 2004).
M. edulis is a bivalve mollusc that can be found over a wide-ranging area comprising the majority of the coastal waters of the northeastern and midatlantic United States and Western Europe. Well known for its role in pollution detection, blue mussels were a logical choice for a GST characterization (U.S. EPA, 1996). The inducibility of the GSTs prompted these investigations, which targeted their use as a biomarker for pollution detection. The initial characterization studies focused on the purification of mussel GSTs through affinity and fast protein liquid chromatographic analysis (FPLC), followed by determination of enzymatic activity with CDNB and subunit molecular mass (Fitzpatrick and Sheehan, 1993). The initial tissues chosen for GST purification are centered on the major feeding and/or suspected detoxification organs such as the gills and digestive gland. The most intense GST specific activity (3.93 μmol min−1mg−1) was found in the gill, and the activity in the digestive gland was approximately one fourth that of the gill. The molecular mass estimation by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) analysis indicated subunit masses of 24.5 kDa and 26.5 kDa.
Further biochemical characterization of M. edulis gill GSTs included isozyme separation/kinetic characterization, immunoblotting, full amino acid sequencing, and substrate specificity analysis. A kinetic analysis of the purified blue mussel isozymes indicated that the most active gill isozyme, the homodimeric GST1-24.5 kDa, resulted in a specific activity of 10.05 μmol min−1mg−1. The other isozyme (26.5 kDa) is also a homodimer but contains none of the aforementioned CDNB activity and is thus referred to as a GSH-binding protein (Fitzpatrick et al., 1995a). Immunoblotting analysis for M. edulis GST1 indicated a positive reaction with the GST pi antisera from rat. The blue mussel GSH-binding protein, on the other hand, exhibited no such reaction with antisera from either the mu or the alpha GST classes. N-terminal amino acid sequencing analysis offered rather interesting results in light of the immunoblotting data. Degradation was employed to carry out N-terminal sequencing up to approximately 20 residues. The sequence of mussel GST1 shares 35%, 55%, and 60% homology with the π class GSTs from salmonid, frog, and mouse, respectively (Dominey et al., 1991; DiIlio et al., 1992; Hayes et al., 1991). Surprisingly, the M. edulis GST1 also shared a 50% sequence homology with the N-terminal sequences from other cephalopod GSTs such as the squid sigma class (Tomarev et al., 1993). Comparison of the GST1 to consensus sequences from the alpha and mu classes indicated a sequence homology of 35% and 50%, respectively. The blue mussel GST1 was determined to most closely resemble the GSTs of the pi class, even though only a limited number of homologus pi class sequences were available for comparison. A comparison to representative mu class sequences resulted in a similar homology. This is yet another indication of the confusion that classification of GSTs can cause without access to the full characterization data on a particular GST family. Full sequence results ultimately did reveal the blue mussel GST1 to be of the pi class, a fact that is much less important than the overall catalytic abilities of the enzyme (Luedeking and Koehler, 2000; Yang et al., 2004).
The N-terminal sequencing results for the mussel GSH-binding protein offered further confusion in that the sequence alignment indicates similar results for all classes of marine GSTs tested. The highest homology (61%) was to that of the mu class consensus sequence. Even though the homology was stronger to the mu class for the binding protein than the GST1 was to the pi class, it was obviously not classified as a mu class GST because it lacked the enzymatic activity. It only shows structural similarity to GSTs in general. The substrate specificity of M. edulis GST1 is the highest for ethacrynic acid, with a specific activity of 1.61 μmol min−1mg−1. However, the classification based on substrate specificity and the N-terminal sequencing is not consistent with inhibitor analysis results. In general, inhibitory concentrations yielding 50% activity with CDNB as substrate (IC50) for pi class GSTs are known to be approximately 4 μM for both ethacrynic acid and hematin (Mannervik et al., 1985). IC50 concentrations for bromosulphophthalein of 2 to 200 μM and 0.05 to 2 μM for hematin are typical for alpha class GSTs (Mannervik et al., 1985). Blue mussel GST1 showed an IC50 pattern in line with that of an alpha class GST, adding confusion to its classification.
Recently, in addition to mussels, hardshell clams are being popularly utilized to investigate the important detoxification roles of GSTs in the marine environment. Quahog (Mercinaria mercinaria) is one kind of hardshell clam that inhabits the coastal waters in the eastern shores of North America. To study marine environment in terms of special types of living populations and exposure to pollution, Blanchette and Singh (1999) purified and characterized GST from quahog. The results showed that GST from quahog total tissue had a specific activity of 38.0 μmol min−1 mg−1, which was higher than that of GSTs from blue mussel (M. edulis gill, 10.05 μmol min−1 mg−1), Mediterranean mussel (M. galloprovincialis digestive gland, 5.8 μmol min−1 mg−1), and oyster (Crassostrea virginica total tissue, 4.2 μmol min−1 mg−1) (Fitzpatrick et al., 1995a, b; Ge and Singh, 1996), but was lower than that of GSTs from Asiatic clam (C. fluminea gill, 206.9 μmol min−1 mg−1), striate beach clam (Atactodea striata hepatopancreas, 108.9 μmol min−1 mg−1), and seashell (Asaphis dichotoma liver, 283.7 μmol min−1 mg−1) (Vidal and Narbonne, 2000; Yang et al., 2002, 2003). The molecular mass estimation by SDS-PAGE analysis showed that quahog GST was a dimer that had multiple forms, with subunit molecular masses of 22, 24, 25, and 27 kDa. Isoelectric focusing analysis resulted in pI values for three isozymes of 5.1, 4.9, and 4.6. In addition, kinetic analysis showed V max and K m values of 48.0 μmol min−1 mg−1 and 0.24 mM, respectively. Compared with mussels, oyster, and clams, quahog GST had larger V max than Mediterranean mussel (M. galloprovincialis total tissue, 0.0222 μmol min−1 mg−1), oyster (C. virginica total tissue, 6.9 μmol min−1 mg−1), Asiatic clam (C. fluminea gill, 0.0772 μmol min−1 mg−1), and seashell (A. dichotoma liver, 33 μmol min−1 mg−1), and smaller K m than blue mussel (M. edulis gill, 0.5 mM), Mediterranean mussel (M. galloprovincialis total tissue, 0.702 mM), oyster (C. virginica total tissue, 0.75 mM), Asiatic clam (C. fluminea total tissue, 0.46 mM), and striate beach clam (A. striata hepatopancreas, 0.43 mM). Moreover, inhibition analysis showed that purified quahog GST was inhibited by both open-chain and closed-chain tetrapyrroles (Blanchette and Singh, 1999).
The GSTs listed in the preceding text comprise the short list of GSTs from aquatic organisms, which can be considered fully characterized. The list of partially characterized GSTs is much more extensive than that of the fully characterized. Tables 4 and 5 list data for partially characterized GSTs from aquatic organisms (Ramage and Nimmo, 1984; Keeran and Lee 1987; Boryslawskyj et al., 1988; Nies et al., 1991; Collier et al., 1992; Makela et al., 1992; Pangrekar and Sikka, 1992; Chetty and Indira, 1995; Forlin et al. 1995; Gallagher et al., 1996b; Ge and Singh, 1996; Martinez-Lara et al., 1996; Lenartova et al. 1997; James et al., 1998; Perez-Lopez et al., 1998; Blanchette and Singh, 1999; El-Demerdash and Elagamy 1999; Kaaya et al., 1999; Angelucci et al., 2000; Gallagher et al., 2000; Melgar-Riol et al., 2001; Blanchette and Singh, 2002; Novoa-Valinas et al., 2002; Yang et al., 2002, 2003). The results from partially characterized GSTs, like those considered fully characterized in the preceding text, only add confusion to the classification attempts. Taken alone, amino acid sequence data, substrate specificities, or gene structure data must be looked upon with caution. Limited results such as those outlined in the preceding text are indicative of an inadequate classification process. Therefore, the classification process and a suggested improvement need to be examined.
Perspectives
The family of GSTs from aquatic organisms has not been well characterized to date. The limited number of in-depth investigations into the structure and characterization of the marine GSTs poses problems to categorize the marine GSTs. Even when a considerable portion of the GST characteristics is known, it is still hard to assign a classification of marine GSTs because confusing and contradictory data are often obtained using the existing classification standards. Conflicting class characteristics can lead to substantial wastes of time and materials in what could be called a search for the “true” GST class. Therefore, it is important that a suitable classification system for marine GSTs be developed. In recent years, more and more studies on marine GSTs focus on the biomarker traits of the enzyme. Therefore, a new classification system of marine GSTs should be established utilizing knowledge of the detoxification and biomarker roles of GSTs in the marine environment.
References
Angelucci S, Sacchetta P, Moio P, Melino S, Petruzzelli R, Gervasi P, Di Ilio C (2000) Purification and characterization of glutathione transferases from the sea bass (Dicentrachus labrax) liver. Arch Biochem Biophys 373, 435–441
Armstrong RN (1991) Glutathione S-transferases: reaction mechanism, structure, and function. Chem Res Toxicol 4, 131–140
Armstrong RN (1997) Structure, catalytic mechanism, and evolution of the glutathione transferases. Chem Res Toxicol 10, 2–18
Armstrong RN, Chen J, Johnson WW, Parsons J, Ji X, Gilliland GL, Tomarev SI, Piatigorsky J (1996) “Structure, mechanism, and evolution of class Mu and Sigma glutathione-S-transferases”. In: Glutathione S-transferases: Structure, Function and Clinical Implications, Vermeulen NPE, Mulder, GJ, Nieuwenhuyse H, Peters WHM, van Bladeren PJ, eds. (London: Taylor and Francis) pp 13–22
Blanchette BN, Singh, BR (1999) Purification and characterization of the glutathione-S-transferases from the Northern quahog Mercinaria mercinaria. Mar Biotechnol 1, 74–80
Blanchette BN, Singh BR (2002) Isolation and characterization of glutathione-S-transferase isozyme Q3 from the Northern quahog, Mercinaria mercinaria. J Protein Chem 21, 151–159
Blanchette BN, Singh BR (2007) A high pressure liquid chromatography-based assay for glutathione-S-transferase class distinction assay. J Biochem Biophys Meth 70, 761–765
Board PG, Baker RT, Chelvanayagam G, Jermin, LS (1997) Zeta, a novel class of glutathione transferases in a range of species from plants to humans. Biochem J 328, 929–935
Board PG, Coggan M, Chelvanayagam G, Easteal S, Jermin LS, Schulte GK, Danley DE, Hoth LR, Griffor MC, Kamath AV, Rosner MH, Chrunyk BA, Perregaux DE, Gabel CA, Geoghegan KF, Pandit J (2000) Identification, characterization, and crystal structure of the omega class glutathione transferases. J Biol Chem 275, 24789–24806
Boryslawskyj M, Garrood AC, Pearson JT, Woodhead D (1988) Evaluation of glutathione S-transferase activity as a stress response to organochlorine compounds, in the freshwater mussel, Sphaerium corneum. Mar Environ Res 24, 101–104
Boyland E, Chasseaud, LF (1969) The role of glutathione and glutathione S-transferases in mercapturic acid biosynthesis. Adv Enzymol Rel Areas Mol Biol 32, 173–179
Brophy PM, Southan C, Barrett J (1989) Glutathione transferases in the tapeworm Moniezia expansa. Biochem J 262, 939–946
Buetler TM, Eaton DL (1992) Glutathione S-transferases: amino acid sequence comparison, classification and phylogenic relationship. J Environ Sci Health C Environ Carcino Ecotoxicol Rev 10, 181–203
Caccuri AM, Antonini G, Nicotra M, Battistoni A, Bello ML, Board PG, Parker MW, Ricci G (1997) Catalytic mechanism and role of hydroxyl residues in the active site of theta class glutathione S-transferases. J Biol Chem 272, 29681–29686
Cameron AD, Sinning I, L’Hermite G, Olin B, Board P, Mannervik B, Jones TA (1995) Structural analysis of human alpha class glutathione transferase in the apo-form and in complexes with ethacrynic acid and its glutathione conjugate. Structure 3, 717–727
Chetty AN, Indira K (1995) Free radical toxicity in a freshwater bivalve (Lamellidens marginalis) under ambient ammonia stress. J Environ Biol 16, 137–142
Collier TK, Singh SV, Awasthi YC, Varanasi U (1992) Hepatic xenobiotic metabolizing enzymes in tow species of benthic fish showing different prevalences of contaminant-associated liver neoplasms. Toxicol Appl Pharmacol 113, 319–324
Contreras-Vergara CA, Harris-Valle C, Sotelo-Mundo RR, Yepiz-Plascencia G (2004) A mu-class glutathione S-transferase from the marine shrimp Litopenaeus vannamei: molecular cloning and active-site structural modeling. Biochem Mol Toxicol 18, 245–252
Dean JV, Devarenne TP, Lee IS, Orlofsky LE (1995) Properties of maize glutathione S-transferase that conjugates coumaric acid and other phenylpropanoids. Plant Physiol 108, 985–994
DiIlio C, Aceto A, Bucciarelli T, Dragani B, Angelucci S, Miranda M, Poma A, Amicarelli F, Barra D, Federici G (1992) Glutathione transferase isoenzymes from Bufo bufo embryos at an early development stage. Biochem J 283, 217–222
Ding Y, Ortelli F, Rossiter LC, Hemingway J, Ranson H (2003) The Anopheles gambiae glutathione transferase supergene family: annotation, phylogeny and expression profiles. BMC Genomics 4, 35–45
Dirr HW, Wallace LA (1999) Role of the C-terminal helix 9 in the stability and ligandin function of class alpha glutathione transferase A1-1. Biochemistry 38, 15631–15640
Dirr H, Reinemer P, Huber R (1994) X-ray crystal structures of cytosolic glutathione S-transferase: implications for protein architecture, substrate recognition and catalytic function. Eur J Biochem 220, 645–661
Dixon DP, Lapthom A, Edwards R (2002) Plant glutathione transferases. Genome Biol 3, reviews 3004.1–3004.10
Doi AM, Pham RT, Hughes EM, Barber DS, Gallagher EP (2004) Molecular cloning and characterization of a glutathione S-transferase from largemouth bass (Micropterus salmoides) liver that is involved in the detoxification of 4-hydroxynonenal. Biochem Pharmacol 67, 2129–2139
Dominey RJ, Nimmo IA, Cronshaw AD, Hayes JD (1991) The major glutathione S-transferase in salmonid fish livers is homologus to the mammalian pi-class GST. Comp Biochem Physiol 100B, 93–98
Doyen P, Vasseur P, Rodiu F (2005) cDNA cloning and expression pattern of pi-class glutathione S-transferase in the freshwater bivalves Unio tumidus and Corbicula fluminea. Comp Biochem Physiol C 140, 300–308
Droog F (1997) Plant glutathione S-transferases, a tale of theta and tau. J Plant Growth Regul 16, 95–107
Droog FNJ, Hooykaas PJJ, vander Zaal EJ (1995) 2,4-Dichlorophenoxyacetic acid and related compounds inhibit two auxin-regulated type III tobacco glutathione S-transferases. Plant Physiol 107, 1139–1146
Edward R, Dixon DP, Walbot V (2000) Plant glutathione S-transferases: enzymes with multiple functions in sickness and health. Trends Plant Sci 5, 193–198
El-Demerdash FM, Elagamy EI (1999) Biologica effects in Tilapia nilotica fish as indicators of population by mercury. Int J Environ Health Res 9, 173–186
Fernandes CL, Chavan SJ, Dong JH, Bornmann WG, Polsky B, Chisari FV, Montali JA, Schmidt Jr DE, Prochaska HJ (1996) “Regulation of glutathione-S-transferases: clues from a worm, a virus, and a mouse with hepatitis”. In: Glutathione S-transferases: Structure, Function and Clinical Implications, Vermeulen NPE, Mulder GJ, Nieuwenhuyse H, Peters WHM, van Bladeren PJ, eds. (London: Taylor and Francis), pp 97–109
Fitzpatrick PJ, Sheehan D (1993) Separation of multiple forms of glutathione S-transferase from the blue mussel, Mytilus edulis. Xenobiotica 23, 851–861
Fitzpatrick PJ, Krag TOB, Hojrup P, Sheehan D (1995a) Characterization of a glutathione S-transferase and a related glutathione-binding protein from gill of the blue mussel, Mytilus edulis. Biochem J 305, 145–150
Fitzpatrick PJ, Sheehan D, Livingstone DR (1995b) Studies on isoenzymes of glutathione S-transferase in the digestive gland of Mytilus galloprovincialis with exposure to pollution. Mar Environ Res 39, 241–244
Förlin L, Lemaire P, Livingstone DR (1995) Comparative studies of hepatic xenobiotic metabolizing and antioxidant enzymes in different fish species. Mar Environ Res 39, 201–204
Fournier D, Bride JM, Poirie M, Berge JB, Plabb FW (1992) Insect glutathione S-transferases. Biochemical characteristics of the major forms from houseflies susceptible and resistant to insecticides. J Biol Chem 267, 1840–1845
Frear DS, Swanson HR (1970) Biosynthesis of S-(4-ethylamino-6-isopropylamino-2-s-triazine) glutathione: partial purification and properties of glutathione-S-transferase from corn. Phytochemistry 9, 2123–2132
Gallagher EP, Sheehy KM (2000) Altered glutathione S-transferase catalytic activities in female brown bullheads from a contaminated central Florida lake. Mar Environ Res 50, 399–403
Gallagher T, Strom A, George S (1995) The cellular distribution of glutathione S-transferase in plaice: an immunocytochemical study. Mar Environ Res 39, 352–353
Gallagher EP, Stapleton PL, Collier TK, Varanasi U, Eaton DL (1996a) Biochemical and molecular characterization of glutathione S-transferase expression in English sole and starry flounder. Mar Environ Res 42, 26–36
Gallagher EP, Stapleton PL, Slone DH, Schlenk D, Eaton DL (1996b) Channel catfish glutathione S-transferase isozyme activity toward (±)-anti-benzo[α]pyrene-trans-7,8-dihydrodiol-9,10-epoxide. Aquat Toxiol 34, 135–150
Gallagher EP, Sheehy KM, Lame MW, Segall HJ (2000) In vitro kinetics of hepatic glutathione S-transferase conjugation in largemouth bass and brown bullheads. Environ Toxicol Chem 19, 319–326
Garcia-Saez I, Parraga A, Phillips MF, Mantle TJ, Coll M (1994) Molecular structure at 1.8 Å of mouse liver class Pi glutathione S-transferase complexed with S-(p-nitrobenzyl)-glutathione and other inhibitors. J Mol Biol 237, 298–314
Ge J, Singh BR (1996) Physicochemical characterization of glutathione S-transferase purified from oyster, Crassostrea virginica. J Mar Biotechnol 4, 150–154
George SG, Young P (1988) Purification and properties of place liver cytosolic glutathione S-transferases. Mar Environ Res 24, 93–96
Gustafsson A, Etahadieh M, Jemth P, Mannervik B (1999) The C-terminal region of human glutathione transferase A1-1 affects the rate of glutathione binding and the ionization of the active site Tyr9. Biochemistry 38, 16268–16275
Habig WH, Pabst MJ, Jakoby WB (1974) Glutathione S-transferases the first enzymatic step in mercapturic acid formation. J Biol Chem 249, 7130–7139
Habig WH, Pabst MJ, Jakoby WB (1976) Glutathione S-transferase AA from rat liver. Arch Biochem Biophys 175, 710–715
Harris J, Coles B, Meyer D, Ketterer B (1991) The isolation and characterization of the major glutathione S-transferase from the squid Loligo vulgaris. Comp Biochem Physiol 98B, 511–515
Hayes JD, McLellan LI (1999) Glutathione and glutathione-dependent enzymes represent a co-ordinately regulated defence against oxidative stress. Free Radic Res 31, 273–300
Hayes JD, Pulford DJ (1995) The glutathione S-transferase supergene family: regulation of GST and the contribution of the isoenzymes to cancer chemprotection and drug resistance. Crit Rev Biochem Mol Biol 30, 445–600
Hayes JD, Kerr LA, Peacock SD, Cronshaw AD, McLellan LI (1991) Hepatic glutathione S-transferases in mice fed on a diet containing the anticarcinogenic antioxidant butylated hydroxyanisol. Isolation of mouse glutathione S-transferase heterodimes by gradient elution of the glutathione sepharose affinity matrix. Biochem J 277, 501–512
Hayes JD, Flanagan JU, Jowsey IR (2005) Glutathione transferases. Annu Rev Pharmacol Toxicol 45, 51–88
Hemingway J (2000) The molecular basis of two contrasting metabolic mechanisms of insecticide resistance. Insect Biochem Mol Biol 30, 1009–1015
Hoarau P, Garello G, Gnassia-Barelli M, Remeo M, Girard JP (2002) Purification and partial characterization of seven glutathione S-transferase isoforms from the clam Ruditapes decussates. Eur J Biochem 269, 4359–4366
Hoarau P, Damiens G, Romeo M, Gnassia-Barelli M, Bebianno MJ (2006) Cloning and expression of a GST-pi gene in Mytilus galloprovincialis. Attempt to use the GST-pi transcript as a biomarker of pollution. Comp Biochem Physiol Part C 143, 196–203
Issemann I, Green S (1990) Activation of a member of the steroid hormone superfamily by peroxisome proliferators. Nature 347, 645–650
Ivanetich KM, Goold RD, Sikakana CN (1990) Explanation of the non-hyperbolic kinetics of the glutathione S-transferases by the simplest steady-state random sequential Bi Bi Mechanism. Biochem Pharmacol 39, 1999–2004
Jakobson I, Warholm M and Mannervik B (1979) The binding of substrates and a product of the enzymatic reaction to glutathione S-transferase A. J Biol Chem 254, 7085–7089
Jakoby WB, Habig WH, Keen JH, Ketley JN, Pabst MJ (1976) Glutathione S-transferases: catalytic aspects. In: Glutathione: Metabolism and Function, Arias IM, Jakoby WB, eds. (New York: Raven Press), p 189
James MO, Sikazwe DN, Gadagbui BKM (1998) Isolation of a pi class glutathione S-transferase (GST) from catfish intestinal mucosa. Mar Environ Res 46, 57–60
Ji X, Zhang P, Armstrong RN, Gilliland GL (1992) The three-dimensional structure of a glutathione S-transferase from the mu class gene. Structural analysis of the binary complex of isoenzyme 3-3 and glutathione at 2.2 Å resolution. Biochemistry 31, 10169–10184
Ji X, von Rosenvinge EC, Johnson WW, Tomarev SI, Piatigorsky J, Armstrong RN, Gilliland GL (1995) Three-dimensional structure, catalytic properties and evolution of a sigma class glutathione transferase from squid, a progenitor of the lens S-crystallins of cephalopods. Biochemistry 34, 5317–5328
Johnson WW, Liu S, Ji X, Gilliland GL, Armstrong RN (1993) Tyrosine 115 participates both in chemical and physical steps of the catalytic mechanism of a glutathione S-transferase. J Biol Chem 268, 11508–11511
Jowsey IR, Thomson RE, Orton TC, Elcombe CR, Hayes JD (2003) Biochemical and genetic characterization of a murine class kappa glutathione S-transferase. Biochem J 373, 559–569
Kaaya A, Najimi S, Ribera D, Narbonne JF, Moukrim A (1999) Characterization of glutathione S-transferases (GST) activities in Perna perna and Mytilus galloprovincialis used as a biomarker of pollution in the Agadir Marine Bay (South of Morocco). Bull Environ Contam Toxicol 62, 623–629
Kanaoka Y, Ago H, Inagaki E, Nanayama T, Miyano M, Kikuno R, Fujii Y, Eguchi N, Toh H, Urade Y, Hayaishi O (1997) Cloning and crystal structure of hematopoietic prostaglandin D synthase. Cell 90, 1085–1095
Keeran WS, Lee RF (1987) The purification and characterization of glutathione S-transferase from the hepatopancreas of the blue crab, Callinectes sapidus. Arch Biochem Biophys 255, 233–243
Konishi T, Kato K, Araki T, Shiraki K, Takaqi M, Tamaru Y (2005a) Molecular cloning and characterization of alpha-class glutathione S-transferase genes from the hepatopancreas of red sea bream, Pagrus major. Comp Biochem Physiol C 140, 309–320
Konishi T, Kato K, Araki T, Shiraki K, Takagi M, Tamaru Y (2005b) A new class of glutathione S-transferase from the hepatopancreas of the red sea bream Pagrus major. Biochem J 388, 299–307
Kudo H, Ueda H, Mochida K, Adachi S, Hara A, Nagasawa H, Doi Y, Fujimoto S, Yamauchi K (1999) Salmonid olfactory system-specific protein (N24) exhibits glutathione S-transferase pi-like structure. J Neurochem 72, 1344–1352
Labrou NE, Mello LV, Clonis YD (2001) Functional and structural roles of the glutathione-binding residues in maize (Zea mays) glutathione S-transferase I. Biochem J 358, 101–110
Labrou NE, Karavangeli M, Tsaftaris A, Clonis YD (2005) Kinetic analysis of maize glutathione S-transferase I catalyzing the detoxification from chloroacetanilide herbicides. Planta 222, 91–97
Ladner JE, Parsons JF, Rife CL, Gilliland GL, Armstrong RN (2004) Parallel evolutionary pathways of glutathione transfearses: structure and mechanism of the mitochondrial class kappa enzyme rGSTK1-1. Biochemistry 43, 352–361
Leaver MJ, George SG (1995) Structure of the plaice glutathione S-transferase A gene. Mar Environ Res 39, 33–37
Leaver MJ, George SG (1996) The repeated glutathione S-transferase genes from a marine fish, the plaice (Pleuronectes platessa). Mar Environ Res 42, 19–23
Leaver MJ, Scott K, George SG (1993) Cloning and characterization of the major hepatic glutathione S-transferases from a marine teleost flatfish, the plaice (Pleuronectes platessa), with structural similarities to plant, insect and mammalian theta class isoenzymes. Biochem J 292, 189–195
Leaver MJ, Wright J, George SG (1997) Structure and expression of a cluster of glutathione S-transferase genes from a marine fish, the plaice (Pleuronectes platessa). Biochem J 321, 405–412
Lee Y, Chang S, Jung S, Kweon H, Lee J (2005) Cloning and expression of alpha class glutathione S-transferase gene from the small hermaphroditic fish Rivulus marmoratus (Cyprinodontiformes, Rivulidae). Mar Pollut Bull 51, 776–783
Lee YM, Seo JS, Jung SO, Kim IC, Lee JS (2006) Molecular cloning and characterization of theta-class glutathione S-transferase (GST-T) from the hermaphroditic fish Rivulus marmoratus and biochemical comparisons with alpha-class glutathione S-transferase (GST-A). Biochem Biophys Res Commun 346, 1053–1061
Lenartova V, Holovska K, Pedrajas JR, Martínez-Lara E, Peinado J, Lopez-Barea J, Rosival I, Kosuth P (1997) Antioxidant and detoxifying fish enzymes as biomarkers of river pollution. Biomarkers 2, 247-252
Liao W, Liang X, Wang L, Lei L, Han B (2006) Molecular cloning and characterization of alpha-class glutathione S-transferase gene from the liver of silver carp, bighead carp, and other major Chinese freshwater fishes. J Biochem Mol Toxicol 20, 114–126
Lin KS, Chuang NN (1993) Anionic glutathione S-transferases in shrimp eyes. Comp Biochem Physiol 105B, 151–156
Lougarre A, Bride JM, Fournier D (1999) Is the insect glutathione S-transferase I gene family intronless? Insect Mol Biol 8, 141–143
Luedeking A, Koehler A (2004) Regulation of expression of multixenobiotic resistence (MXR) genes by environmental factors in the blue mussel Mytilus edulis. Aquat Toxicol 69, 1–10
Makela TP, Lindstrom-Seppa P, Oikari AOJ (1992) Organochlorine residues and physiological conduction of the freshwater mussel (Anodonta anatina) caged in the river Prelinen, Eastern Finland, receiving Pulp Mill Effluent. Aqua Fennica 22, 49–58
Mannervik B (1985) The isoenzymes of glutathione transferase. Adv Enzymol Relat Areas Mol Biol 57, 357–417
Mannervik B, Danielson UH (1988) Glutathione transferases-transferase and catalytic activity. CRC Crit Rev Biochem 23, 283–337
Mannervik B, Alin P, Guthenberg C, Jensson H, Tahir MK, Warholm M, Jornvall H (1985) Identification of three classes of cytosolic glutathione transferases common to several mammalian species: correlation between structural data and enzymatic properties. Proc Natl Acad Sci USA 82, 7202–7206
Marrs KA (1996) The functions and regulation of glutathione S-transferases in plants. Annu Rev Plant Physiol Plant Mol Biol 47, 127–158
Marrs KA, Alfenito MR, Lloyd AM, Walbot V (1995) A glutathione S-transferase involved in vacuolar transfer encoded by maize gene Bronze-2. Nature 375, 397–400
Martinez E, Givel F, Wahli W (1991) A common ancestor DNA motif for invertebrate and vertebrate hormone response elements. EMBO J 10, 263–268
Martinez-Lara E, Toribio F, Lopez-Barea J, Barcena JA (1996) Glutathione S-transferase isoenzyme patterns in gilthead seabream (Sparus aurata) exposed to environmental contaminants. Comp Biochem Physiol Part C 113, 215–220
Mathews CK, van Holde KE (1990) In Biochemistry, 1st ed. (New York: Benjamin/Cummings), p 711
Melgar-Riol MJ, Novoa-Valinas MC, Garcia-Fernandez MA, Perez-Lopez M (2001) Glutathione S-transferases from rainbow trout live rand freshly isolated hepatocytes: purification and characterization. Comp Biochem Physiol C 128, 227–235
Meyer DJ, Coles B, Pemble SE, Gilmore KS, Fraser GM, Ketterer B (1991) Theta, a new class of glutathione transferases purified from rat and man liver. Biochem J 274, 409–414
Morel F, Rauch C, Petit E, Piton A, Theret N, Coles B, Guillouzo A (2004) The human glutathione transferase kappa: gene and protein characterization and evidence for a peroxisomal localization. J Biol Chem 279, 16246–16253
Morgenstern R, Guthenberg C, DePierre JW (1982) Microsomal glutathione S-transferase: purification, initial characterization and demonstration that it is not identical to cytosolic GSTs A, B and C. Eur J Biochem 128, 243–248
Nam YK, Cho YS, Choi BN, Kim KH, Kim SK, Kim DS (2005) Alteration of antioxidant enzymes at the mRNA level during short-term starvation of rockbream Oplegnathus fasciatus. Fish Sci 71, 1385–1387
Neuefeind T, Huber R, Dasenbrock H, Prade L, Bieseler B (1997a) Crystal structure of glutathione S-transferase-I in complex with lactoylglutathione: evidence for an induced-fit mechanism. J Mol Biol 274, 446–453
Neuefeind T, Huber R, Reinemer P, Knaeblein J, Prade L, Mann K, Bieseler B (1997b) Cloning, sequencing, crystallization and X-ray structure of glutathione S-transferase-III from Zea mays var. mutin: a leading enzyme in detoxification of maize herbicides. J Mol Biol 274, 577–587
Nies E, Almar MM, Hermeneqildo C, Monsalve E, Romero FJ (1991) The activity of glutathione S-transferase in hepatopancreas of Procambarus clarkia: seasonal variations and the influence of environmental pollutants. Comp Biochem Physiol C 100, 65–66
Nishiihira J, Ishibashi T, Sakai M, Tsuda S, Hikichi K (1993) Identification of the hydrophobic ligand binding region in recombinant glutathione S-transferase P and its binding effect on the conformational state of the enzyme. Arch Biochem Biophys 302, 128–133
Novoa-Valinas MC, Perez-Lopez M, Melgar MJ (2002) Comparative study of the purification and characterization of the cytosolic glutathione S-transferases from two salmonid species: Atlantic salmon (Salmo salar) and brown trout (Salmo trutta). Comp biochem Physiol C 131, 207–213
Oakley AJ, Bello ML, Battistoni A, Ricci G, Rossjohn J, Villar HO, Parker MW (1997) The structures of human glutathione transferase P1-1 in complex with glutathione and various inhibitors at high resolution. J Mol Biol 274, 84–100
Overbaugh JM, Lau EP, Marino VA, Fall R (1988) Purification and preliminary characterization of a monomeric glutathione S-transferase from Tetrahymena thermophilia. Arch Biochem Biophys 261, 227–234
Pangrekar J, Sikka HC (1992) Xenobiotic metabolizing enzyme activity in the liver and kidney of the brown bullhead (Ictalurus nebulosus). Mar Environ Res 34, 287–291
Pemble SE, Taylor JB (1992) An evolutionary perspective on glutathione transferases inferred from class theta glutathione transferase cDNA sequences. Biochem J 287, 957–963
Pemble SE, Wardle AF, Taylor JB (1996) Glutathione S-transferase class kappa: characterization by the cloning of rat mitochondrial GST and identification of a human homologue. Biochem J 319, 749–754
Perez-Lopez M, Rouimi P, Debrauwer L, Cravedi JP (1998) Glutathione S-transferase subunits pattern in rainbow trout isolated hepatocytes. Mar Environ Res 46, 385–389
Perez-Lopez M, Anglade P, Bec-Ferte MP, Debrauwer L, Perdu E, Carvedi JP, Rouimi P (2000) Characterization of hepatic and extrahepatic glutathione S-transferases in rainbow trout (Oncorhynchus mykiss) and their induction by 3,3´,4,4´-tetrachlorobiphenyl. Fish Physiol Biochem 22, 21–32
Pickett CB (1989) Glutathione S-transferases: gene structure, regulation, and biological function. Annu Rev Biochem 58, 743–764
Polekhina G, Board PG, Blackburn AC, Parker MW (2001) Crystal structure of maleylacetoacetate isomerase/glutathione transferase zeta reveals the molecular basis for its remarkable catalytic promiscuity. Biochemistry 40, 1567–1576
Prapanthadara L, Ranson H, Somboon P, Hemingway J (1998) Cloning expression and characterization of an insect class I glutathione S-transferase from Anopheles dirus species B. Insect Biochem Mol Biol 28, 321–329
Prochaska HJ, DeLong MJ, Talalay P (1985) On the mechanism of induction of cancer-protective enzymes: a unifying proposal. Proc Natl Acad Sci USA 82, 8232–8236
Raghunathan S, Chandross RJ, Kretsinger RH, Allison TJ, Penington CJ, Rule GS (1994) Crystal structure of human class mu glutathione transferase GSTM2-2. Effects of lattice packing on the conformational heterogeneity. J Mol Biol 238, 815–832
Ramage PIN, Nimmo IA (1984) The substrate specificities and subunit compositions of the hepatic glutathione S-transferases of rainbow trout (Salmo gairdneri). Comp Biochem Physiol B 78, 189–194
Ranson H, Prapanthadara L, Hemingway J (1997) Cloning and characterization of two glutathione S-transferases from a DDT-resistant strain of Anopheles gambiae. Biochem J 324, 97–102
Reinemer P, Dirr HW, Landenstein R, Huber R, LoBello M, Federici G, Parker MW (1992) Three-dimensional structure of class pi glutathione S-transferase from human placenta in complex with S-hexylglutathione at 2.8 Å resolution. J Mol Biol 227, 214–226
Reinemer P, Prade L, Hof P, Neuefeind T, Huber R, Zettl R, Palme K, Schell J, Koelln I, Bartunik HD, Bieseler B (1996) Three-dimensional structure of glutathione S-transferase from Arabidopsis thaliana at 2.2 Å resolution: structural characterization of herbicide-conjugating plant glutathione S-transferases and a novel active site architecture. J Mol Biol 255, 289–309
Robinson A, Huttley GA, Booth HS, Board PG (2004) Modelling and bioinformatics studies of the human kappa class glutathione transferase predict a novel third transferase family with homology to prokaryotic 2-hydroxychromene-2-carboxylate isomerases. Biochem J 379, 541–552
Rossjohn J, McKinstry WJ, Oakley AJ, Verger D, Flanagan J, Chelvanayagam G, Tan KL, Board PG, Parker MW (1998) Human theta class glutathione transferase: the crystal structure reveals a sulfate-binding pocket within a buried active site. Structure 6, 309–322
Rushmore TH, Pickett CB (1993) Glutathione S-transferase, structure, regulation and therapeutic implications. J Biol Chem 268, 11475–11478
Salinas AE, Wong MG (1999) Glutathione S-transferases-a review. Curr Med Chem 6, 279–309
Shah DM, Hironaka CM, Wiegand RC, Harding EI, Krivi GG, Tiemeier DC (1986) Structural analysis of a maize gene coding for glutathione S-transferase involved in herbicide detoxification. Plant Mol Biol 6, 203–211
Sheehan D, Meade G, Foley VM, Dowd CA (2001) Structure, function and evolution of glutathione transferases: implications for classification of nonmammalian members of an ancient enzyme superfamily. Biochem J 360, 1–16
Sinning I, Kleywegt GJ, Cowan SW, Reinemer P, Dirr HW, Huber R, Gilliland GL, Armstrong RN, Ji X, Board PG, Olin B, Mannervik B, Jones TA (1993) Structure determination and refinement of human alpha class glutathione transferase A1-1, and a comparison with the mu and pi class enzymes. J Mol Biol 232, 192–212
Stenersen J, Kobro S, Bjerke M, Arend U (1987) Glutathione transferases in aquatic and terrestrial animals from nine phyla. Comp Biochem Physiol C 86, 73–82
Stevens JM, Armstrong RN, Dirr HW (2000) Electrostatic interactions affecting the active site of class sigma glutathione S-transferase. Biochem J 347, 193–197
Strausberg R, Feingold EA (2002) Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences. Proc Natl Acad Sci USA 99, 16899–16903
Suzuki T, Takagi Y, Osanai H, Li L, Takeuchi M, Katoh Y, Kobayashi M, Yamamoto M (2005) Pi class glutathione S-transferase genes are regulated by Nrf2 through an evolutionarily conserved regulatory element in zebrafish. Biochem J 388, 65–73
Syvanen M, Zhou ZH, Wang JY (1994) Glutathione transferase gene family from the housefly Musca domestica. Mol Gen Genet 245, 25–31
Tang SS, Chang GG (1996) Kinetic mechanism of octopus hepatopancreatic glutathione transferase in reverse micelles. Biochem J 315, 599–606
Tomarev SI, Zinovieva RD, Guo K, Piatigorssky J (1993) Squid glutathione S-transferase. Relationship with other glutathione S-transferases and the S-crystallins of cephalopods. J Biol Chem 268, 4534–4542
Tong Z, Board PG, Anders MW (1998) Glutathione transferase zeta catalyses the oxygenation of the carcinogen dichloroacetic acid to glyoxylic acid. Biochem J 331, 371–374
Toung YPS, Hsieh TS, Tu CPD (1991) The Drosophila glutathione S-transferase 1-1 is encoded by an intronless gene 87B. Biochem Biophys Res Commun 178, 1205–1211
US EPA (1996) New Bedford Harbor long-term monitoring assessment report: baseline sampling. U.S.EPA, office of research and development, Narragansett, RI. EPA 600/R-96/097
van der Oost J, Nederkoorn PH, Stouthamer AH, Westerhoff HV, van Spanning RJ (1996) An alternative model for heam ligation in nitrate reductase and analogous respiratory cytochrome b complexes. Mol Microbiol 22, 193–196
Vidal ML, Narbonne JF (2000) Characterization of glutathione S-transferase activity in the Asiatic Clam Corbicula fluminea. Bull Environ Contam Toxicol 64, 455–462
Vidal ML, Rouimi P, Debrauwer L, Narbonne JF (2002) Purification and characterization of glutathione S-transferases from the freshwater clam Corbicula fluminea (Müller). Comp Biochem Physiol C 131, 477–489
Vuilleumier S (1997) Bacterial glutathione S-transferases: What are they good for? J Bacteriol 179, 1431–1441
Wilce MCJ, Qakley AJ, Rossjohn J, Feil SC, LoBello M, Ricci G, Board PG, Parker MW (1996) “Crystallographic studies of Pi- and Theta-class glutathione-S-transferases”. In: Glutathione S-transferases: Structure, Function and Clinical Implications, Vermeulen NPE, Mulder GJ, Nieuwenhuyse H, Peters WHM, van Bladeren PJ, eds. (London: Taylor and Francis), pp 39–56
Woods IG, Wilson C, Friedlander B, Chang P, Reyes DK, Nix R, Kelly PD, Chu F, Postlethwait JH, Talbot WS (2005) The zebrafish gene map defines ancestral vertebrate chromosomes. Genome Res 15, 1307–1314
Xiao G, Liu S, Ji X, Johnson WW, Chen J, Parsons JF, Stevens WJ, Gilliland GL, Armstrong RN (1996) First sphere and second sphere electrostatic effects in the active site of a class mu glutathione transferase. Biochemistry 35, 4753–4765
Yang H, Nie L, Zhu S, Zhou X (2002) Purification and characterization of a novel glutathione S-transferase from Asaphis dichotoma. Arch Biochem Biophys 403, 202–208
Yang H, Zeng Q, Nie L, Zhu S, Zhou X (2003) Purification and characterization of a novel glutathione S-transferase from Atactodea striata. Biochem Biophys Res Commun 307, 626–631
Yang HL, Zeng QY, Li EQ, Zhu SG, Zhou XW (2004) Molecular cloning, expression and characterization of glutathione-S-transferase from Mytilus edulis. Comp Biochem Physiol B 139, 175–182
Yu SJ (1996) Insect glutathione S-transferases. Zool Stud 35, 9–19
Acknowedgments
This work was supported in part by the Camille and Henry Dreyfus Foundation and a DoD/Army Contract No. W911NF-06-1-0095. The authors thank Dr. Kai Zhang and Dr. Haihong Wang for their help in analyzing the gene data and crystal structure of GSTs.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Blanchette, B., Feng, X. & Singh, B.R. Marine Glutathione S-Transferases. Mar Biotechnol 9, 513–542 (2007). https://doi.org/10.1007/s10126-007-9034-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10126-007-9034-0