Abstract
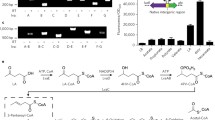
An extremely thermophilic bacterium, Geobacillus thermoleovorans B23, is capable of degrading a broad range of alkanes (with carbon chain lengths ranging between C11 and C32) at 70 °C. Whole-genome sequence analysis revealed that unlike most alkane-degrading bacteria, strain B23 does not possess an alkB-type alkane monooxygenase gene. Instead, it possesses a cluster of three ladA-type genes, ladAαB23, ladAβB23, and ladB B23, on its chromosome, whose protein products share significant amino acid sequence identities, 49.8, 34.4, and 22.7 %, respectively, with that of ladA alkane monooxygenase gene found on a plasmid of Geobacillus thermodetrificans NG 80-2. Each of the three genes, ladAαB23, ladAβB23, and ladB B23, was heterologously expressed individually in an alkB1 deletion mutant strain, Pseudomonas fluorescens KOB2Δ1. It was found that all three genes were functional in P. fluorescens KOB2Δ1, and partially restored alkane degradation activity. In this study, we suggest that G. thermoleovorans B23 utilizes multiple LadA-type alkane monooxygenases for the degradation of a broad range of alkanes.





Similar content being viewed by others
Abbreviations
- C12:
-
Dodecane
- C16:
-
Hexadecane
References
Altschul SF, Madden TL, Schäffer JZ, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Boonmak C, Takahashi Y, Morikawa M (2013) Draft genome sequence of Geobacillus thermoleovorans strain B23. Genome A 1(6):e00944-13
Darling AE, Mau B, Perna NT (2010) Progressive mauve: multiple genome alignment with gene gain, loss, and rearrangement. PLoS One 5:e11147
Das N, Chandran P (2011) Microbial degradation of hydrocarbon contaminants: an overview. Biotechnol Res Int, Article ID 941810
Eichhorn E, Davey CA, Sargent DF, Leisinger T, Richmond J (2002) Crystal structure of Escherichia coli alkanesulfonate monooxygenase SsuD. J Mol Biol 324:457–468
Ellis HR (2010) The FMN-dependent two-component monooxygenase systems. Arch Biochem Biophys 497:1–12
Felsenstein J (1985) Confidence limits on phylogenesis: an approach using the bootstrap. Evolution 39:783–791
Feng L, Wang W, Cheng J, Ren Y, Zhao G, Gao C, Tang Y, Liu X, Han W, Peng X, Liu R, Wang L (2007) Genome and proteome of long-chain alkane degrading Geobacillus thermodenitrificans NG80-2 isolated from a deep-subsurface oil reservoir. Proc Natl Acad Sci USA 104:5602–5607
Højberg O, Schnider U, Winteler HV, Sørensen J, Haas D (1999) Oxygen-sensing reporter strain of Pseudomonas fluorescens for monitoring the distribution of low-oxygen habitats in soil. Appl Environ Microbiol 65:4085–4093
Inoue H, Nojima H, Okayama H (1990) High efficiency transformation of Escherichia coli with plasmids. Gene 96:23–28
Kato T, Haruki M, Imanaka T, Morikawa M, Kanaya S (2001a) Isolation and characterization of long-chain-alkane degrading Bacillus thermoleovorans from deep subterranean petroleum reservoirs. J Biosci Bioeng 91:64–70
Kato T, Miyanaga A, Haruki M, Imanaka T, Morikawa M, Kanaya S (2001b) Gene cloning of an alcohol dehydrogenase from thermophilic alkane degrading Bacillus thermoleovorans B23. J Biosci Bioeng 91:100–102
Kato T, Miyanaga A, Kanaya S, Morikawa M (2009) Alkane inducible proteins in Geobacillus thermoleovorans B23. BMC Microbiol 9:60
Kato T, Miyanaga A, Kanaya S, Morikawa M (2010) Gene cloning and characterization of an aldehyde dehydrogenase from long-chain alkane-degrading Geobacillus thermoleovorans B23. Extremophiles 14:33–39
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Lageveen RG, Huisman GW, Preusting H, Ketelaar P, Eggink G, Witholt B (1988) Formation of polyesters by Pseudomonas oleovorans: effect of substrates on formation and composition of poly-(R)-3-hydroxyalkanoates and poly-(R)-3-hydroxyalkenoates. Appl Environ Microbiol 54:2924–2932
Li L, Liu X, Yang W, Xu F, Wang W, Feng L, Bartlam M, Wang L, Rao Z (2008) Crystal structure of long-chain alkane monooxygenase (LadA) in complex with coenzyme FMN: unveiling the long-chain alkane hydroxylase. J Mol Biol 376:453–465
Marmur J (1961) A procedure for the isolation of deoxyribonucleic acid from microorganisms. J Mol Biol 3:208–218
Morikawa M (2010) Dioxygen activation responsible for oxidation of aliphatic and aromatic hydrocarbon compounds: current state and variants. Appl Microbiol Biotechnol 87:1595–1603
Nie Y, Fang H, Li Y, Chi CQ, Tang YQ, Wu XL (2013) The genome of the moderate halophile Amycolicicoccus subflavus DQS3-9A1T reveals four alkane hydroxylation systems and provides some clues on the genetic basis for its adaptation to a petroleum environment. PLoS One 8:e70986
Rojo F (2009) Degradation of alkanes by bacteria. Environ Microbiol 11:2477–2490
Rost B (1999) Twilight zone of protein sequence alignments. Prot Eng 12:85–94
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Smits THM, Seeger MA, Witholt B, van Beilen JB (2001) New alkane-responsive expression vectors for Escherichia coli and Pseudomonas. Plasmid 46:16–24
Smits THM, Balada SB, Witholt B, van Beilen JB (2002) Functional analysis of alkane hydroxylase from Gram-negative and Gram-positive bacteria. J Bacteriol 184:1733–1742
Takei D, Washio K, Morikawa M (2008) Identification of alkane hydroxylase genes in Rhodococcus sp. strain TMP2 that degrades a branched alkane. Biotechnol Lett 30:1447–1452
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tourova TP, Nazina TZ, Mikhailova EM, Rodionova TA, Ekimov AN, Mashukova AV, Poltaraus AB (2008) alkB homologs in thermophilic bacteria of the genus Geobacillus. Mol Biol 42:217–226
van Beilen JB, Funhoff E (2007) Alkane hydroxylases involved in microbial alkane degradation. Appl Microbiol Biotechnol 74:13–21
van der Ploeg JR, Iwanicka-Nowickka R, Bykowski T, Hryniewicz MM, Leisinger T (1999) The Escherichia coli ssuEADCB gene cluster is required for the utilization of sulfur from aliphatic sulfonates and is regulated by the transcriptional activator CbI. J Biol Chem 274(41):29358–29365
Yanisch-Perron C, Vieira J, Messing J (1985) Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103–109
Acknowledgments
Pseudomonas fluorescens KOB2Δ1 and pCom8 were kind gifts from Dr. Theo H. M. Smits (Agroscope, Wädenswil, Switzerland). This work was supported by KAKENHI (19380189) to M.M.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by L. Huang.
Rights and permissions
About this article
Cite this article
Boonmak, C., Takahashi, Y. & Morikawa, M. Cloning and expression of three ladA-type alkane monooxygenase genes from an extremely thermophilic alkane-degrading bacterium Geobacillus thermoleovorans B23. Extremophiles 18, 515–523 (2014). https://doi.org/10.1007/s00792-014-0636-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00792-014-0636-y