Abstract
The histone-like DNA-binding proteins (HU) serve as model molecules for protein thermostability studies, as they function in different bacteria that grow in a wide range of temperatures and show sequence diversity under a common fold. In this work, we report the cloning of the hutth gene from Thermus thermophilus, the purification and crystallization of the recombinant HUTth protein, as well as its X-ray structure determination at 1.7 Å. Detailed structural and thermodynamic analyses were performed towards the understanding of the thermostability mechanism. The interaction of HUTth protein with plasmid DNA in solution has been determined for the first time with MST. Sequence conservation of an exclusively thermophilic order like Thermales, when compared to a predominantly mesophilic order (Deinococcales), should be subject, to some extent, to thermostability-related evolutionary pressure. This hypothesis was used to guide our bioinformatics and evolutionary studies. We discuss the impact of thermostability adaptation on the structure of HU proteins, based on the detailed evolutionary analysis of the Deinococcus–Thermus phylum, where HUTth belongs. Furthermore, we propose a novel method of engineering thermostable proteins, by combining consensus-based design with ancestral sequence reconstruction. Finally, through the structure of HUTth, we are able to examine the validity of these predictions. Our approach represents a significant advancement, as it explores for the first time the potential of ancestral sequence reconstruction in the divergence between a thermophilic and a mainly mesophilic taxon, combined with consensus-based engineering.







Similar content being viewed by others
Abbreviations
- IPTG:
-
Isopropyl thio-β-d-galactoside
- E. coli :
-
Escherichia coli
- B. :
-
Bacillus
- G. :
-
Geobacillus
- T. :
-
Thermus
- Tvo:
-
Thermoplasma volcanium
- SDS-PAGE:
-
Sodium dodecyl sulphate–polyacrylamide gel electrophoresis
- MSA:
-
Multiple sequence alignment
- MST:
-
Microscale thermophoresis
- Tris:
-
Tris(hydroxymethyl)aminomethane
- T m :
-
Protein melting temperature
- DBD:
-
DNA-binding domain
- HTH:
-
Helix-turn-helix
- DS:
-
Dimerization signal
- EMSA:
-
Electrophoretic mobility shift assay
References
Aki TS, Adhya S (1997) Repressor induced site-specific binding of HU for transcriptional regulation. EMBO J 16:3666–3674
Ashkenazy H, Penn O, Doron-Faigenboim A, Cohen O, Cannarozzi G, Zomer O, Pupko T (2012) FastML: a web server for probabilistic reconstruction of ancestral sequences. Nucleic Acids Res 40(W1):W580–W584. doi:10.1093/nar/gks498
Balandina A, Claret L, Hengge-Aronis R, Rouviere-Yaniv J (2001) The Escherichia coli histone-like protein HU regulates rpoS translation. Mol Microbiol 39:1069–1079
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The protein data bank. Nucleic Acids Res 28:235–242
Berthold V, Geider K (1976) Interaction of DNA with DNA-binding proteins. The characterization of protein HD from Escherichia coli and its nucleic acid complexes. Eur J Biochem 71:443–449
Bhowmick T, Ghosh S, Dixit K, Ganesan V, Ramagopal UA, Dey D, Nagaraja V (2014) Targeting mycobacterium tuberculosis nucleoid-associated protein HU with structure-based inhibitors. Nat Commun 5:4124. doi:10.1038/ncomms5124
Biltonen RL, Freire E (1978) Thermodynamic characterization of conformational states of biological macromolecules using differential scanning calorimetry. CRC Crit Rev Biochem 5:85–124
Boelens R, Vis H, Vorgias CE, Wilson KS, Kaptein R (1996) Structure and dynamics of the DNA binding protein HU from Bacillus stearothermophilus by NMR spectroscopy. Biopolymers 40:553–559
Bonnefoy E, Takahashi M, Yaniv JR (1994) DNA-binding parameters of the HU protein of Escherichia coli to cruciform DNA. J Mol Biol 242:116–129
Boubrik F, Rouviere-Yaniv J (1995) Increased sensitivity to gamma irradiation in bacteria lacking protein HU. Proc Natl Acad Sci USA 92:3958–3962
Boyko K, Gorbacheva M, Rakitina T, Korzhenevskiy D, Vanyushkina A, Kamashev D, Lipkin A, Popov V (2015) Expression, purification, crystallization and preliminary X-ray crystallographic analysis of the histone-like HU protein from Spiroplasma melliferum KC3. Acta Crystallogr F Struct Biol Commun 71:24–27. doi:10.1107/S2053230X14025333
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Broyles SS, Pettijohn DE (1986) Interaction of the Escherichia coli HU protein with DNA. Evidence for formation of nucleosome-like structures with altered DNA helical pitch. J Mol Biol 187:47–60
Castaing B, Zelwer C, Laval J, Boiteux S (1995) HU protein of Escherichia coli binds specifically to DNA that contains single-strand breaks or gaps. J Biol Chem 270:10291–10296
Christodoulou E, Vorgias CE (1998) Cloning, overproduction, purification and crystallization of the DNA binding protein HU from the hyperthermophilic eubacterium Thermotoga maritima. Acta Crystallogr D 54:1043–1045
Christodoulou E, Vorgias CE (2002) The thermostability of DNA-binding protein HU from mesophilic, thermophilic, and extreme thermophilic bacteria. Extremophiles 6:21–31
Christodoulou E, Rypniewski WR, Vorgias CE (2003) High-resolution X-ray structure of the DNA-binding protein HU from the hyperthermophilic Thermotoga maritima and the determinants of its thermostability. Extremophiles 7:111–122
Cole MF, Gaucher EA (2011) Utilizing natural diversity to evolve protein function: applications towards thermostability. Curr Opin Chem Biol 15:399–406. doi:10.1016/j.cbpa.2011.03.005
Coste F, Hervouet N, Oberto J, Zelwer C, Castaing B (1999) Crystallization and preliminary X-ray diffraction analysis of the homodimeric form 2 of the HU protein from Escherichia coli. Acta Crystallogr D 55:1952–1954
Dame RT (2005) The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin. Mol Microbiol 56:858–870
Dame RT, Goosen N (2002) HU, promoting or counteracting DNA compaction? FEBS Lett 529:151–156
Damodaran S (2003) In situ measurement of conformational changes in proteins at liquid interfaces by circular dichroism spectroscopy. Anal Bioanal Chem 376:182–188
Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27:1164–1165. doi:10.1093/bioinformatics/btr088
Dri AM, Rouviere-Yaniv J, Moreau PL (1991) Inhibition of cell division in hupA hupB mutant bacteria lacking HU protein. J Bacteriol 173:2852–2863
Dri AM, Moreau PL, Rouvière-Yaniv J (1992) Role of the histone-like proteins OsmZ and HU in homologous recombination. Gene 120:11–16
Drlica K, Rouviere-Yaniv J (1987) Histonelike proteins of bacteria. Microbiol Rev 51:301–319
Durney MA, Wechselberger RW, Kalodimos CG, Kaptein R, Vorgias CE, Boelens R (2004) An alternate conformation of the hyperthermostable HU protein from Thermotoga maritima has unexpectedly high flexibility. FEBS Lett 563:49–54
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Emsley P, Lohkamp B, Scott WG, Cowtan K (2010) Features and development of Coot. Acta Crystallogr D Biol Crystallogr 66:486–501. doi:10.1107/S0907444910007493
Esser D, Rudolph R, Jaenicke R, Bohm G (1999) The HU protein from Thermotoga maritima: recombinant expression, purification and physicochemical characterization of an extremely hyperthermophilic DNA-binding protein. J Mol Biol 291:1135–1146
Evans PR, Murshudov GN (2013) How good are my data and what is the resolution? Acta Crystallogr D Biol Crystallogr 69:1204–1214. doi:10.1107/S0907444913000061
Fernández S, Rojo F, Alonso JC (1997) The Bacillus subtilis chromatin-associated protein Hbsu is involved in DNA repair and recombination. Mol Microbiol 23:1169–1179
Freire E (1995) Differential scanning calorimetry. Methods Mol Biol 40:191–218
Glyakina AV, Garbuzynskiy SO, Lobanov MY, Galzitskaya OV (2007) Different packing of external residues can explain differences in the thermostability of proteins from thermophilic and mesophilic organisms. Bioinformatics 23:2231–2238
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224. doi:10.1093/molbev/msp259
Greenfield NJ (1996) Methods to estimate the conformation of proteins and polypeptides from circular dichroism data. Anal Biochem 235:1–10
Grove A (2011) Functional evolution of bacterial histone-like HU proteins. Curr Issues Mol Biol 13:1–12
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. doi:10.1093/sysbio/syq010
Guo F, Adhya S (2007) Spiral structure of Escherichia coli HUalphabeta provides foundation for DNA supercoiling. Proc Natl Acad Sci USA 104:4309–4314
Haluzi H, Goitein D, Koby S, Mendelson I, Teff D, Mengeritsky G, Giladi H, Oppenheim AB (1991) Genes coding for integration host factor are conserved in gram-negative bacteria. J Bacteriol 173:6297–6299
Hodges-Garcia Y, Hagerman PJ, Pettijohn DE (1989) DNA ring closure mediated by protein HU. J Biol Chem 264:14621–14623
Holck A, Kleppe K (1985) Affinity of protein HU for different nucleic acids. FEBS Lett 185:121–124
Huisman O, Faelen M, Girard D, Jaffe A, Toussaint A, Rouviere-Yaniv J (1989) Multiple defects in Escherichia coli mutants lacking HU protein. J Bacteriol 171:3704–3712
Hwang DS, Kornberg A (1992) Opening of the replication origin of Escherichia coli by DnaA protein with protein HU or IHF. J Biol Chem 267:23083–23086
Kabsch W (2010) XDS. Acta Crystallogr D Biol Crystallogr 66:125–132. doi:10.1107/S0907444909047337
Kamashev D, Rouviere-Yaniv J (2000) The histone-like protein HU binds specifically to DNA recombination and repair intermediates. EMBO J 19:6527–6535
Kano Y, Imamoto F (1990) Requirement of integration host factor (IHF) for growth of Escherichia coli deficient in HU protein. Gene 89:133–137
Kano Y, Wada M, Nagase T, Imamoto F (1986) Genetic characterization of the gene hupB encoding the HU-1 protein of Escherichia coli. Gene 45:37–44
Karshikoff A, Nilsson L, Ladenstein R (2015) Rigidity versus flexibility: the dilemma of understanding protein thermal stability. FEBS J 282:3899–3917. doi:10.1111/febs.13343
Kawamura S, Abe Y, Ueda T, Masumoto K, Imoto T, Yamasaki N, Kimura M (1998) Investigation of the structural basis for thermostability of DNA-binding protein HU from Bacillus stearothermophilus. J Biol Chem 273:19982–19987
Kim do H, Im H, Jee JG, Jang SB, Yoon HJ, Kwon AR, Kang SM, Lee BJ (2014) β-Arm flexibility of HU from Staphylococcus aureus dictates the DNA-binding and recognition mechanism. Acta Crystallogr D Biol Crystallogr 70:3273–3289. doi:10.1107/S1399004714023931
Krissinel E, Henrick K (2007) Inference of macromolecular assemblies from crystalline state. J Mol Biol 372:774–797
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Lavoie BD, Chaconas G (1994) A second high affinity HU binding site in the phage Mu transpososome. J Biol Chem 269:15571–15576
Lehmann M, Wyss M (2001) Engineering proteins for thermostability, the use of sequence alignments versus rational design and directed evolution. Curr Opin Biotechnol 12:371–375
Li S, Waters R (1998) Escherichia coli strains lacking protein HU are UV sensitive due to a role for HU in homologous recombination. J Bacteriol 180:3750–3756
Liu D, Yumoto H, Murakami K, Hirota K, Ono T, Nagamune H, Kayama S, Matsuo T, Miyake Y (2008) The essentiality and involvement of Streptococcus intermedius histone-like DNA-binding protein in bacterial viability and normal growth. Mol Microbiol 68:1268–1282. doi:10.1111/j.1365-2958.2008.06232.x
Luijsterburg MS, Noom MC, Wuite GJ, Dame RT (2006) The architectural role of nucleoid-associated proteins in the organization of bacterial chromatin: a molecular perspective. J Struct Biol 156:262–272
Macvanin M, Adhya S (2012) Architectural organization in E. coli nucleoid. Biochim Biophys Acta 1819:830–835. doi:10.1016/j.bbagrm.2012.02.012
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2015) CDD: NCBI's conserved domain database. Nucleic Acids Res 43:D222–226. doi:10.1093/nar/gku1221
Maugini E, Tronelli D, Bossa F, Pascarella S (2009) Structural adaptation of the subunit interface of oligomeric thermophilic and hyperthermophilic enzymes. Comput Biol Chem 33:137–148
McRee DE (1999) XtalView/Xfit—A versatile program for manipulating atomic coordinates and electron density. J Struct Biol 125:156–165
Micka B, Marahiel MA (1992) The DNA-binding protein HBsu is essential for normal growth and development in Bacillus subtilis. Biochimie 74:641–650
Mouw KW, Rice PA (2007) Shaping the Borrelia burgdorferi genome: crystal structure and binding properties of the DNA-bending protein Hbb. Mol Microbiol 63:1319–1330
Murshudov GN, Vagin AA, Dodson EJ (1997) Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr D 53:240–255
Oberto J, Nabti S, Jooste V, Mignot H, Rouviere-Yaniv J (2009) The HU regulon is composed of genes responding to anaerobiosis, acid stress, high osmolarity and SOS induction. PLoS One 4:e4367. doi:10.1371/journal.pone.0004367
Orfaniotou F, Tzamalis P, Thanassoulas A, Stefanidi E, Zees A, Boutou E, Vlassi M, Nounesis G, Vorgias CE (2009) The stability of the archaeal HU histone-like DNA-binding protein from Thermoplasma volcanium. Extremophiles 13:1–10
Padas PM, Wilson KS, Vorgias CE (1992) The DNA-binding protein HU from mesophilic and thermophilic bacilli: gene cloning, overproduction and purification. Gene 117:39–44
Painbeni E, Caroff M, Rouviere-Yaniv J (1997) Alterations of the outer membrane composition in Escherichia coli lacking the histone-like protein HU. Proc Natl Acad Sci USA 94:6712–6717
Pettijohn DE (1988) Histone-like proteins and bacterial chromosome structure. J Biol Chem 263:12793–12796
Pontiggia A, Negri A, Beltrame M, Bianchi ME (1993) Protein HU binds specifically to kinked DNA. Mol Microbiol 7:343–350
Privalov PL, Potekhin SA (1986) Scanning microcalorimetry in studying temperature-induced changes in proteins. Method Enzymol 131:4–51
Ramstein J, Hervouet N, Coste F, Zelwer C, Oberto J, Castaing B (2003) Evidence of a thermal unfolding dimeric intermediate for the Escherichia coli histone-like HU proteins: thermodynamics and structure. J Mol Biol 331:101–121
Risso VA, Gavira JA, Gaucher EA, Sanchez-Ruiz JM (2014) Phenotypic comparisons of consensus variants versus laboratory resurrections of Precambrian proteins. Proteins Struct Funct Bioinf 82:887–896
Robinson-Rechavi M, Alibés A, Godzik A (2006) Contribution of electrostatic interactions, compactness and quaternary structure to protein thermostability: lessons from structural genomics of Thermotoga maritima. J Mol Biol 356:547–557
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. doi:10.1093/sysbio/sys029
Rouviere-Yaniv J, Yaniv M, Germond JE (1979) E. coli DNA binding protein HU forms nucleosome like structure with circular double-stranded DNA. Cell 17:265–274
Ruiz-Sanz J, Filimonov VV, Christodoulou E, Vorgias CE, Mateo PL (2004) Thermodynamic analysis of the unfolding and stability of the dimeric DNA-binding protein HU from the hyperthermophilic eubacterium Thermotoga maritima and its E34D mutant. Eur J Biochem 271:1497–1507
Sælensminde G, Halskau Ø Jr, Jonassen I (2009) Amino acid contacts in proteins adapted to different temperatures: hydrophobic interactions and surface charges play a key role. Extremophiles 13:11–20
Scopes RK (1974) Measurement of protein by spectrophotometry at 205 nm. Anal Biochem 59:277–282
Swinger KK, Lemberg KM, Zhang Y, Rice PA (2003) Flexible DNA bending in HU-DNA cocrystal structures. EMBO J 22:3749–3760
Tanaka I, Appelt K, Dijk J, White SW, Wilson KS (1984) 3-A resolution structure of a protein with histone-like properties in prokaryotes. Nature 310:376–381
Tina KG, Bhadra R, Srinivasan N (2007) PIC: protein interactions calculator. Nucleic Acids Res 35:W473–W476. doi:10.1093/nar/gkm423 (Web Server issue)
Vagin A, Teplyakov A (1997) MOLREP: an automated program for molecular replacement. J Appl Crystallogr 30:1022–1025
Vis H, Mariani M, Vorgias CE, Wilson KS, Kaptein R, Boelens R (1995) Solution structure of the HU protein from Bacillus stearothermophilus. J Mol Biol 254:692–703
Wada M, Kano Y, Ogawa T, Okazaki T, Imamoto F (1988) Construction and characterization of the deletion mutant of hupA and hupB genes in Escherichia coli. J Mol Biol 204:581–591
White SW, Appelt K, Wilson KS, Tanaka I (1989) A protein structural motif that bends DNA. Proteins 5:281–288
White SW, Wilson KS, Appelt K, Tanaka I (1999) The high-resolution structure of DNA-binding protein HU from Bacillus stearothermophilus. Acta Crystallogr D Biol Crystallogr 55:801–809
Wienken CJ, Baaske P, Rothbauer U, Braun D, Duhr S (2010) Protein-binding assays in biological liquids using microscale thermophoresis. Nat Commun 1:100. doi:10.1038/ncomms1093
Wijma HJ, Floor RJ, Janssen DB (2013) Structure- and sequence-analysis inspired engineering of proteins for enhanced thermostability. Curr Opin Str Biol 23:588–594. doi:10.1016/j.sbi.2013.04.008
Wilson KS, Vorgias CE, Tanaka I, White SW, Kimura M (1990) The thermostability of DNA-binding protein HU from Bacilli. Protein Eng 4:11–22
Winn MD, Ballard CC, Cowtan KD, Dodson EJ, Emsley P, Evans PR, Keegan RM, Krissinel EB, Leslie AG, McCoy A, McNicholas SJ, Murshudov GN, Pannu NS, Potterton EA, Powell HR, Read RJ, Vagin A, Wilson KS (2011) Overview of the CCP4 suite and current developments. Acta Crystallogr D Biol Crystallogr 67:235–242. doi:10.1107/S0907444910045749
Yasuzawa K, Hayashi N, Goshima N, Kohno K, Imamoto F, Kano Y (1992) Histone-like proteins are required for cell growth and constraint of supercoils in DNA. Gene 122:9–15
Yu J, Zhou Y, Tanaka I, Yao M (2010) Roll: a new algorithm for the detection of protein pockets and cavities with a rolling probe sphere. Bioinformatics 26:46–52
Zentgraf H, Berthold V, Geider K (1977) Interaction of DNA with DNA binding proteins II. Displacement of Escherichia coli DNA unwinding protein and the condensed structure of DNA complexed with protein HD. Biochim Biophys Acta BBA Nucleic Acids Protein Synth 474:629–638
Zillner K, Jerabek-Willemsen M, Duhr S, Braun D, Längst G, Baaske P (2012) Microscale thermophoresis as a sensitive method to quantify protein, nucleic acid interactions in solution. Methods Mol Biol 815:241–252. doi:10.1007/978-1-61779-424-7_18
Acknowledgments
We acknowledge technical support by the SPC facility at EMBL Hamburg in the frame of Biostruct-X (EE FP7). We would also like to acknowledge technical assistance from A. Tsoka. F. S. has been supported in part by the ARISTEIA I program, administered by the General Secretariat of Research and Technology of Greece, co-financed by the European Social Fund and the State of Greece. P. S. A. is supported by a Ph.D. fellowship from Paris Diderot University and funds by the Ph.D. Program Frontieres du Vivant (FdV)–Program Bettencourt.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by L. Huang.
Electronic supplementary material
Below is the link to the electronic supplementary material.
792_2016_859_MOESM1_ESM.pdf
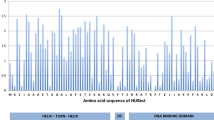
Supplementary Figure 1: The amino acid primary structure of the HUTth protein and the corresponding hubest gene (PDF 28 kb)
792_2016_859_MOESM2_ESM.pdf
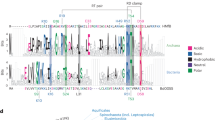
Supplementary Figure 2: Sequence alignment of the mesophilic HUBsu, with the thermophilics HUBst and HUTvo and the hyperthermophilic HUTmar and HUTth. The secondary structure elements are also indicated. The amino acids that are related with the thermostability are indicated by * (PDF 197 kb)
792_2016_859_MOESM3_ESM.pdf
Supplementary Figure 3: HU gene trees for the Deinococcus–Thermus phylum, respectively. Growth temperature is included at the end of gene names. Maximum likelihood (A) and Bayesian (B) gene trees possessed very similar topologies. However, due to the few informative positions in HU proteins, many branches remained unresolved and/or were poorly supported. The Deinococcus branch is comparatively very long, and even includes some internal long branches. Most species bear multiple homologs, some of which are situated in plasmids, and sequence conservation are generally lower. We attempted to remove some of the long branches, but Deinococcus is very sensitive to taxon sampling, thus the topology becomes altered and new long branches appear. The branch of the Thermus genus was inadequately resolved probably, because of very high identity (>90 %) among sequences. Truepera radiovictrix is consistently grouped with the Thermales instead of its commonly accepted position in the Deinococcales. It could be that conservation of sequence due to thermostability in a short sequence can create such an artifact (PDF 336 kb)
792_2016_859_MOESM4_ESM.pdf
Supplementary Table 1: HUTth homodimer. The H-bonding interactions between the protomers. †The solvent numbering refers to the deposited structure with pdb code 5EKA (PDF 344 kb)
792_2016_859_MOESM5_ESM.pdf
Supplementary Table 2: Intermolecular contacts between subunits of closely related HU-DNA-binding protein structures. The computation of the type and number of interactions were carried out at the server http://pic.mbu.iisc.ernet.in/ designed by Tina et al. 2007. 5EKA HU-DNA-binding protein from Thermus thermophilus; 1B8Z HU from Thermotoga maritima; 1HUU DNA-binding protein HU from G. stearothermophilus (B. stearothermophilus); 1P71 Anabaena HU-DNA corcrystal; 3RHI DNA-binding protein HU from B. anthracis; 4QJU crystal structure of DNA-bound nucleoid associated protein, SAV1473; and 2O97 crystal structure of E. coli HU heterodimer (PDF 45 kb)
792_2016_859_MOESM6_ESM.pdf
Supplementary Table 3: Calculation of buried surfaces and cavities between subunits of closely related HU-DNA-binding protein structures. The computations of the surfaces were carried out with PISA (http://www.ebi.ac.uk/pdbe/pisa/) (Krissinel et al. 2007). The computations of the cavities were carried out with POCASA (altair.sci.hokudai.ac.jp/g6/service/pocasa/) (Yu et al. 2010) (PDF 42 kb)
Rights and permissions
About this article
Cite this article
Papageorgiou, A.C., Adam, P.S., Stavros, P. et al. HU histone-like DNA-binding protein from Thermus thermophilus: structural and evolutionary analyses. Extremophiles 20, 695–709 (2016). https://doi.org/10.1007/s00792-016-0859-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-016-0859-1