Abstract
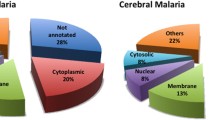
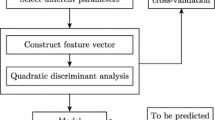
As a parasitic protozoan, Plasmodium falciparum (P. falciparum) can cause malaria. The mitochondrial proteins of malaria parasite play important roles in the discovery of anti-malarial drug targets. Thus, accurate identification of mitochondrial proteins of malaria parasite is a key step for understanding their functions and finding potential drug targets. In this work, we developed a sequence-based method to identify the mitochondrial proteins of malaria parasite. At first, we extended adjoining dipeptide composition to g-gap dipeptide composition for discretely formulating the protein sequences. Subsequently, the analysis of variance (ANOVA) combined with incremental feature selection (IFS) was used to pick out the optimal features. Finally, the jackknife cross-validation was used to evaluate the performance of the proposed model. Evaluation results showed that the maximum accuracy of 97.1 % could be achieved by using 101 optimal 5-gap dipeptides. The comparison with previous methods demonstrated that our method was accurate and efficient.



Similar content being viewed by others
Abbreviations
- ANOVA:
-
Analysis of variance
- auROC:
-
Area under the receiver operating characteristic
- IFS:
-
Incremental feature selection
- MCC:
-
Matthews correlation coefficient
- ROC:
-
Receiver operating characteristic
- Sn:
-
Sensitivity
- Sp:
-
Specificity
- Acc:
-
Overall accuracy
- SVM:
-
Support vector machine
References
Bender A, van Dooren GG, Ralph SA, McFadden GI, Schneider G (2003) Properties and prediction of mitochondrial transit peptides from Plasmodium falciparum. Mol Biochem Parasitol 132:59–66. doi:10.1016/j.molbiopara.2003.07.001
Chen YL, Li QZ, Zhang LQ (2012) Using increment of diversity to predict mitochondrial proteins of malaria parasite: integrating pseudo-amino acid composition and structural alphabet. Amino Acids 42:1309–1316. doi:10.1007/s00726-010-0825-7
Coordinators NR (2014) Database resources of the national center for biotechnology information. Nucleic Acids Res 42:D7–17. doi:10.1093/nar/gkt1146
Ding C, Yuan LF, Guo SH, Lin H, Chen W (2012) Identification of mycobacterial membrane proteins and their types using over-represented tripeptide compositions. J Proteomics 77:321–328. doi:10.1016/j.jprot.2012.09.006
Ding H et al (2013) Prediction of Golgi-resident protein types by using feature selection technique. Chemometr Intell Lab 124:9–13. doi:10.1016/j.chemolab.2013.03.005
Ding H, Deng EZ, Yuan LF, Liu L, Lin H, Chen W, Chou KC (2014a) iCTX-Type: a sequence-based predictor for identifying the types of conotoxins in targeting ion channels. Biomed Res Int 2014:286419. doi:10.1155/2014/286419
Ding H, Feng PM, Chen W, Lin H (2014b) Identification of bacteriophage virion proteins by the ANOVA feature selection and analysis. Mol BioSyst 10:2229–2235. doi:10.1039/c4mb00316k
Fan RE, Chen PH, Lin CJ (2005) Working set selection using second order information for training support vector machines. J Mach Learn Res 6:1889–1918
Guda C, Guda P, Fahy E, Subramaniam S (2004) MITOPRED: a web server for the prediction of mitochondrial proteins. Nucleic Acids Res 32:W372–W374. doi:10.1093/nar/gkh374
Guo SH, Deng EZ, Xu LQ, Ding H, Lin H, Chen W, Chou KC (2014) iNuc-PseKNC: a sequence-based predictor for predicting nucleosome positioning in genomes with pseudo k-tuple nucleotide composition. Bioinformatics 30:1522–1529. doi:10.1093/bioinformatics/btu083
Hayat M, Tahir M, Khan SA (2014) Prediction of protein structure classes using hybrid space of multi-profile Bayes and bi-gram probability feature spaces. J Theor Biol 346:8–15. doi:10.1016/j.jtbi.2013.12.015
Jia C, Liu T, Chang AK, Zhai Y (2011) Prediction of mitochondrial proteins of malaria parasite using bi-profile Bayes feature extraction. Biochimie 93:778–782. doi:10.1016/j.biochi.2011.01.013
Jia C, Lin X, Wang Z (2014) Prediction of protein S-nitrosylation sites based on adapted normal distribution bi-profile Bayes and Chou’s pseudo amino acid composition. Int J Mol Sci 15:10410–10423. doi:10.3390/ijms150610410
Kumar M, Verma R, Raghava GPS (2006) Prediction of mitochondrial proteins using support vector machine and hidden Markov model. J Biol Chem 281:5357–5363. doi:10.1074/jbc.M511061200
Lin H, Chen W (2011) Prediction of thermophilic proteins using feature selection technique. J Microbiol Methods 84:67–70. doi:10.1016/j.mimet.2010.10.013
Lin H, Chen W, Yuan LF, Li ZQ, Ding H (2013) Using over-represented tetrapeptides to predict protein submitochondria locations. Acta Biotheor 61:259–268. doi:10.1007/s10441-013-9181-9
Nanni L, Lumini A, Brahnam S (2014) An empirical study of different approaches for protein classification. Sci World J 2014:236717. doi:10.1155/2014/236717
Qiu WR, Xiao X, Lin WZ, Chou KC (2014) iMethyl-PseAAC: identification of protein methylation sites via a pseudo amino acid composition approach. Biomed Res Int 2014:947416. doi:10.1155/2014/947416
Saha I et al (2014) Ensemble learning prediction of protein-protein interactions using proteins functional annotations. Mol BioSyst 10:820–830. doi:10.1039/c3mb70486f
Vaidya AB, Mather MW (2009) Mitochondrial evolution and functions in malaria parasites. Annu Rev Microbiol 63:249–267. doi:10.1146/annurev.micro.091208.073424
Verma R, Varshney GC, Raghava GP (2010) Prediction of mitochondrial proteins of malaria parasite using split amino acid composition and PSSM profile. Amino Acids 39:101–110. doi:10.1007/s00726-009-0381-1
World Health Organization (2014). http://www.who.int/en/
Acknowledgments
We would like to thank the anonymous reviewers for their valuable suggestions. This work was supported by the National Nature Scientific Foundation of China (No. 61301260) and the Fundamental Research Funds for the Central Universities (No. ZYGX2012J113).
Conflict of interest
The authors declare that there is no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Ding, H., Li, D. Identification of mitochondrial proteins of malaria parasite using analysis of variance. Amino Acids 47, 329–333 (2015). https://doi.org/10.1007/s00726-014-1862-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-014-1862-4