Abstract
The medicinal effects and techniques for cultivating Anoectochilus formosanus are well-documented, but little is known about the mycorrhizal fungi associated with A. formosanus. Rhizoctonia (Thanatephorus) anastomosis group 6 (AG-6) was the most common species isolated from fungal pelotons in native A. formosanus and represented 67 % of the sample. Rhizoctonia (Ceratobasidium) AG-G, P, and R were also isolated and represent the first occurrence in the Orchidaceae. Isolates of AG-6, AG-R, and AG-P in clade I increased seed germination 44–91 % and promoted protocorm growth from phases III to VI compared to asymbiotic treatments and isolates of AG-G in clade II and Tulasnella species in clade III. All isolates in clades I to III formed fungal pelotons in tissue-cultured seedlings of A. formosanus, which exhibited significantly greater growth than nonmycorrhizal seedlings. An analysis of the relative effect of treatment (\( {\widehat{p}}_i \)) showed that the low level of colonization (\( {\widehat{p}}_i = 0.30\hbox{--} 0.47 \)) by isolates in clade I resulted in a significant increase in seedling growth compared to isolates in clades II (0.63–0.82) and III (0.63–0.75). There was also a negative correlation (r = −0.8801) with fresh plant weight and fungal colonization. Our results suggest that isolates in clade I may represent an important group associated with native populations of A. formosanus and can vary in their ability to establish a symbiotic association with A. formosanus. The results presented here are potentially useful for advancing research on the medicinal properties, production, and conservation of A. formosanus in diverse ecosystems.
Similar content being viewed by others
Introduction
The genus Anoectochilus (Orchidaceae), which comprises more than 40 species, is widespread throughout tropical regions. Several species of this genus are used in Chinese folk medicines, such as Anoectochilus formosanus Hayata and A noectochilus koshunensis Hayata, which are terrestrial orchids and distributed only in Taiwan and Okinawa (Japan) (Gutiérrez 2010).
Because of its ornamental value and medicinal properties (Tseng et al. 2006; Wu et al. 2007), the number of native A. formosanus has been greatly decreased due to increased harvesting by intentional picking and uprooting from native habitats (Cheng and Chang 2009). A. formosanus collected from the wild are often sold for more than 300 USD per kilogram (Chang et al. 2007). Using plant tissue techniques (Ho et al. 1987), the propagation and cultivation of A. formosanus has been established in Taiwan for at least 20 years. However, pot culture and transplantation of the tissue-cultured plantlets in soil without colonized mycorrhizal fungi have been challenging due to poor plant growth and increased susceptibility to mites (Chang et al. 2007) and plant pathogenic fungi Fusarium oxysporum f. sp. anoectochili (Huang et al. 2014) and the oomycete species of Pythium (e.g., P ythium aphanidermatum, Pythium myriotylum, and Pythium splendens) (Chang 1999) during ex vitro growth. Therefore, we hypothesized that plant survival could be increased by inoculating seeds or tissue-cultured plantlets with mycorrhizal fungi. However, there is a paucity of knowledge regarding the mycorrhizal associations between native A. formosanus and its associated mycorrhizal fungi.
The beneficial association of symbiotic endomycorrhizal fungi with orchids is well-documented (Rasmussen 2002). In nature, the seeds of orchids are minute and contain few stored food reserves, and colonization by a mycorrhizal fungus provides nutrients that are important for seed germination and seedling establishment (Dearnaley 2007). In either green or in achlorophyllous species of orchids, the dependency on mycorrhizal fungi can continue into adulthood (Gebauer and Meyer 2003; Julou et al. 2005; Rasmussen and Rasmussen 2007). Recently, the applications of mycorrhizal association for horticultural and conservation purposes have gained considerable attention (Zettler et al. 2007; Swarts and Dixon 2009).
With few exceptions (Selosse et al. 2004), the majority of orchid mycorrhizal (OM) fungi belong to early diverging lineages in the phylum Basidiomycota (Moncalvo et al. 2006; Smith and Read 2008). Rasmussen (2002) suggested that photosynthetic orchids associate with a wide range of Rhizoctonia species. The form-genus Rhizoctonia D.C. includes Thanatephorus Donk, Ceratobasidium D.P. Rogers, Tulasnella Schröeter, and Sebacina Tul. (Sneh et al. 1991; González García et al. 2006; Smith and Read 2008). The specific association of photosynthetic orchids with Rhizoctonia fungi in the Tulasnellaceae and Ceratobasidiaceae (e.g., cantharelloid clade) has been previously reported (Otero et al. 2002; Ma et al. 2003; Suarez et al. 2006). Certain photosynthetic orchids, even when sampled over a wide range, have a single dominant species of Rhizoctonia fungus (McCormick et al. 2004; Shefferson et al. 2005, 2010; Irwin et al. 2007). However, fully mycoheterotrophic (MH) orchids, which are achlorophyllous and nutritionally dependent on their mycorrhizal fungi, can be colonized by several different ectomycorrhizal (ECM) fungi (e.g., Russulaceae and Thelephoraceae fungi) (Roy et al. 2009; Kennedy et al. 2011) as well as saprobic and parasitic fungi (Smith and Read 2008). By contrast, fully MH orchids revealed associations with more diverse fungal lineages (Dearnaley et al. 2012). Recently, certain reports have highlighted that Rhizoctonia fungi also have the ability to associate with orchids as OM fungi and with autotrophic plants as ECM fungi at the same time to form a tripartite symbiosis (Yagame et al. 2008, 2012; Bougoure et al. 2009).
The mycorrhizal specificity of orchids with Rhizoctonia fungi has been the subject of debate for many years (Leake 2005). Molecular and sequence-based analyses have provided an efficient and rapid means of characterizing and identifying strains of endomycorrhizal Rhizoctonia fungi that associate with orchids (Otero et al. 2002; Ma et al. 2003; Suarez et al. 2006; Taylor and McCormick 2008). Phylogenetic analysis of the fungal ribosomal DNA-internal transcribed spacer (rDNA-ITS) sequences has also suggested that both binucleate Rhizoctonia spp. (teleomorph = Ceratobasidium) and multinucleate Rhizoctonia spp. (teleomorph = Thanatephorus) can be divided into different anastomosis groups (AGs), which are determined by examining macroscopic somatic hyphal interactions between a representative tester strain and an unknown Rhizoctonia isolate (Carling et al. 1999; Sharon et al. 2008).
The primary objective of this study was to characterize and identify Rhizoctonia fungi sampled from fungal pelotons in A. formosanus collected from native populations in different geographic regions of Taiwan. Subsequent experiments were conducted to determine if these fungi promoted seed germination and seedling growth of A. formosanus. Furthermore, we investigated the degree of colonization caused by endomycorrhizal fungi under an in vitro condition. No information is currently available on the effects of the density of colonized endomycorrhizal fungi in A. formosanus. The results obtained from this study provide information on the characterization and colonization of endomycorrhizal fungi to A. formosanus and represent potentially useful data for medicinal orchid production and conservation in native habitats.
Materials and methods
Isolation of endomycorrhizal fungi
Samples of native A. formosanus collected from seven sites in northern and central Taiwan were investigated for mycorrhizal fungi. Locations of plants, sampling dates, and original sources of the plant samples are presented in Table 1.
Each individual of A formosanus was divided into three separate parts: the roots, rhizome, and junction between root and rhizome. Sections of cortex tissue with fungal pelotons were excised, surface-disinfested in a 1 % solution of NaOCl for 1 min, rinsed in sterile water, and transferred to tap water agar (TWA) and potato dextrose agar (PDA) acidified with 0.2 % lactic acid to inhibit growth of bacteria. After incubation for 24–48 h at 24–26 °C, cortical cells were examined daily for the presence of fungal growth, and hyphal tips were excised and transferred to a fresh plate of PDA for morphological identification and to potato dextrose broth (PDB) to generate mycelium for subsequent DNA extraction and molecular identification. In addition, tissues with fungal pelotons were stored in 1.5-ml Eppendorf tubes at −80 °C for DNA extraction.
Morphological characterization
Isolates with hyphal characteristics of Rhizoctonia fungi (e.g., branching near the septum of the cells in vegetative hyphae, constriction of hyphae and formation of a septum at a short distance from the point of origin, and absence of clamp connections) were cultured on PDA at 25 °C in the dark (Sneh et al. 1991). Hyphae from each isolate were placed in a solution consisting of one drop of 0.5 % (w/v) Safranin O and one drop of 3 % KOH (Bandoni 1979) and examined with a compound light microscope with an attached digital camera to determine the number of nuclei per hyphal cell (Carling et al. 1999).
Seed harvest and symbiotic germination
Initially, six plants of A. formosanus Hayata from Wufong (Hsinchu) were cultured in a greenhouse at the National Chung Hsing University. Artificial cross-pollination was performed in six individual plants at the beginning of December 2011 and 2012. The pollinated inflorescences were covered with nylon mesh (48 μm) to avoid invasion of insects into the capsules. Capsules were harvested from 50 days after pollination. One capsule from each of three randomly selected plants was collected.
Seeds from the collected fruits were sown on the same day to avoid loss of viability. Capsules were surface-disinfested in 1 % NaOCl with 0.3 % of Tween 80 for 15 min, rinsed three times in sterile distilled water, and split in half with a sterile scalpel. Seeds were picked out of the capsules with sterilized forceps, and approximately 50 to 120 seeds were added to the surface of 0.25 % oatmeal medium (OMA) in a 6-cm-diameter Petri dish (Warcup 1981). A 1-cm3 block of mycelium obtained from an actively growing culture was placed in the center of the Petri dish on the surface of OMA. Each plate was inoculated with an isolate of Rhizoctonia fungi sampled from A. formosanus and also from other native orchids, including isolate Cno3-2 (accession no. JX514374 from Zeuxine sp.), Cno10-3 (accession no. JX514376 from Cheirostylis hungyehensis), CalS1-2 (accession no. JX514384 from Calanthe sylvatica), Eno3-3 (accession no. JX514377 from Goodyera procera), Sno6-1 (accession no. JX514393 from Cymbidium ensifolium), Sno3-3 (accession no. JX514385 from Liparis nakaharai), Sno5-12 (accession no. JX514386 from Cleisostoma paniculatum), and R02 (from Calanthe arisanensis kindly provided by Chang, Doris CN) that served as controls for the experiments. Modified Hyponex medium (Hy) (Chang and Chou 2001) and 1/2 strength Murashige-Skoog (MS) media (1962) were also used for asymbiotic seed germination and to assess seed vigor. Five replicates of each treatment were prepared for each isolate, and the experiment was repeated once. Plates were sealed with parafilm and incubated in darkness at 25 °C for 30 days to promote seed germination and initial protocorm development.
After 10 days of culture, seeds in the symbiotic and asymbiotic treatments were collected from the surface of agar plates, fixed overnight at 4 °C in 2 % paraformaldehyde plus 2.5 % glutaraldehyde in 0.2 M phosphate buffer at pH 7.0, rinsed three times in phosphate buffer, and placed in 1 % OsO4 for 2 h at 4 °C. Tissue was rinsed in phosphate buffer, dehydrated in a graded ethanol series, and embedded in LR white resin. For staining of prepared tissue sections, sections (1 μm) were stained with periodic acid-Schiff’s (PAS) reagent for bright-field microscopy. Carbohydrates (e.g., cellulose, starch, and glycogen) stain purplish red and nuclei may stain light pink (Ruzin 1999).
Germination (%) was determined 30 days after seed sowing. The process of symbiotic development from seed germination to protocorm development with differentiated shoots was divided into seven phases (Fig. 3) as follows: 0 = an undifferentiated embryo in the testa; I = embryo absorbed water and had slight swelling in the testa; II = embryo swelling was observed with the production of epidermal hairs that ruptured the testa; III = increased embryo swelling and rupturing of the testa that was <1,000 μm in length; IV = embryo was between 1,000 and 2,000 μm in length; V = embryo was between 2,000 and 3,000 μm in length; and VI = embryo length >3,000 μm. The emergence of the swollen embryo from the seed coat was used as the primary criterion to define seed germination (phase III).
DNA extraction, rDNA-ITS sequencing, and phylogenetic analyses
For DNA extraction, tissues with fungal pelotons and mycelium of each isolate from liquid culture were frozen at −80 °C before use. DNA was extracted from the frozen samples using the DNeasy Plant mini kit (Qiagen, Hilden, Germany). Total DNA was recovered in 50 μL of distilled deionized water or TE buffer and stored at −20 °C. The fungal rDNA-ITS sequence (ITS1-5.8S-ITS2) was amplified using primer sets ITS1/ITS4 (White et al. 1990) and ITS1-OF/ITS4-OF (Taylor and McCormick 2008).
PCR reactions were performed using 50 pM of each primer, 500 ng of template DNA, 2.5 units of Taq polymerase, 200 mM each of dNTPs, and 1.5 mM of MgCl2. Reactions were performed in a GeneAmpR PCR System 2700 (Applied Biosystems, Foster City, CA, USA) under the following thermoprofile: initial denaturation at 94 °C for 2 min, followed by 30 cycles of denaturation at 94 °C for 40 s, annealing at 55 °C for 60 s, extension at 72 °C for 60 s, and final extension at 72 °C for 5 min. PCR products were purified using the MinElute PCR Purification Kit (Qiagen). Clean samples were eluted with 50 μL sterile water and quantified for conducting sequencing reactions by using a NanoDropTM 1000 Spectrophotometer (ND-1000, Thermo Fisher Scientific, Waltham, MA). Cycle sequencing was conducted using BigDye version 3.1 chemistry, and sequencing was done on an ABI 3100 Genetic Analyzer (Applied Biosystems). Both strands of DNA were sequenced. The sequences obtained in this study are available from GenBank under the provided accession numbers (Table 1 and Supplemental data 1).
Sequences were subjected to a BLAST search (Altschul et al. 1979) using the National Center for Biotechnology Information database (http://www.ncbi.nih.gov/BLAST). Well-characterized and closely related taxa obtained from BLAST searches were also included in the phylogenetic analyses. Multiple sequence alignments of each ITS sequence were made with ClustalW (Thompson et al. 1994) packaged with BioEdit Sequence Alignment Editor (Ibis Biosciences, Carlsbad, CA, USA). Phylogenetic trees were generated using maximum parsimony (MP) analysis (MEGA 5) (Tamura et al. 2011). Genetic distances of MP search method were calculated using the Subtree-Pruning-Regrafting (SPR). One thousand bootstrap replicates were performed to estimate the node reliability of the phylogenetic trees. Bootstrap values >70 % were considered significant and represented well-supported groups (clades).
Colonization of endomycorrhizal fungi in tissue-cultured plantlets
A seed-derived A. formosanus plant with a well-developed rhizome and shoot was selected (Shiau et al. 2002), and leaves and roots were removed from the plant. A single rhizome was cut into several lengths (1.5 cm), and each rhizome contained a node that had the potential to differentiate and produce a new seedling. Each small rhizome was placed on the surface of MS medium containing 0.5 ppm 1-naphthaleneacetic acid (NAA) and 3 ppm 6-benzylaminopurine (BA) at 24 °C in the dark for 30 days (Ho et al. 1987). Tissue-cultured seedlings with uniform size (~1.5–2 cm) produced from each node were chosen for colonization experiments, and the asymbiotic (control) and symbiotic treatments were arranged in a completely randomized design with eight replicates of each treatment. The fungus was inoculated onto the center of a slant tube containing 0.25 % OMA (Warcup 1981) and allowed to colonize the entire medium before each tissue-cultured seedling was transferred to the base of the slant tube. There were eight replicate slant tubes per treatment and the experiment was repeated twice. Slant tubes were incubated in a growth chamber (12 h day/night photoperiod and daytime illuminance of approximately 3,500 lx corresponded to ca. 47.25 μmol m−2 s−1 using cool white fluorescent lamps) at 25 °C for 4 months (Ho et al. 1987; Shiau et al. 2002).
After 4 months, tissue-cultured seedlings were removed from each slant tube, and agar was carefully removed from the surface of the rhizome and roots and weighed to determine fresh weight. Generally, each plantlet had four to eight nodes. For our experiments, we randomly selected four plantlets in each of the fungal treatments to examine each node for fungal pelotons. Each node of an A. formosanus rhizome containing a branching root (>3 mm) was scored based on the colonized percentage of the entire surface (fungal pelotons) using category values from 0 to 4 (Fig. 6), where 0 = no fungal pelotons were observed, 1 = cortex was slightly colonized and formed a few pelotons at the junction of the rhizome and branching root; 2 = more fungal pelotons formed at the upper and lower site in the branching root; 3 = the basal part of the branching root (1–3 mm) was full of fungal pelotons, which have spread to the rhizome; and 4 = fungal pelotons were full of branching root and rhizome. From each root or rhizome with fungal colonization, a small piece of tissue (0.5 cm in length) was excised and macerated in 1 ml of sterilized distilled water with a sterilized glass rod to isolate the intracellular fungal pelotons. Individual pelotons were collected and plated on TWA and PDA to isolate the fungus.
Statistical analysis
The experimental data in mycorrhizal synthesis of tissue-cultured seedlings experiments were analyzed using SAS software version 9.4 (SAS Institute Inc., Cary, NC). The ordinal data of a 0-to-4 scale for fungal colonization were analyzed using the nonparametric methodology of Brunner et al. (2002) as described by Shah and Madden (2004). Rank assigned to each observation was determined by PROC RANK and then PROC MIXED to calculate test statistics and levels of significance. Estimated relative treatment effects were calculated using \( {\widehat{p}}_i = 1/N \) (\( {\overline{R}}_i - 1/2 \)) in which values are between 0 and 1, where i is the treatment, N is the number of observations, and \( \overline{R} \) is the mean rank. Ninety-five percent confidence intervals (CI) were generated using LD_CI macro (Brunner et al. 2002). A smaller value of \( {\widehat{p}}_i \) for a treatment indicates lower value for the degree of fungal colonization. Values of \( {\widehat{p}}_i \) between two treatments are significantly different if the CI for each \( {\widehat{p}}_i \) does not overlap.
Data on plant weight were analyzed by analysis of variance (ANOVA). Mean separation was performed by protected post hoc Tukey’s test (P = 0.05) after a significant F test in SAS ver. 9.4 (SAS Institute, Cary, NC) was determined.
Results
Endomycorrhizal fungi from A. formosanus
All samples of terrestrial A. formosanus collected from central and northern regions of Taiwan had fungal pelotons at the junction of the root and rhizome (Fig. 1). To make sure no pertinent information was lost, tissue and peloton isolation (Rasmussen 2002) and direct PCR (Taylor and McCormick 2008) were applied to check and identify the fungal pelotons that existed in cortical cells of A. formosanus. A. formosanus samples from a given location often yielded the same cultural type of Rhizoctonia, except for the sample from Alishan which contained two different cultural types of Rhizoctonia fungi (Table 1). A representative isolate of Rhizoctonia was selected from each sample site by using hyphal anastomosis and nuclear staining criteria (Sneh et al. 1991). These isolates showing unique cultural characteristics were subjected to further morphological and molecular characterization. In this study, five of the nine isolates sampled from native A. formosanus possessed multinucleate hyphal cells, while the remaining four isolates had binucleate hyphal cells. Among the latter, both isolates ANOF 6 and ANOF G2 were tentatively identified as Epulorhiza (teleomorph = Tulasnella) based on the colony and hyphal characteristics described by Currah and Zelmer (1992). No teleomorph stage of any isolate of Rhizoctonia was observed throughout the study.
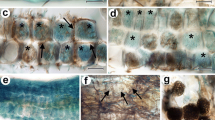
Light microscopic observation of root or rhizome sections of native A. formosanus Hayata (a) without staining (b–d) and stained with trypan blue (e–g). b–c Longitudinal and transverse sections showed that fungal pelotons (black arrowheads) formed at the junction of root and rhizome, and starch granules (white arrowhead) accumulated at a nearby vascular bundle in the rhizome. d Transverse sections of root close to the rhizome also formed fungal pelotons. e–f Transverse sections containing pelotons stained with trypan blue, pelotons are dark blue, and plant tissues are light blue. Fungal mycelia entered the cortex through a structure called “hair cushion” (asterisk) formed only at the junction of root and rhizome. g Fungal mycelia (small arrow) can be seen in a single rhizoid of the hair cushion (asterisk). Scale bars represent 2 cm (a), 1,000 μm (b–e), 400 μm (f), and 20 μm (g)
Phylogenetic analyses
Both ITS1/4 and ITS1-OF/ITS4-OF primer sets were used to amplify rDNA-ITS sequences of endomycorrhizal fungi observed in orchid tissues. Although some amplification of fungi in the phylum Ascomycota was observed (i.e., Fusarium spp.), Rhizoctonia fungi (teleomorphs = Tulasnella, Ceratobasidium, and Thanatephorus) were amplified selectively with primer set ITS1-OF/ITS4-OF which also exhibited more sensitivity than the ITS1/4 primer set in our study. Amplified rDNA-ITS sequences of isolates and uncultured isolates (direct amplification) were sequenced, and the results of BLAST searches were conducted and checked twice. In this study, we found that endomycorrhizal fungi could be easily isolated from the junction of root and rhizome of A. formosanus (Fig. 1) and successfully detected at this site by direct amplification with PCR.
The MP analysis divided the endomycorrhizal Rhizoctonia from native A. formosanus into three major clades (clades I to III) with high bootstrap values. These clades have been further analyzed and categorized into different AGs (Fig. 2), as rDNA-ITS sequences were proven to reflect the classification of AGs within Rhizoctonia spp. (Sharon et al. 2006; Sharon et al. 2008). Results of the BLAST searches are shown in Supplemental data 1. Clade I contained isolates from native A. formosanus in the northern regions of Taiwan, with the exception of ANOF 7 from the central regions (Alishan). Three sequences of uncultured isolates (TANOF D1, TANOF D2, and TANOF 8) and five sequences of isolates (ANOF 0, ANOF 4, ANOF D1, ANOF D2, and ANOF 7) in clade I shared high rDNA-ITS sequence similarity (97 to 99 % identity) with uncultured isolates of Rhizoctonia fungi in the Ceratobasidiaceae sampled from orchid (Goodyera spp.) roots (Shefferson et al. 2010). Phylogenetic analysis with maximum parsimony suggested that eight sequences grouped with Rhizoctonia solani AG-6 (teleomorph = Thanatephorus) (Pope and Carter 2001; Gonzalez et al. 2001) and formed a well-supported subgroup (subgroup 1) within clade I. The identification of isolates of R. solani AG-6 was also verified according to the anastomosis group criteria.
Phylogeny of Rhizoctonia isolates from orchids and related fungi based on rDNA-ITS sequences analysis using the maximum parsimony (MP) method. One ITS sequence from Sebacina vermifera (AF20272) was defined as an outgroup taxon. Numbers at the nodes indicate bootstrap values (1,000 replicates; only values above 70 % are presented) from the corresponding MP tree. The following sequences were direct detection from tissues containing fungal pelotons: TANOF 2, TANOF 3, TANOF D1, TANOF D2, TANOF 6, TANOF G2, TANOF 8, and TANOF 10
Both isolate ANOF 3 and uncultured isolate TANOF 3 in clade I were sampled from different tissues of the same individual plant. They were grouped in subgroup 2 with bootstrap support of greater than 70 %, sharing high sequence similarity (97–99 % identity) with isolates of binucleate Rhizoctonia AG-R (teleomorph = Ceratobasidium) previously deposited in GenBank (accession numbers HQ269817, HQ269823, and AB286942) (Sharon et al. 2008; Copes et al. 2011). Comparison of the rDNA-ITS sequences suggested that isolate ANOF 3 belongs to AG-R. Uncultured isolate TANOF 10 in clade I formed a well-supported subgroup (subgroup 3) and had sequence similarity to sequences from isolates of binucleate Rhizoctonia AG-P (GenBank accession numbers AB286941 and AB286938) with 94 % identity (Sharon et al. 2008) and binucleate Rhizoctonia AG-U (accession numbers HQ269810, HQ269820, and HQ269825) with 94 % identity (Copes et al. 2011). Subgroups 2 and 3 were closely related to each other and also related to eight other sequences in subgroup 1 obtained from A. formosanus (Fig. 2).
The rDNA-ITS sequences of isolates ANOF 2 and uncultured isolate TANOF 2 were sampled from different tissues of the same individual plant collected in the central region of Taiwan and grouped in clade II, showing 99 % identity to rDNA-ITS sequences from isolates of binucleate Rhizoctonia sp. AG-G (accession numbers DQ102399, DQ102401, DQ102402, AY927320, and AY927327) (Manici and Bonora 2007; Sharon et al. 2007). Isolate ANOF 2 was identified as Rhizoctonia AG-G (teleomorph = Ceratobasidium) based on hyphal anastomosis and rDNA-ITS sequence analyses.
Both isolate ANOF 6 and uncultured isolate TANOF 6 from native A. formosanus in the central regions of Taiwan (e.g., Nantou and Alishan) are grouped in clade III, showing 95 % sequence identity to sequences from three isolates of Tulasnella calospora previously deposited in GenBank (accession numbers GU166403, GU166407, and GU166410) (Nontachaiyapoom et al. 2010). Furthermore, isolates ANOF G2 and uncultured isolate TANOF G2 also showed high sequence similarity (98 % identity) to isolates of T. calospora (accession number GU166403), uncultured members of Tulasnellaceae (accession number JX024734) from Dactylorhiza spp. orchids with 99 % identity (Jacquemyn et al. 2012), and uncultured members of Tulasnellaceae (accession numbers JX545220 and JX545219) from Dendrobium spp. orchids with 98 % sequence identity (Xing et al. 2013). The sequence comparison supported that these isolates could be identified as T. calospora.
Symbiotic seed germination
The emergence of a swollen embryo from the seed coat was used as the criterion to define seed germination (phase III, Fig. 3). The embryo began to swell when hyphae invaded the basal cells (Fig. 4a). Hyphae penetrated into the inner cortical parenchyma and subepidermal parenchyma cells and formed pelotons (Fig. 4b). The meristematic region was not colonized (Fig. 4b).
Stages in the development of A. formosanus protocorms. a Phase 0, an ungerminated seed invested by membranous testa. b Phase I, embryo absorbed water and swelled slightly in testa. c Phase II, embryo swollen slightly was observed with production of an epidermal hairs that ruptured the testa. d Phase III, embryo was large enough to burst the testa and was less than 1,000 μm in length. e Phase IV, the length of the embryo was between 1,000 and 2,000 μm. f Phase V, the length of the embryo was between 2,000 and 3,000 μm. g Phase VI, the embryo was more than 3,000 μm in length. Scale bars represent 2,500 μm
Structures of A. formosanus protocorm after 10 days of culture (a–c) and 60 days of culture (d–f) with fungus (symbiotic) and noninoculated (asymbiotic) treatments. a Direct observation by light microscopy. b, c Sections were stained with periodic acid-Schiff (PAS) reagent. Carbohydrates stained purplish red and nuclei lightly stained. a, b Fungal mycelia have penetrated into basal cell(s) or rhizoid(s) (black arrowheads) and then into the inner cortical parenchymal cells and have formed fungal pelotons (white arrowheads). The meristematic region was not colonized (asterisk). c Non-inoculated treatment is the same size as the inoculated treatment but contains many granules (small arrowhead), which are almost absent in fungal treatments. d–f Protocorms growing on 1/2 strength MS (d) and 0.25 % OMA (e) without fungal inoculation were smaller and flatter than those in fungal treatments on 0.25 % OMA (f). Scale bar represents 50 μm (a–c) and 1,000 μm (d–f)
Germination for the asymbiotic seeds was 89 % on Hy medium, 75 % on 1/2 strength MS, and 13 % on 0.25 % OMA. However, most of the germinated seeds only grew to phase III (Fig. 5) after 30 days of incubation. Seeds sown on 0.25 % OMA without fungal inoculation remained in phase III after 60 days of culture (Fig. 4e). For the remaining two asymbiotic treatments (Hy medium and 1/2 strength MS), growth of protocorms into stages IV–V was observed after 60 days of culture (Fig. 4d), but they were smaller and flatter in width than those in the symbiotic treatment (0.25 % OMA) (Fig. 4f). Fungal pelotons formed in germinated A. formosanus seeds, indicating that germination was associated with the presence of the fungus (Figs. 4 and 5). Increased germination was observed with isolates found in clade I and ranged from 44 to 91 % compared to asymbiotic treatment (13 %) in 0.25 % OMA and promoted growth of protocorms to phases IV and V. However, isolate ANOF 7 in clade I from central regions of Taiwan (Alishan) did not increase seed germination (14 %).
Asymbiotic and symbiotic seed germination of A. formosanus Hayata, recorded 30 days after sowing seeds on 0.25 % oatmeal agar (OMA), 1/2 strength Murashige-Skoog (MS) medium, and Hyponex medium for asymbiotic control and on 0.25 % OMA for symbiotic treatments. Phase III was used to define whether a seed had germinated. Percentage of germinated seeds was presented as ( ) in phase III, (
) in phase III, ( ) in phase IV, (
) in phase IV, ( ) in phase V, and (
) in phase V, and ( ) in phase VI
) in phase VI
Both isolates ANOF G2 and ANOF 6 (teleomorph = Tulasnella) in clade III sampled from A. formosanus in central regions of Taiwan (Nantou and Alishan) did not increase seed germination (3–10 %) in 0.25 % OMA (Fig. 5). Isolates Sno3-1 and Sno6-1, which were closely related to isolates ANOF G2 and ANOF 6 in clade III, were sampled from the northern region (Sanxia) and also did not increase seed germination (0–2 %). Results suggested that isolates of T. calospora can form fungal pelotons in cortical cells of native A. formosanus growing in central regions but did not increase seed germination under our laboratory-based assay conditions. Isolate ANOF 2 in clade II, which was also sampled from A. formosanus in central Taiwan, increased seed germination (59 %); however, almost all the germinated seeds grew slowly into phase III after 30 days of co-culture compared to the other treatments with fungi in clade I (Fig. 5).
Colonization of endomycorrhizal fungi
In general, tissue-cultured seedling inoculated with any isolate of Rhizoctonia increased the fresh plant weight compared to asymbiotic treatment after 120 days of co-culture in vitro (Table 2). However, plant growth was affected by different isolates of Rhizoctonia fungi. By sectioning each part of rhizome and roots, the results showed that plantlets had massive pelotons at the junction of root and rhizome (Fig. 6). Calculating the relative treatment effects for a 0-to-4 scale of fungal colonization suggested that isolates in clade I significantly promote the growth of seedlings with a relatively low level of colonization (\( {\widehat{p}}_i = 0.30\hbox{--} 0.47 \)) compared to the other isolates in clades II (\( {\widehat{p}}_i = 0.63\hbox{--} 0.82 \)) and III (\( {\widehat{p}}_i = 0.63\hbox{--} 0.75 \)) with 95 % confidence interval. Our data also indicated that the increase due to the treatment effects on fungal colonization was negatively correlated (r = −0.8801) with fresh weight of tissue-cultured seedling (Supplemental data II).
Hand sections of symbiotic and asymbiotic treatments at each node of A. formosanus showing different levels of colonization after 120 days of culture. a 0 = fungal pelotons were absent and white arrowhead stands for starch granules. b 1 = cortex was slightly colonized and formed a few pelotons (black arrowheads) at the junction of rhizome and branching root. c 2 = more fungal pelotons have formed at the upper and lower site in the branching root. d 3 = the basal part of the branching root (1–3 mm) was full of fungal pelotons, which have spread to rhizome. e 4 = branching root and rhizome were full of fungal pelotons. Scale bars represent 1,000 μm
Discussion
Characterization and identification of endomycorrhizal fungi
The endomycorrhizal fungi sampled from green orchids has been recognized as belonging to the form genus of Rhizoctonia that include species of Thanatephorus, Ceratobasidium, Tulasnella, and Sebacina (Sneh et al. 1991; Roberts 1999; Dearnaley 2007; Smith and Read 2008). All isolates from fungal pelotons in native A. formosanus belonged to the form genus Rhizoctonia, including Thanatephours, Ceratobasidium, and Tulasnella. In our study, none of the isolates from A. formosanus were identified as Sebacina, which can form endomycorrhizae with several green orchids in Australia, especially in fully mycoheterotrophic orchids (McKendrick et al. 2002; Selosse et al. 2002; Taylor et al. 2003).
The method of AG determination is a commonly used procedure to characterize and identify isolates of Ceratobasidium and Thanatephorus since the sexual stage of these fungi is often challenging to induce in the laboratory (Sneh et al. 1991; Carling et al. 1999). Many AGs have been found on orchids and increased our understanding of orchid mycorrhizal symbionts (Ramsay et al. 1987). Currently, the multinucleate R. solani (teleomorph = Thanatephorus) contains 14 AGs (González García et al. 2006), while the binucleate Rhizoctonia spp. (teleomorph = Ceratobasidium) contains at least 16 AGs (Sharon et al. 2008). Three AGs of Thanatephorus (AG-5, AG-6, and AG-12) and four AGs of Ceratobasidium (AG-A, AG-C, AG-E, and AG-I) have been reported to form mycorrhizal associations with orchids in nature (Sneh et al. 1991; Carling et al. 1999; Pope and Carter 2001). Fungi in the same AG may possess similar morphological characteristics or specialized host ranges (Sneh et al. 1991; Keijer et al. 1997). With molecular analysis and AG determination, isolates from native A. formosanus were identified as multinucleate R. solani AG-6, binucleate Rhizoctonia sp. AG-G, AG-P, and AG-R, and Rhizoctonia repens (teleomorph = T. calospora) (Fig. 2). Sharon et al. (2008) have suggested that AG-P are closely related with AG-U based on rDNA-ITS sequence similar and may represent the same anastomosis group. However, additional analyses with multiple genetic loci are needed to better substantiate these findings.
Previously on the studies on phylogenetic relationships of Rhizoctonia fungi, the genus Ceratobasidium was paraphyletic and R. solani AG-6 often grouped with isolates of Ceratobasidium (Gonzalez et al. 2001; Moncalvo et al. 2006). These observations were similar to those reported by Sharon et al. (2008). R. solani AG-6 was the predominant mycorrhizal fungus isolated from fungal pelotons of A. formosanus in the northern regions of Taiwan and represented 67 % of the sample. In this study, we found that Ceratobasidium (i.e., AG-R and AG-P) was grouped in clade I with AG-6 and all of isolates from northern regions except isolate ANOF 7 from central regions. The results suggest that isolates in clade I represent the predominant endomycorrhizal symbionts associated with A. formosanus in nature (Fig. 2). Interestingly, some isolates of AG-P and AG-R were previously recorded as plant pathogenic fungi (Sharon et al. 2008; Yang et al. 2008), but certain isolates were also used for protecting plants against pathogenic fungi (Sneh and Ichielevich-Auster 1998; Jabaji-Hare and Neate 2005). A recent study reported from Veldre et al. (2013) suggested that the sequences of Rhizoctonia fungi derived from soil (representing putative saprobes) and orchid mycorrhizae clustered together, but remained distinct from pathogens, and that autotrophic orchids generally form root symbioses with available Ceratobasidiaceae isolates in soil.
Isolates ANOF 2 (AG-G) and both isolates of ANOF 6 and ANOF D2 (Tulasnella) from the central regions of Taiwan were distributed in clades II and III, respectively. Isolate ANOF 7 (AG-6) in clade I was also sampled from A. formosanus in central regions of Taiwan. It cannot be stated with certainty that the diversity of Rhizoctonia spp. in A. formosanus from central regions was different from northern regions due to our limited sample size as native A. formosanus is challenging to collect in central Taiwan. However, the environment for plant growth (i.e., substrate and seasonal changes) has been reported to influence the fungal preference and diversity of orchid-associated fungi (Porras-Alfaro and Bayman 2007; Selosse and Roy 2009; Otero et al. 2011).
Symbiotic seed germination
The method of symbiotic seed germination demonstrates relationships between seeds and endomycorrhizal fungi. A. formosanus embryo formed fungal pelotons at the mycorrhizome after 10 days of culture (Fig. 4). The disappearance of starch grains in amyloplasts has frequently been reported by several researchers when they observed that a mycorrhizal fungus invaded cells of orchid roots and protocorms (Richardson et al. 1992; Uetake et al. 1992).
Analyzing the data of seed germination (Fig. 5) with the result of phylogenetic analysis (Fig. 2) suggested that isolates sampled from A. formosanus in northern regions of Taiwan in clade I (i.e., subgroups 1 to 3), except for isolate ANOF 7, effectively increase seed germination and promote the growth of protocorm into phases IV–V. It is worth mentioning that both isolates CalS1-2 and Cno10-3 were sampled previously from other green orchids. Our data suggests that A. formosanus seeds possess mycorrhizal preference with isolates in clade I under in vitro experimental conditions. Additionally, isolates of Ceratobasidium AG-A in northern regions of Taiwan also showed symbiotic ability with A. formosanus seeds similar to the responses observed for fungi in clade I. The ability of isolates of Ceratobasidium to stimulate orchid seed germination and seedling growth was predicted by phylogenetic relationships (Otero et al. 2005), suggesting a genetic basis for mycorrhizal symbiosis. Nevertheless, isolate ANOF 7 in clade I (subgroup 1) from the central region of Taiwan (Alishan) did not increase seed germination, implying that isolates from different geographic regions varied in symbiotic ability within the population of A. formosanus. However, additional studies are needed to ascertain the genetic diversity of isolates sampled from different geographic locations to increase the predictability of phylogenetic-based methods for identifying potential endomycorrhizal symbionts.
Tulasnelloid fungi especially species of Tulasnella are well-studied endomycorrhizal fungus associated with many green orchids and promote seed germination (Suarez et al. 2006). Our results suggested that isolates of AG-G in clade II and Tulasnella species in clade III sampled from A. formosanus in central regions and other green orchids in northern regions did not increase germination of A. formosanus seeds and promote increased plant development (Fig. 5). Although seeds inoculated with ANOF 2 expressed a moderate increase in seed germination, only a few seeds grew to phase IV after 60 days of inoculation and exhibited limited growth. Therefore, the mycorrhizal specificity of A. formosanus was also possibly associated with the geographic origin of the orchid seeds and/or Rhizoctonia fungus. This difference in the mycorrhizal specificity of A. formosanus may also be explained by the fact that seeds instead of adult plants were utilized for the symbiotic experiments, i.e., the difference in specificity was due to physiological differences in the developmental stage of the orchid (Masuhara et al. 1993). Although our data suggest that mycorrhizal specificity of A. formosanus seeds with Rhizoctonia fungi in clade I may occur, further studies of seed germination in the same or other geographic locations under in situ conditions are warranted to better document and substantiate this observation.
Colonization of endomycorrhizal fungi
Mycorrhizal specificity in relation to orchid diversity has been studied in field conditions (Masuhara and Katsuya 1994; McKendrick et al. 2002; McCormick et al. 2004). Because the populations of native A. formosanus have greatly decreased the occurrence of this species in central and northern Taiwan, we conducted a synthesis study by using tissue-cultured seedlings when a pure culture can be obtained from fungal pelotons to further demonstrate functionality and fungal–plant interactions. As mentioned above, isolates in clade II and clade III expressed low or no ability to increase seed germination in vitro. However, when using tissue-cultured seedlings, all isolates we studied formed fungal pelotons at the junction of roots and rhizome. The results showed that mycorrhizal specificity of A. formosanus seeds was different from tissue-cultured seedlings.
A. formosanus will increase the efficacy of photosynthesis by inoculating endomycorrhizal Rhizoctonia fungi (Cheng and Chang 2009). Indeed, when a protocorm grows into a seedling with a differentiated green shoot, the presence of chlorophyll may not be sufficient to enable full autotrophy (Smith and Read 2008). It has been demonstrated previously that the pathway for carbon transfer from the fungus to plant is retained into adulthood in autotrophic species of orchids and suggests that green orchids are partially mycoheterotrophic (“mixotrophic”) (Gebauer and Meyer 2003; Julou et al. 2005; Cameron et al. 2006). However, it is unlikely that endomycorrhizal fungi derive other nutritional benefits from their association with the orchid (Leake 2005).
In general, all isolates examined in this study significantly increased the fresh weight of tissue-cultured seedlings compared to the asymbiotic treatment (Table 2), irrespective of their genetic relatedness, geographic origin, and ability to promote seed germination of A. formosanus. However, we found that fungal colonization was negatively correlated with fresh weight of plantlets (r = −0.8801) (Supplemental II). Isolates in clade I (i.e., subgroups 1 to 3) increased seed germination and significantly promoted the growth of tissue-cultured seedlings with low relative treatment effects ranging from 0.30 to 0.47. Although the fresh weights of plantlets with high relative treatment effects caused by isolate Eno3-3 (\( {\widehat{p}}_i = 0.82 \)) sampled from orchid G. procera and isolate Sno5-12 (\( {\widehat{p}}_i = 0.87 \)) from C. paniculatum were greatly decreased, all plants were alive and had greener leaves than those in asymbiotic treatments. Endomycorrhizal Rhizoctonia fungi might be considered to be parasitic on orchids when the net cost of the symbiosis exceeds any net benefits, and parasitism can be induced developmentally, environmentally, or possibly genotypically. Therefore mycorrhizal symbiosis was described as a mutualism–parasitism continuum (Johnson et al. 1997; Newton et al. 2010).
The colonization by isolates in clade II (\( {\widehat{p}}_i = 0.63\hbox{--} 0.82 \)) and clade III (\( {\widehat{p}}_i = 0.63\hbox{--} 0.75 \)) was similar and limited the growth of A. formosanus plantlets, which did not decay and continued growing with a slight brown coloration at the base of the rhizome. This basal part of rhizome had cortical cells that were completely colonized with fungal pelotons. Fungitoxic phytoalexins, cell wall-degrading enzymes, and species of reactive oxygen can occur in tubers or rhizomes and are hypothesized to regulate or prevent fungal colonization (Stoessl and Arditti 1984; Beyrle et al. 1995). However, their role in the process of peloton digestion and formation is not clear (Smith and Read 2008). Paduano et al. (2011) demonstrated that pectin is exclusively found in the interface formed around Ceratobasidium sp. but not Russula sp. and that the native green orchid Limodorum abortivum expressed different cellular responses to the mycorrhizal fungi. Using tissue-cultured seedlings as symbiotic materials for further investigation of the fungal–plant interface associated with different levels of colonization under laboratory condition may potentially uncover important key interactions for partially mycoheterotrophic A. formosanus in regulating fungal colonization, growth, and development.
Conclusion
The population of native A. formosanus is associated with endomycorrhizal Rhizoctonia fungi, especially R. solani AG-6. These isolates were divided into clades I to III and differed in their ability to colonize orchid seeds and tissue-cultured seedlings in vitro assays. Our data suggest that the majority of isolates sampled from native A. formosanus in northern regions of Taiwan in clade I have ability to increase seed germination successfully and also effectively promote the growth of tissue-cultured seedlings with a low relative treatment effects for the degree of colonization. By comparison with isolates in clade I, an increased value of the relative treatment effect exhibited by isolates in clades II and III reduced the growth of A. formosanus. Our findings will be potentially useful for medicinal orchid production and conservation of A. formosanus in diverse and ecologically sensitive native habitats.
References
Altschul SF, Madden TL, Schäffer AA, Zhang JH, Zhang Z, Miller W, Lipman DJ (1979) Gapped BLAST and PSI–BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bandoni RJ (1979) Safranin O as a rapid nuclear stain for fungi. Mycologia 71:873–874
Beyrle HF, Smith SE, Peterson RL, Franco CMM (1995) Colonization of Orchis morio protocorms by a mycorrhizal fungus: effects of nitrogen nutrition and glyphosate in modifying the responses. Can J Bot 73:1128–1140
Bougoure JJ, Ludwig M, Brundrett M, Grierson P (2009) Identity and specificity of the fungi forming mycorrhizas with the rare myco-heterotrophic orchid Rhizanthella gardneri. Mycol Res 113:1097–1106
Brunner E, Domhof S, Langer F (2002) Nonparametric analysis of longitudinal data in factorial experiments. Wiley, New York
Cameron DD, Leake JR, Read DJ (2006) Mutualistic mycorrhiza in orchids: evidence from plant–fungus carbon and nitrogen transfers in the green-leaved terrestrial orchid Goodyera repens. New Phytol 171:405–416
Carling DE, Pope EJ, Brainard KA, Carter DA (1999) Characterization of mycorrhizal isolates of Rhizoctonia solani from an orchid, including AG-12, a new anastomosis group. Phytopathology 89:942–946
Chang SF (1999) Integrated control of basal stem rot and stem rot of Taiwan Anoectochilus. Taichung, Taiwan: National Chung Hsing University, Master thesis. [http://nchuir.lib.nchu.edu.tw/handle/309270000/113175]
Chang CN, Chou LC (2001) Seed germination of Haemaria discolor var. dawsoniana and the use of mycorrhizae. Symbiosis 30:29–40
Chang CN, Chou LC, Lee GC (2007) New cultivation methods for Anoectochilus formosanus Hayata. Orchid Sci Biotech 1:56–60
Cheng SF, Chang DCN (2009) Growth responses and changes of active components as influenced by elevations and orchid mycorrhizae on Anoectochilus formosanus Hayata. Bot Stud 50:459–466
Copes WE, Rodriguez-Carres M, Toda T, Rinehart TA, Cubeta MA (2011) Seasonal prevalence of species of binucleate Rhizoctonia fungi in growing medium, leaf litter, and stems of container-grown Azalea. Plant Dis 95:705–711
Currah RS, Zelmer C (1992) A key and notes for the genera of fungi mycorrhizal with orchids and a new species of the genus Epulorhiza. Rep Tottori Mycol Inst 30:43–59
Dearnaley JDW (2007) Further advances in orchid mycorrhizal research. Mycorrhiza 17:475–486
Dearnaley JDW, Martos F, Swlosse MA (2012) 12 orchid mycorrhizas: molecular ecology, physiology, evolution and conservation aspects. In: Hock B (ed) Fungal associations. The Mycota. A comprehensive treatise on fungi as experimental systems for basic and applied research, vol 9. Springer, Heidelberg, pp 207–230
Gebauer G, Meyer M (2003) N-15 and C-13 natural abundance of autotrophic and myco-heterotrophic orchids provides insight into nitrogen and carbon gain from fungal association. New Phytol 160:209–223
González García V, Onco P, Rubio Susan V (2006) Biology and systematics of the form genus Rhizoctonia. Span J Agric Res 4:55–79
Gonzalez D, Carling DE, Kuninaga S, Vilgalys R, Cubeta MA (2001) Ribosomal DNA systematics of Ceratobasidium and Thanatephorus with Rhizoctonia anamorphs. Mycologia 93:1138–1150
Gutiérrez RMP (2010) Orchids: a review of uses in traditional medicine, its phytochemistry and pharmacology. J Med Plants Res 4:592–638
Ho CK, Chang SH, Chen ZZ (1987) Tissue culture and acclimatization in Anoectochilus formosanus Hay. Bull Taiwan For Res Inst 2:83–104
Huang LW, Wang CJ, Lin YS, Chung WC, Chung WH (2014) Stem rot of jewel orchids caused by a new forma specialis, Fusarium oxysporum f. sp. anoectochili in Taiwan. Plant Pathol 63:539–547
Irwin MJ, Bougoure JJ, Dearnaley JDW (2007) Pterostylis nutans (Orchidaceae) has a specific association with two Ceratobasidium root associated fungi across its range in Eastern Australia. Mycoscience 48:231–239
Jabaji-Hare S, Neate SM (2005) Nonpathogenic binucleate Rhizoctonia spp. and benzothiadiazole protect cotton seedlings against Rhizoctonia damping-off and Alternaria leaf spot in cotton. Phytopathology 95:1030–1036
Jacquemyn H, Deja A, De hert K, Cachapa Bailarote B, Lievens B (2012) Variation in mycorrhizal associations with tulasnelloid fungi among populations of five Dactylorhiza species. PLoS ONE 7:e42212
Johnson N, Graham JH, Smith F (1997) Functioning of mycorrhizal associations along the mutualism–parasitism continuum. New Phytol 135:575–585
Julou T, Burghardt B, Gebauer G, Berveiller D, Damesin C, Selosse MA (2005) Mixotrophy in orchids: insights from a comparative study of green individuals and nonphotosynthetic individuals of Cephalanthera damasonium. New Phytol 166:639–653
Keijer J, Korsman MG, Dullemans AM, Houterman PM, de Bree J, Van Silfhout CH (1997) In vitro analysis of host plant specificity in Rhizoctonia solani. Plant Pathol 46:659–669
Kennedy AH, Taylor DL, Watson LE (2011) Mycorrhizal specificity in the fully mycoheterotrophic Hexalectris Raf. (Orchidaceae: Epidendroideae). Mol Ecol 20:1303–1316
Leake JR (2005) Plants parasitic on fungi: unearthing the fungi in myco-heterotrophs and debunking the ‘saprophytic’ plant myth. Mycologist 19:113–122
Ma M, Tan TK, Wong SM (2003) Identification and molecular phylogeny of Epulorhiza isolates from tropical orchids. Mycol Res 107:1041–1049
Manici LM, Bonora P (2007) Molecular genetic variability of Italian binucleate Rhizoctonia spp. isolates from strawberry. Eur J Plant Pathol 118:31–42
Masuhara G, Katsuya K (1994) In situ and in vitro specificity between Rhizoctonia spp. and Spiranthes sinensis (Persoon) Ames, var. amoena (M. Bieberstein) Hara (Orchidaceae). New Phytol 127:711–718
Masuhara G, Katsuya K, Yamaguchi K (1993) Potential for symbiosis of Rhizoctonia solani and binucleate Rhizoctonia with seeds of Spiranthes sinensis var. amoena in vitro. Mycol Res 97:746–752
McCormick MK, Whigham DF, O'Neill J (2004) Mycorrhizal diversity in photosynthetic terrestrial orchids. New Phytol 163:425–438
McKendrick S, Leake J, Taylor DL, Read D (2002) Symbiotic germination and development of the myco-heterotrophic orchid Neottia nidus-avis in nature and its requirement for locally distributed Sebacina spp. New Phytol 154:233–247
Moncalvo JM, Nilsson RH, Koster B, Dunham SM, Bernauer T, Matheny PB, Porter TM, Margaritescu S, Weiß M, Garnica S (2006) The cantharelloid clade: dealing with incongruent gene trees and phylogenetic reconstruction methods. Mycologia 98:937–948
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Newton AC, Fitt BD, Atkins SD, Walters DR, Daniell TJ (2010) Pathogenesis, parasitism and mutualism in the trophic space of microbe-plant interactions. Trends Microbiol 18:365–373
Nontachaiyapoom S, Sasirat S, Manoch L (2010) Isolation and identification of Rhizoctonia-like fungi from roots of three orchid genera, Paphiopedilum, Dendrobium, and Cymbidium, collected in Chiang Rai and Chiang Mai provinces of Thailand. Mycorrhiza 20:459–471
Otero JT, Ackerman JD, Bayman P (2002) Diversity and host specificity of endophytic Rhizoctonia-like fungi from tropical orchids. Am J Bot 89:1852–1858
Otero JT, Bayman P, Ackerman JD (2005) Variation in mycorrhizal performance in the epiphytic orchid Tolumnia variegata in vitro: the potential for natural selection. Evol Ecol 19:29–43
Otero JT, Thrall PH, Clements M, Burdon JJ, Miller JT (2011) Codiversification of orchids (Pterostylidinae) and their associated mycorrhizal fungi. Aust J Bot 59:480–497
Paduano C, Rodda M, Ercole E, Girlanda M, Perotto S (2011) Pectin localization in the Mediterranean orchid Limodorum abortivum reveals modulation of the plant interface in response to different mycorrhizal fungi. Mycorrhiza 21:97–104
Pope EJ, Carter DA (2001) Phylogenetic placement and host specificity of mycorrhizal isolates belonging to AG-6 and AG-12 in the Rhizoctonia solani species complex. Mycologia 93:712–719
Porras-Alfaro A, Bayman P (2007) Mycorrhizal fungi of Vanilla: diversity, specificity and effects on seed germination and plant growth. Mycologia 99:510–525
Ramsay RR, Sivasithamparam K, Dixon KW (1987) Anastomosis groups among Rhizoctonia-like endophytic fungi in southwestern Australian Pterostylis species (Orchidaceae). Lindleyana 2:161–166
Rasmussen HN (2002) Recent developments in the study of orchid mycorrhiza. Plant Soil 244:149–163
Rasmussen HN, Rasmussen FN (2007) Trophic relationships in orchid mycorrhiza—diversity and implications for conservation. Lankesteriana 7:334–341
Richardson KA, Peterson RL, Currah RS (1992) Seed reserves and early symbiotic protocorm development of Platanthera hyperborea (Orchidaceae). Can J Bot 70:291–300
Roberts P (1999) Rhizoctonia-forming fungi. The Herbarium, Royal Botanic Gardens, Kew
Roy M, Yagame T, Yamato M, Iwase K, Heinz C, Faccio A, Bonfante P, Selosse MA (2009) Ectomycorrhizal Inocybe species associate with the mycoheterotrophic orchid Epipogium aphyllum but not its asexual propagules. Ann Bot 104:595–610
Ruzin SE (1999) Plant microtechnique and microscopy. Oxford University Press, Oxford
Selosse MA, Roy M (2009) Green plants that feed on fungi: facts and questions about mixotrophy. Trends Plant Sci 14:64–70
Selosse MA, Bauer R, Moyersoen B (2002) Basal hymenomycetes belonging to the Sebacinaceae are ectomycorrhizal on temperate deciduous trees. New Phytol 155:183–195
Selosse MA, Faccio A, Scappaticci G, Bonfante P (2004) Chlorophyllous and achlorophyllous specimens of Epipactis microphylla, (Neottieae, Orchidaceae) are associated with ectomycorrhizal septomycetes, including truffles. Microb Ecol 47:416–426
Shah DA, Madden LV (2004) Nonparametric analysis of ordinal data in designed factorial experiments. Phytopathology 94:33–43
Sharon M, Kuninaga S, Hyakumachi M, Sneh B (2006) The advancing identification and classification of Rhizoctonia spp. using molecular and biotechnological methods compared with the classical anastomosis grouping. Mycoscience 47:299–316
Sharon M, Freeman S, Kuninaga S, Sneh B (2007) Genetic diversity, anastomosis groups and virulence of Rhizoctonia spp. from strawberry. Eur J Plant Pathol 117:247–265
Sharon M, Kuninaga S, Hyakumachi M, Naito S, Sneh B (2008) Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience 49:93–114
Shefferson RP, Weiß M, Kull T, Taylor DL (2005) High specificity generally characterises mycorrhizal association in rare lady’s slipper orchids, genus Cypripedium. Mol Ecol 14:613–626
Shefferson RP, Cowden CC, McCormick MK, Yukawa T, Ogura-Tsujita Y, Hashimoto T (2010) Evolution of host breadth in broad interactions: mycorrhizal specificity in East Asian and North American rattlesnake plantains (Goodyera spp.) and their fungal hosts. Mol Ecol 19:3008–3017
Shiau YJ, Sagare AP, Chen UC, Yang SR, Tsay HS (2002) Conservation of Anoectochilus formosanus Hayata by artificial cross-pollination and in vitro culture of seeds. Bot Bull Acad Sin 43:123–130
Smith SE, Read DJ (2008) Mycorrhizal symbiosis, 3rd edn. Academic, London
Sneh B, Ichielevich-Auster M (1998) Induced resistance of cucumber seedlings caused by some non-pathogenic Rhizoctonia (np-R) isolates. Phytoparasitica 26:27–38
Sneh B, Burpee L, Ogoshi A (1991) Identification of Rhizoctonia species. American Phytopathological Society, St Paul, Minnesota
Stoessl A, Arditti J (1984) Orchid phyloalexins. In: Arditti J (ed) Orchid biology, reviews and perspectives III. Cornell University Press, Ithaca
Suarez JP, Weiss M, Abele A, Garnica S, Oberwinkler F, Kottke I (2006) Diverse tulasnelloid fungi form mycorrhizas with epiphytic orchids in an Andean cloud forest. Mycol Res 110:1257–1270
Swarts ND, Dixon KW (2009) Terrestrial orchid conservation in the age of extinction. Ann Bot 104:543–556
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Taylor DL, McCormick MK (2008) Internal transcribed spacer primers and sequences for improved characterization of basidiomycetous orchid mycorrhizas. New Phytol 177:1020–1033
Taylor DL, Bruns TD, Szaro TM, Hodges SA (2003) Divergence in mycorrhizal specialization within Hexalectris spicata (Orchidaceae), a nonphotosynthetic desert orchid. Am J Bot 90:1168–1179
Thompson JD, Higgins DG, Gibson TJ (1994) ClustalW: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tseng CC, Shang HF, Wang LF, Su B, Hsu CC, Kao HY, Cheng KT (2006) Antitumor and immunostimulating effects of Anoectochilus formosanus Hayata. Phytomedicine 13:366–370
Uetake Y, Kobayashi K, Ogoshi A (1992) Ultrastructural changes during the symbiotic development of Spiranthes sinensis (Orchidaceae) protocorms associated with binucleate Rhizoctonia anastomosis group C. Mycol Res 96:199–209
Veldre V, Abarenkov K, Bahram M, Martos F, Selosse MA, Tamm H, Kõljalg U, Tedersoo L (2013) Evolution of nutritional modes of Ceratobasidiaceae (Cantharellales, Basidiomycota) as revealed from publicly available ITS sequences. Fungal Ecol 6:256–268
Warcup J (1981) The mycorrhizal relationships of Australian orchids. New Phytol 87:371–381
White TJ, Bruns TD, Lee SB, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Wu JB, Lin WL, Hsieh CC, Ho HY, Tsay HS, Lin WC (2007) The hepatoprotective activity of kinsenoside from Anoectochilus formosanus. Phytother Res 21:58–61
Xing XX, Ma X, Deng Z, Chen J, Wu F, Guo S (2013) Specificity and preference of mycorrhizal associations in two species of the genus Dendrobium (Orchidaceae). Mycorrhiza 23:317–324
Yagame T, Yamato M, Suzuki A, Iwase K (2008) Ceratobasidiaceae mycorrhizal fungi isolated from non-photosynthetic orchid Chamaegastrodia sikokiana. Mycorrhiza 18:97–101
Yagame T, Orihara T, Selosse MA, Yamato M, Iwase K (2012) Mixotrophy of Platanthera minor, an orchid associated with ectomycorrhiza-forming Ceratobasidiaceae fungi. New Phytol 193:178–187
Yang G, Conner R, Cai H, Li F, Chen Y (2008) First report of rhizome blight of ginger caused by binucleate Rhizoctonia AG-R in China. Plant Dis 92:312–312
Zettler LW, Poulter SB, McDonald KI, Stewart SL (2007) Conservation-driven propagation of an epiphytic orchid (Epidendrum nocturnum) with a mycorrhizal fungus. HortSci 42:135–139
Acknowledgments
We are grateful to Mr. Jin-Fu Xu for providing the orchid species, Dr. Tzu-Shing Deng and Dr. Chi-Ni Hsia for their advice on plant tissue culturing techniques, and Dr. Hao-Wei Kao for guidance on the phylogenetic analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Highlights
• We characterized the endomycorrhizal Rhizoctonia fungi in native A. formosanus.
• R. solani AG-6 are common species isolated from fungal pelotons in native A. formosanus.
• Isolates of AG-G, P, and R represent the first occurrence in the Orchidaceae.
• Rhizoctonia fungi in clade I increased seed germination and promoted seedling growth.
• Fresh plant weight and fungal colonization were negatively correlated.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Jiang, JH., Lee, YI., Cubeta, M.A. et al. Characterization and colonization of endomycorrhizal Rhizoctonia fungi in the medicinal herb Anoectochilus formosanus (Orchidaceae). Mycorrhiza 25, 431–445 (2015). https://doi.org/10.1007/s00572-014-0616-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-014-0616-1